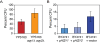Incipient balancing selection through adaptive loss of aquaporins in natural Saccharomyces cerevisiae populations - PubMed (original) (raw)
Incipient balancing selection through adaptive loss of aquaporins in natural Saccharomyces cerevisiae populations
Jessica L Will et al. PLoS Genet. 2010.
Abstract
A major goal in evolutionary biology is to understand how adaptive evolution has influenced natural variation, but identifying loci subject to positive selection has been a challenge. Here we present the adaptive loss of a pair of paralogous genes in specific Saccharomyces cerevisiae subpopulations. We mapped natural variation in freeze-thaw tolerance to two water transporters, AQY1 and AQY2, previously implicated in freeze-thaw survival. However, whereas freeze-thaw-tolerant strains harbor functional aquaporin genes, the set of sensitive strains lost aquaporin function at least 6 independent times. Several genomic signatures at AQY1 and/or AQY2 reveal low variation surrounding these loci within strains of the same haplotype, but high variation between strain groups. This is consistent with recent adaptive loss of aquaporins in subgroups of strains, leading to incipient balancing selection. We show that, although aquaporins are critical for surviving freeze-thaw stress, loss of both genes provides a major fitness advantage on high-sugar substrates common to many strains' natural niche. Strikingly, strains with non-functional alleles have also lost the ancestral requirement for aquaporins during spore formation. Thus, the antagonistic effect of aquaporin function-providing an advantage in freeze-thaw tolerance but a fitness defect for growth in high-sugar environments-contributes to the maintenance of both functional and nonfunctional alleles in S. cerevisiae. This work also shows that gene loss through multiple missense and nonsense mutations, hallmarks of pseudogenization presumed to emerge after loss of constraint, can arise through positive selection.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
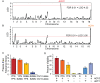
Figure 1. Yeast aquaporins underlie freeze-thaw tolerance in yeast.
(A) Genome scan of YPS163 x S288c identified a major QTL on the left arm of Chromosome 12. (B) A second scan in which the first QTL was treated as a fixed term identified a second QTL on the right arm of Chromosome 16. LOD scores at which FDR = 0.01 are indicated by horizontal red lines. Additional peaks in Figure 1B were not significant when both QTL on Chromosomes 12 and 16 were held fixed, suggesting they may be false positives. (C) Effect plot from QTL mapping. (D) Freeze-thaw tolerance was measured in parental strains, YPS163 (‘YPS’) mutants, and S288c-derivative BY4741 (‘S’) harboring empty vector or a plasmid-borne copy of AQY1 or AQY2 from YPS163, as described in Methods. The average and standard deviation of biological quadruplicates is shown.
Figure 2. Multiple independent losses of AQY2 and AQY1 in diverse S. cerevisiae isolates.
Bayesian trees of the recapitulated AQY proteins (where gaps were treated as missing data before translation) for Aqy2 (left) and Aqy1 (right). Trees were generated using Mr. Bayes 3.1 and a mixed amino acid replacement model with invgamma rates. All nodes displayed posterior probabilities >0.92. Strains with full-length proteins are nearly identical and do not resolve in the tree. Each star represents the appearance of a different deletion, including the Malaysian G528 (grey), 11-bp (yellow), and Asian G25 (blue) deletions in AQY2, and the amino-terminal Malaysian (green) and A881 (orange) deletions in AQY1. Appearance of the aqy1-V121M polymorphism is highlighted with a pink circle. Strains are color coded according to their niche as shown in the key. Freeze-thaw tolerance scores are listed to the right of strains shown in the Aqy1 tree (with +++ for tolerance and–for sensitivity, see Methods for details).
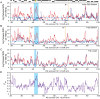
Figure 3. Skewed patterns of variation surrounding the AQY2 locus.
The average number of pairwise nucleotide differences per 1,000 bp sliding window of step size of 100 bp was plotted on Chromosome 12. Variation within (blue curve) and between (red curve) groups of (A) 4 strains harboring the Asian G25 AQY2 deletion, (B) 16 strains with the 11-bp deletion allele, and (C) 6 strains containing the full-length AQY2. Horizontal blue lines represent the genome-wide average of pair-wise variation within each group, and vertical lines represent manually defined breakpoints in trends. Regions identified with skewed between-group minus within-group variation (see Methods) are highlighted in blue. (D) A plot of FST based on the three groupings in (a–c); horizontal grey line represents the genome-wide average. Gene positions are shown above plots as black boxes, with AQY2 highlighted in yellow.
Figure 4. Strains lacking AQY genes show a fitness advantage under high osmolarity.
Cells were grown on solid agar plates containing 1.5 M sorbitol for 2 days and the number of colony-forming units (CFU) was compared to a no-stress control plate. (A) CFU for YPS163 and YPS163 aqy1Δ aqy2Δ grown on rich medium plus 1.5 M sorbitol; (B) S288c-derivative BY4741 harboring the empty vector or a plasmid expressing the YPS163 allele of AQY1 or AQY2, grown on selective medium with 1.5 M sorbitol. The average and standard deviation of biological triplicates is shown.
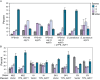
Figure 5. AQY function is required for sporulation in YPS163.
(A) Denoted strains were sporulated as described in Methods for 2 days, and the number of events with 4, 3, 2 spores or ‘other’ (representing unsporulated or unscorable cells) was recorded. Each plot represents the average and standard deviation of biological triplicate. Results from S. paradoxus strain NRRL Y-17217 are shown here. (B) Vineyard strain M22, sake strain K1, SK1, and S228c derivative BY4743 (BY) harboring empty vector or pYPS_ AQY1 plasmid were sporulated as described for 2–3 d before scoring.
Similar articles
- Functional relevance of water and glycerol channels in Saccharomyces cerevisiae.
Sabir F, Loureiro-Dias MC, Soveral G, Prista C. Sabir F, et al. FEMS Microbiol Lett. 2017 May 1;364(9). doi: 10.1093/femsle/fnx080. FEMS Microbiol Lett. 2017. PMID: 28430948 Review. - Aquaporin-mediated improvement of freeze tolerance of Saccharomyces cerevisiae is restricted to rapid freezing conditions.
Tanghe A, Van Dijck P, Colavizza D, Thevelein JM. Tanghe A, et al. Appl Environ Microbiol. 2004 Jun;70(6):3377-82. doi: 10.1128/AEM.70.6.3377-3382.2004. Appl Environ Microbiol. 2004. PMID: 15184134 Free PMC article. - Aquaporin expression correlates with freeze tolerance in baker's yeast, and overexpression improves freeze tolerance in industrial strains.
Tanghe A, Van Dijck P, Dumortier F, Teunissen A, Hohmann S, Thevelein JM. Tanghe A, et al. Appl Environ Microbiol. 2002 Dec;68(12):5981-9. doi: 10.1128/AEM.68.12.5981-5989.2002. Appl Environ Microbiol. 2002. PMID: 12450819 Free PMC article. - Polymorphism of Saccharomyces cerevisiae aquaporins.
Laizé V, Tacnet F, Ripoche P, Hohmann S. Laizé V, et al. Yeast. 2000 Jul;16(10):897-903. doi: 10.1002/1097-0061(200007)16:10<897::AID-YEA583>3.0.CO;2-T. Yeast. 2000. PMID: 10870101 - Water Transport in Yeasts.
Sabir F, Prista C, Madeira A, Moura T, Loureiro-Dias MC, Soveral G. Sabir F, et al. Adv Exp Med Biol. 2016;892:107-124. doi: 10.1007/978-3-319-25304-6_5. Adv Exp Med Biol. 2016. PMID: 26721272 Review.
Cited by
- Mixing of vineyard and oak-tree ecotypes of Saccharomyces cerevisiae in North American vineyards.
Hyma KE, Fay JC. Hyma KE, et al. Mol Ecol. 2013 Jun;22(11):2917-30. doi: 10.1111/mec.12155. Epub 2013 Jan 3. Mol Ecol. 2013. PMID: 23286354 Free PMC article. - A haplotype method detects diverse scenarios of local adaptation from genomic sequence variation.
Lange JD, Pool JE. Lange JD, et al. Mol Ecol. 2016 Jul;25(13):3081-100. doi: 10.1111/mec.13671. Epub 2016 Jun 6. Mol Ecol. 2016. PMID: 27135633 Free PMC article. - A tradeoff drives the evolution of reduced metal resistance in natural populations of yeast.
Chang SL, Leu JY. Chang SL, et al. PLoS Genet. 2011 Mar;7(3):e1002034. doi: 10.1371/journal.pgen.1002034. Epub 2011 Mar 31. PLoS Genet. 2011. PMID: 21483812 Free PMC article. - Wild Yeast for the Future: Exploring the Use of Wild Strains for Wine and Beer Fermentation.
Molinet J, Cubillos FA. Molinet J, et al. Front Genet. 2020 Nov 2;11:589350. doi: 10.3389/fgene.2020.589350. eCollection 2020. Front Genet. 2020. PMID: 33240332 Free PMC article. Review. - The population genomics of adaptive loss of function.
Monroe JG, McKay JK, Weigel D, Flood PJ. Monroe JG, et al. Heredity (Edinb). 2021 Mar;126(3):383-395. doi: 10.1038/s41437-021-00403-2. Epub 2021 Feb 11. Heredity (Edinb). 2021. PMID: 33574599 Free PMC article. Review.
References
- Mitchell-Olds T, Willis JH, Goldstein DB. Which evolutionary processes influence natural genetic variation for phenotypic traits? Nat Rev Genet. 2007;8:845–856. - PubMed
- Clark AG, Glanowski S, Nielsen R, Thomas PD, Kejariwal A, et al. Inferring nonneutral evolution from human-chimp-mouse orthologous gene trios. Science. 2003;302:1960–1963. - PubMed
- Nielsen R, Bustamante C, Clark AG, Glanowski S, Sackton TB, et al. A scan for positively selected genes in the genomes of humans and chimpanzees. PLoS Biol. 2005;3:e170. doi: 10.1371/journal.pbio.0030170. - DOI - PMC - PubMed
- Sabeti PC, Reich DE, Higgins JM, Levine HZ, Richter DJ, et al. Detecting recent positive selection in the human genome from haplotype structure. Nature. 2002;419:832–837. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases