Functions and regulation of RNA editing by ADAR deaminases - PubMed (original) (raw)
Review
Functions and regulation of RNA editing by ADAR deaminases
Kazuko Nishikura. Annu Rev Biochem. 2010.
Abstract
One type of RNA editing converts adenosines to inosines (A-->I editing) in double-stranded RNA (dsRNA) substrates. A-->I RNA editing is mediated by adenosine deaminase acting on RNA (ADAR) enzymes. A-->I RNA editing of protein-coding sequences of a limited number of mammalian genes results in recoding and subsequent alterations of their functions. However, A-->I RNA editing most frequently targets repetitive RNA sequences located within introns and 5' and 3' untranslated regions (UTRs). Although the biological significance of noncoding RNA editing remains largely unknown, several possibilities, including its role in the control of endogenous short interfering RNAs (esiRNAs), have been proposed. Furthermore, recent studies have revealed that the biogenesis and functions of certain microRNAs (miRNAs) are regulated by the editing of their precursors. Here, I review the recent findings that indicate new functions for A-->I editing in the regulation of noncoding RNAs and for interactions between RNA editing and RNA interference mechanisms.
Figures
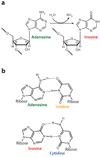
Figure 1
Deamination of adenosine to inosine by ADAR. (a) A hydrolytic deamination reaction converts adenosine to inosine. (b) Adenosine base pairs with uridine, whereas inosine base pairs, as if it were guanosine, in a Watson-Crick-bonding configuration with cytidine.
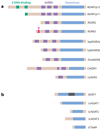
Figure 2
ADAR and ADAT family members. (a) The ADAR family is shown. Three vertebrate ADAR family members (ADAR1–3) are known. Two ADAR1 translation products (ADAR1p150 and ADAR1p110 isoforms) are known. Vertebrate ADARs, squid SqADAR2a and SqADAR2b (splicing isoforms), Drosophila melanogaster dADAR, and two C. elegans members (_Ce_ADR1 and _Ce_ADR2) share common functional domains: two to three repeats of the dsRNA-binding domain (dsRBD) and a catalytic deaminase domain. Certain structural features, such as Z-DNA-binding domains and the arginine-rich R domain, are unique to particular ADAR members. (b) The ADAT family is shown. Presented is the only known mammalian ADAT family member (ADAT1), its yeast homolog (_Sc_ADAT1), two additional yeast ADAT families (_Sc_ADAT2 and _Sc_ADAT3), and the only known bacterial ADAT family member (_Ec_TadA). The unique mammalian ADAT1 sequence is located within the deaminase domain (gray bar). ADARs target dsRNA. ADATs target tRNA, even though they lack any known RNA-binding motifs. Yeast _Sc_ADAT2 and _Sc_ADAT3 form active heterodimers, whereas ADAR1 and ADAR2 form active homodimers.
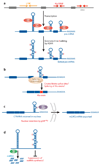
Figure 3
Possible regulatory functions for noncoding RNA editing. (a) Extensive A→I editing of an RNA duplex structure that consists of inverted Alu or LINE repeats. The inverted Alu or LINE repeats in introns and untranslated regions (UTRs) form intramolecular RNA duplexes genome wide, which are then subjected to A→I RNA editing by ADAR. (b) Splicing machinery interprets an inosine as a guanosine. Therefore, splice sites might be created or deleted by A→I editing of intronic Alu fold-back dsRNAs, leading to the inclusion or exclusion of Alu exons (102). (c) A→I editing of a short interspersed element (SINE) fold-back dsRNA present within the 3′ UTR of cationic amino acid transporter 2 (CAT2) nuclear (CTN) RNA. Its binding to p54nrb might be involved in the regulatory mechanism that retains this RNA within nuclear speckles (118). When cells are placed under stress, CTN-RNA is cleaved and de novo polyadenylated at an alternative site to release the protein-coding mCAT2 mRNA, which is then translated for CAT2 proteins (118). (d) A→I editing of Alu or LINE fold-back dsRNAs might suppress the generation of esiRNAs and consequent RNAi-mediated silencing of retrotransposon activities and/or genes (mRNAs) harboring these sequences within UTRs in trans.
Figure 4
Interaction between RNA editing and RNAi pathways. Two possible ways in which RNA editing and RNAi pathways may interact. (a) The introduction of many I·U base pairs and the alteration of the dsRNA structure by ADAR leads to the generation of fewer esiRNAs by Dicer, because dsRNAs that contain many I·U base pairs become resistant to Dicer cleavage. Alternatively, extensively edited dsRNAs with multiple I·U base pairs may be rapidly degraded by Tudor staphylococcal nuclease (Tudor-SN), resulting in the generation of fewer esiRNAs. (b) In addition, a fraction of already processed siRNAs might be sequestered by certain ADAR gene family members, reducing the effective siRNA concentration. For instance, cytoplasmic ADAR1p150 binds siRNA tightly. Gene silencing by siRNA is significantly more effective in the absence of ADAR1, indicating that ADAR1p150 is a cellular factor that limits siRNA potency in mammalian cells by decreasing the effective siRNA concentration and its incorporation into RNA-induced silencing complex.
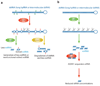
Figure 5
Regulation of miRNA processing and expression by RNA editing. (a) The Drosha/DGCR8 complex cleaves pri-miRNAs in the nucleus, producing ~70-nt pre-miRNA intermediates, which are exported by exportin-5 (Exp5) and RanGTP (Ran) into the cytoplasm. The Dicer/TRBP complex executes a second cleavage, generating mature miRNAs. (b) Drosha cleavage of pri- to pre-miRNA is suppressed by A→ editing of certain sites, such as the +4 and +5 sites of pri-miRNA-142 (miR-142). In addition, highly edited pri-miRNA-142 is degraded by Tudor staphylococcal nuclease (Tudor-SN) (138). (c) A→I editing of certain sites might lead to inhibition of the Dicer/TRBP cleavage step, as shown for pri-miR-151 editing. (d) Editing of pri-miRNAs at certain sites might lead to the expression of edited mature miRNAs and silencing of a different set of target genes owing to “seed” sequence alterations, as shown for miR-376.
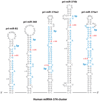
Figure 6
A→I RNA editing of pri-miR-376 RNAs. Hairpin structures of five human pri-miR-376 cluster member RNAs that are frequently edited in brain and other tissues. Regions processed into mature miRNAs are highlighted in blue. The two most highly edited adenosines (+4 and +44 sites) are highlighted in red. Editing sites that are numbered with the 5′ end of the human miR-376a1–5p sequence are counted as +1.
Similar articles
- ADAR editing in double-stranded UTRs and other noncoding RNA sequences.
Hundley HA, Bass BL. Hundley HA, et al. Trends Biochem Sci. 2010 Jul;35(7):377-83. doi: 10.1016/j.tibs.2010.02.008. Epub 2010 Apr 8. Trends Biochem Sci. 2010. PMID: 20382028 Free PMC article. Review. - ADAR-mediated RNA editing in non-coding RNA sequences.
Yang Y, Zhou X, Jin Y. Yang Y, et al. Sci China Life Sci. 2013 Oct;56(10):944-52. doi: 10.1007/s11427-013-4546-5. Epub 2013 Sep 5. Sci China Life Sci. 2013. PMID: 24008387 Review. - An I for an A: Dynamic Regulation of Adenosine Deamination-Mediated RNA Editing.
Vesely C, Jantsch MF. Vesely C, et al. Genes (Basel). 2021 Jul 1;12(7):1026. doi: 10.3390/genes12071026. Genes (Basel). 2021. PMID: 34356042 Free PMC article. Review. - Noncoding regions of C. elegans mRNA undergo selective adenosine to inosine deamination and contain a small number of editing sites per transcript.
Wheeler EC, Washburn MC, Major F, Rusch DB, Hundley HA. Wheeler EC, et al. RNA Biol. 2015;12(2):162-74. doi: 10.1080/15476286.2015.1017220. RNA Biol. 2015. PMID: 25826568 Free PMC article. - Editor meets silencer: crosstalk between RNA editing and RNA interference.
Nishikura K. Nishikura K. Nat Rev Mol Cell Biol. 2006 Dec;7(12):919-31. doi: 10.1038/nrm2061. Nat Rev Mol Cell Biol. 2006. PMID: 17139332 Free PMC article. Review.
Cited by
- Genome sequence-independent identification of RNA editing sites.
Zhang Q, Xiao X. Zhang Q, et al. Nat Methods. 2015 Apr;12(4):347-50. doi: 10.1038/nmeth.3314. Epub 2015 Mar 2. Nat Methods. 2015. PMID: 25730491 Free PMC article. - The role of RNA editing enzyme ADAR1 in human disease.
Song B, Shiromoto Y, Minakuchi M, Nishikura K. Song B, et al. Wiley Interdiscip Rev RNA. 2022 Jan;13(1):e1665. doi: 10.1002/wrna.1665. Epub 2021 Jun 8. Wiley Interdiscip Rev RNA. 2022. PMID: 34105255 Free PMC article. Review. - Transient overexpression of exogenous APOBEC3A causes C-to-U RNA editing of thousands of genes.
Sharma S, Patnaik SK, Kemer Z, Baysal BE. Sharma S, et al. RNA Biol. 2017 May 4;14(5):603-610. doi: 10.1080/15476286.2016.1184387. Epub 2016 May 5. RNA Biol. 2017. PMID: 27149507 Free PMC article. - Natural antisense transcripts in the biological hallmarks of cancer: powerful regulators hidden in the dark.
Zhao S, Zhang X, Chen S, Zhang S. Zhao S, et al. J Exp Clin Cancer Res. 2020 Sep 14;39(1):187. doi: 10.1186/s13046-020-01700-0. J Exp Clin Cancer Res. 2020. PMID: 32928281 Free PMC article. Review. - NERD-seq: a novel approach of Nanopore direct RNA sequencing that expands representation of non-coding RNAs.
Saville L, Wu L, Habtewold J, Cheng Y, Gollen B, Mitchell L, Stuart-Edwards M, Haight T, Mohajerani M, Zovoilis A. Saville L, et al. Genome Biol. 2024 Aug 28;25(1):233. doi: 10.1186/s13059-024-03375-8. Genome Biol. 2024. PMID: 39198865 Free PMC article.
References
- Baltimore D. Our genome unveiled. Nature. 2001;409:814–816. - PubMed
- Maniatis T, Reed R. An extensive network of coupling among gene expression machines. Nature. 2002;416:499–506. - PubMed
- Benne R, Van den Burg J, Brakenhoff JP, Sloof P, Van Boom JH, Tromp MC. Major transcript of the frameshifted coxII gene from trypanosome mitochondria contains four nucleotides that are not encoded in the DNA. Cell. 1986;46:819–826. - PubMed
- Gott JM, Emeson RB. Functions and mechanisms of RNA editing. Annu. Rev. Genet. 2000;34:499–531. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials