Family-based association study of ITGB3 in autism spectrum disorder and its endophenotypes - PubMed (original) (raw)
doi: 10.1038/ejhg.2010.180. Epub 2010 Nov 24.
Federica Lombardi, Roberto Sacco, Paolo Curatolo, Barbara Manzi, Riccardo Alessandrelli, Roberto Militerni, Carmela Bravaccio, Carlo Lenti, Monica Saccani, Cindy Schneider, Raun Melmed, Tiziana Pascucci, Stefano Puglisi-Allegra, Karl-Ludvig Reichelt, Francis Rousseau, Patricia Lewin, Antonio M Persico
Affiliations
- PMID: 21102624
- PMCID: PMC3062005
- DOI: 10.1038/ejhg.2010.180
Family-based association study of ITGB3 in autism spectrum disorder and its endophenotypes
Valerio Napolioni et al. Eur J Hum Genet. 2011 Mar.
Abstract
The integrin-β 3 gene (ITGB3), located on human chromosome 17q21.3, was previously identified as a quantitative trait locus (QTL) for 5-HT blood levels and has been implicated as a candidate gene for autism spectrum disorder (ASD). We performed a family-based association study in 281 simplex and 12 multiplex Caucasian families. ITGB3 haplotypes are significantly associated with autism (HBAT, global P=0.038). Haplotype H3 is largely over-transmitted to the affected offspring and doubles the risk of an ASD diagnosis (HBAT P=0.005; odds ratio (OR)=2.000), at the expense of haplotype H1, which is under-transmitted (HBAT P=0.018; OR=0.725). These two common haplotypes differ only at rs12603582 located in intron 11, which reaches a P-value of 0.072 in single-marker FBAT analyses. Interestingly, rs12603582 is strongly associated with pre-term delivery in our ASD patients (P=0.008). On the other hand, it is SNP rs2317385, located at the 5' end of the gene, that significantly affects 5-HT blood levels (Mann-Whitney U-test, P=0.001; multiple regression analysis, P=0.010). No gene-gene interaction between ITGB3 and SLC6A4 has been detected. In conclusion, we identify a significant association between a common ITGB3 haplotype and ASD. Distinct markers, located toward the 5' and 3' ends of the gene, seemingly modulate 5-HT blood levels and autism liability, respectively. Our results also raise interest into ITGB3 influences on feto-maternal immune interactions in autism.
© 2011 Macmillan Publishers Limited All rights reserved 1018-4813/11
Figures
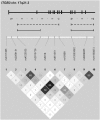
Figure 1
ITGB3 exon–intron structure, genotyped SNPs, and linkage disequilibrium expressed in _r_2. Haplotype blocks defined according to the confidence interval, four-gamete rule, and solid spine of LD algorithms are shown above by solid, broken, and dotted/broken lines, respectively.
References
- American Psychiatric Association American Psychiatric Association: Diagnostic and Statistical Manual of Mental Disorders4th edition.Publisher; 1994
- Persico AM, Bourgeron T. Searching for ways out of the autism maze: genetic, epigenetic and environmental clues. Trends Neurosci. 2006;29:349–358. -PubMed
- Klauck SM. Genetics of autism spectrum disorder. Eur J Hum Genet. 2006;14:714–720. -PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources