Dnmt3a-dependent nonpromoter DNA methylation facilitates transcription of neurogenic genes - PubMed (original) (raw)
Dnmt3a-dependent nonpromoter DNA methylation facilitates transcription of neurogenic genes
Hao Wu et al. Science. 2010.
Abstract
DNA methylation at proximal promoters facilitates lineage restriction by silencing cell type-specific genes. However, euchromatic DNA methylation frequently occurs in regions outside promoters. The functions of such nonproximal promoter DNA methylation are unclear. Here we show that the de novo DNA methyltransferase Dnmt3a is expressed in postnatal neural stem cells (NSCs) and is required for neurogenesis. Genome-wide analysis of postnatal NSCs indicates that Dnmt3a occupies and methylates intergenic regions and gene bodies flanking proximal promoters of a large cohort of transcriptionally permissive genes, many of which encode regulators of neurogenesis. Surprisingly, Dnmt3a-dependent nonproximal promoter methylation promotes expression of these neurogenic genes by functionally antagonizing Polycomb repression. Thus, nonpromoter DNA methylation by Dnmt3a may be used for maintaining active chromatin states of genes critical for development.
Figures
Fig. 1
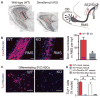
Essential roles of Dnmt3a in postnatal neurogenesis. (A) Cell nuclei staining of olfactory bulb (OB) in WT and KO mice at postnatal day (P) 24. The schematic on the right outlines the postnatal neurogenesis from SEZ/SVZ NSCs, and the red boxes denote the portion of the rostral migratory stream (RMS) with newborn neurons entering the OB. Note that both the OB size and the number of newborn neurons in KO mice (red box) are reduced. Arrows indicate radially migrating immature neurons in the postnatal OB. Scale bars, 200 μm. (B) Immunohistochemistry and quantification of Dcx+ neuroblasts in WT and KO RMS (P21 to P24). Hpf, high-power field; n, number of fields from three pairs of littermate mice. Error bars indicate SEM (*P < 0.01). Scale bars, 50 μm. (C) Immunocytochemistry of the immature neuronal marker Tuj1 in differentiating WT and KO SEZ/SVZ cultures. Scale bars, 50 μm. (D) Quantification of Tuj1+ neurons in differentiating WT, KO, and rescued NSC cultures. Error bars indicate SEM (*P < 0.01, n = 10 to 15).
Fig. 2
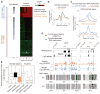
Dnmt3a occupies and methylates defined regions at both transcriptionally active and inactive genes. (A) Heat map representations of CpG islands (blue/white) and Dnmt3a (red/green) are shown for both Dnmt3a targets and nontargets. Red/blue represents enrichment, whereas green/white represents no enrichment. Genes were rank-ordered by CpG-island length within 4-kb regions flanking TSSs. FDR, false discovery rate; n, number of genes. (B) Distribution of Dnmt3a binding-sites within regions flanking TSSs of H3K4me3-high/CpG-rich and H3K4me3-low/CpG-poor targets. (C) Distribution of Dnmt3a-dependent DNA methylation within regions flanking TSSs of CpG-rich and CpG-poor genes. (D) Dnmt3a occupancy and Dnmt3a-dependent DNA methylation within representative CpG-rich/H3K4me3-high (Gbx2) and CpG-poor/ H3K4me3-low (Gfap) targets. Regions validated by bisulphite sequencing are shown in gray. IP, immunoprecipitation; WCE, whole-cell extract. (E) Box plots of expression levels in WT postnatal NSCs are shown for genes associated with Dnmt3a, H3K4me3, H3K27me3, and proximal promoter methylation (sperm-specific genes). Middle bars, medians; notches, standard errors; boxes, interquartile ranges; and whiskers, 10th and 90th percentiles.
Fig. 3
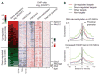
Dnmt3a promotes transcription through nonpromoter methylation. (A) Heat map representations of relative changes in expression (from four experiments), DNA methylation, H3K4me3, H3K27me3, and total histone H3 levels (panH3) between KO and WT NSCs are shown for differentially expressed Dnmt3a targets, which are ranked by changes in expression between KO and WT NSCs. n, number of targets. (B) Distributions of DNA demethylation and H3K27me3 increase in KO NSCs are shown for up-regulated targets (red), down-regulated targets (green), targets without expression changes (blue), and nontargets (gray).
Fig. 4
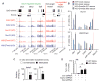
Dnmt3a promotes transcription by antagonizing Polycomb repression. (A) Ezh2, Suz12, and H3K27me3 occupancy at representative genes in WT and KO NSCs. Regions analyzed by ChIP-qPCR (in fig. S11E) are shaded in gray. (B) Quantitative PCR analysis of Suz12 and H3K27me3 at representative targets in WT, KO, and rescued NSCs. (C) Relative binding of PRC2 subunits on methylated or unmethylated chromatin arrays was quantified by densitometry (*P = 0.027, n = 4 experiments; **P = 0.0013, n = 6). (D) Quantification of Tuj1+ neurons in control and PRC2-deficient differentiating NSCs. Error bars indicate SEM of randomly selected fields from two independent cultures (*P < 0.05; **P < 0.01).
Similar articles
- Targeting of EZH2 to a defined genomic site is sufficient for recruitment of Dnmt3a but not de novo DNA methylation.
Rush M, Appanah R, Lee S, Lam LL, Goyal P, Lorincz MC. Rush M, et al. Epigenetics. 2009 Aug 16;4(6):404-14. doi: 10.4161/epi.4.6.9392. Epub 2009 Aug 29. Epigenetics. 2009. PMID: 19717977 - Dnmt3L antagonizes DNA methylation at bivalent promoters and favors DNA methylation at gene bodies in ESCs.
Neri F, Krepelova A, Incarnato D, Maldotti M, Parlato C, Galvagni F, Matarese F, Stunnenberg HG, Oliviero S. Neri F, et al. Cell. 2013 Sep 26;155(1):121-34. doi: 10.1016/j.cell.2013.08.056. Cell. 2013. PMID: 24074865 - Control of epigenetic states by WT1 via regulation of de novo DNA methyltransferase 3A.
Szemes M, Dallosso AR, Melegh Z, Curry T, Li Y, Rivers C, Uney J, Mägdefrau AS, Schwiderski K, Park JH, Brown KW, Shandilya J, Roberts SG, Malik K. Szemes M, et al. Hum Mol Genet. 2013 Jan 1;22(1):74-83. doi: 10.1093/hmg/dds403. Epub 2012 Oct 5. Hum Mol Genet. 2013. PMID: 23042785 Free PMC article. - The histone mark H3K36me2 recruits DNMT3A and shapes the intergenic DNA methylation landscape.
Weinberg DN, Papillon-Cavanagh S, Chen H, Yue Y, Chen X, Rajagopalan KN, Horth C, McGuire JT, Xu X, Nikbakht H, Lemiesz AE, Marchione DM, Marunde MR, Meiners MJ, Cheek MA, Keogh MC, Bareke E, Djedid A, Harutyunyan AS, Jabado N, Garcia BA, Li H, Allis CD, Majewski J, Lu C. Weinberg DN, et al. Nature. 2019 Sep;573(7773):281-286. doi: 10.1038/s41586-019-1534-3. Epub 2019 Sep 4. Nature. 2019. PMID: 31485078 Free PMC article. - Emerging Insights into the Distinctive Neuronal Methylome.
Clemens AW, Gabel HW. Clemens AW, et al. Trends Genet. 2020 Nov;36(11):816-832. doi: 10.1016/j.tig.2020.07.009. Epub 2020 Aug 21. Trends Genet. 2020. PMID: 32839016 Free PMC article. Review.
Cited by
- Imprecise DNMT1 activity coupled with neighbor-guided correction enables robust yet flexible epigenetic inheritance.
Wang Q, Yu G, Ming X, Xia W, Xu X, Zhang Y, Zhang W, Li Y, Huang C, Xie H, Zhu B, Xie W. Wang Q, et al. Nat Genet. 2020 Aug;52(8):828-839. doi: 10.1038/s41588-020-0661-y. Epub 2020 Jul 20. Nat Genet. 2020. PMID: 32690947 - Epigenetic regulation of stemness maintenance in the neurogenic niches.
Montalbán-Loro R, Domingo-Muelas A, Bizy A, Ferrón SR. Montalbán-Loro R, et al. World J Stem Cells. 2015 May 26;7(4):700-10. doi: 10.4252/wjsc.v7.i4.700. World J Stem Cells. 2015. PMID: 26029342 Free PMC article. Review. - The Adult Ventricular-Subventricular Zone (V-SVZ) and Olfactory Bulb (OB) Neurogenesis.
Lim DA, Alvarez-Buylla A. Lim DA, et al. Cold Spring Harb Perspect Biol. 2016 May 2;8(5):a018820. doi: 10.1101/cshperspect.a018820. Cold Spring Harb Perspect Biol. 2016. PMID: 27048191 Free PMC article. Review. - Epigenetic control of neurotransmitter expression in olfactory bulb interneurons.
Banerjee K, Akiba Y, Baker H, Cave JW. Banerjee K, et al. Int J Dev Neurosci. 2013 Oct;31(6):415-23. doi: 10.1016/j.ijdevneu.2012.11.009. Epub 2012 Dec 3. Int J Dev Neurosci. 2013. PMID: 23220178 Free PMC article. Review. - Mutations inDNMT3A and loss of RKIP are independent events in acute monocytic leukemia.
Fried I, Wölfler A, Quehenberger F, Hoefler G, Sill H, Zebisch A. Fried I, et al. Haematologica. 2012 Dec;97(12):1936-7. doi: 10.3324/haematol.2012.068429. Epub 2012 Aug 8. Haematologica. 2012. PMID: 22875620 Free PMC article. No abstract available.
References
- Jaenisch R, Bird A. Nat Genet. 2003;33:245. - PubMed
- Suzuki MM, Bird A. Nat Rev Genet. 2008;9:465. - PubMed
- Okano M, Bell DW, Haber DA, Li E. Cell. 1999;99:247. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases