Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities - PubMed (original) (raw)
. 2009 Dec;75(23):7537-41.
doi: 10.1128/AEM.01541-09. Epub 2009 Oct 2.
Sarah L Westcott, Thomas Ryabin, Justine R Hall, Martin Hartmann, Emily B Hollister, Ryan A Lesniewski, Brian B Oakley, Donovan H Parks, Courtney J Robinson, Jason W Sahl, Blaz Stres, Gerhard G Thallinger, David J Van Horn, Carolyn F Weber
Affiliations
- PMID: 19801464
- PMCID: PMC2786419
- DOI: 10.1128/AEM.01541-09
Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities
Patrick D Schloss et al. Appl Environ Microbiol. 2009 Dec.
Abstract
mothur aims to be a comprehensive software package that allows users to use a single piece of software to analyze community sequence data. It builds upon previous tools to provide a flexible and powerful software package for analyzing sequencing data. As a case study, we used mothur to trim, screen, and align sequences; calculate distances; assign sequences to operational taxonomic units; and describe the alpha and beta diversity of eight marine samples previously characterized by pyrosequencing of 16S rRNA gene fragments. This analysis of more than 222,000 sequences was completed in less than 2 h with a laptop computer.
Figures
FIG. 1.
Description and comparison of the eight samples analyzed by Sogin et al. (27). The dendrogram to the left represents the similarity of the samples based on the membership-based Jaccard coefficient calculated using Chao1 estimated richness values. The dendrogram on the right represents the similarity of the samples based on the structure-based θYC coefficient. The distance from the tip of the dendrogram to the root is 0.50 for both trees.
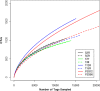
FIG. 2.
Rarefaction curves describing the dependence of discovering novel OTUs as a function of sampling effort for OTUs defined at a 0.10 distance cutoff. The curves for FS312 and FS396 climb to 3,095 and 2,804 OTUs after sampling of 54,894 and 80,769 sequences, respectively.
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials