Carrier testing for severe childhood recessive diseases by next-generation sequencing - PubMed (original) (raw)
. 2011 Jan 12;3(65):65ra4.
doi: 10.1126/scitranslmed.3001756.
Darrell L Dinwiddie, Neil A Miller, Shannon L Hateley, Elena E Ganusova, Joann Mudge, Ray J Langley, Lu Zhang, Clarence C Lee, Faye D Schilkey, Vrunda Sheth, Jimmy E Woodward, Heather E Peckham, Gary P Schroth, Ryan W Kim, Stephen F Kingsmore
Affiliations
- PMID: 21228398
- PMCID: PMC3740116
- DOI: 10.1126/scitranslmed.3001756
Carrier testing for severe childhood recessive diseases by next-generation sequencing
Callum J Bell et al. Sci Transl Med. 2011.
Abstract
Of 7028 disorders with suspected Mendelian inheritance, 1139 are recessive and have an established molecular basis. Although individually uncommon, Mendelian diseases collectively account for ~20% of infant mortality and ~10% of pediatric hospitalizations. Preconception screening, together with genetic counseling of carriers, has resulted in remarkable declines in the incidence of several severe recessive diseases including Tay-Sachs disease and cystic fibrosis. However, extension of preconception screening to most severe disease genes has hitherto been impractical. Here, we report a preconception carrier screen for 448 severe recessive childhood diseases. Rather than costly, complete sequencing of the human genome, 7717 regions from 437 target genes were enriched by hybrid capture or microdroplet polymerase chain reaction, sequenced by next-generation sequencing (NGS) to a depth of up to 2.7 gigabases, and assessed with stringent bioinformatic filters. At a resultant 160x average target coverage, 93% of nucleotides had at least 20x coverage, and mutation detection/genotyping had ~95% sensitivity and ~100% specificity for substitution, insertion/deletion, splicing, and gross deletion mutations and single-nucleotide polymorphisms. In 104 unrelated DNA samples, the average genomic carrier burden for severe pediatric recessive mutations was 2.8 and ranged from 0 to 7. The distribution of mutations among sequenced samples appeared random. Twenty-seven percent of mutations cited in the literature were found to be common polymorphisms or misannotated, underscoring the need for better mutation databases as part of a comprehensive carrier testing strategy. Given the magnitude of carrier burden and the lower cost of testing compared to treating these conditions, carrier screening by NGS made available to the general population may be an economical way to reduce the incidence of and ameliorate suffering associated with severe recessive childhood disorders.
Conflict of interest statement
Competing interests: L.Z. is an employee of Illumina Inc. At the time the research was performed, G.P.S. was an employee of Illumina Inc. C.C.L., H.E.P., and V.S. are employees of Life Technologies. U.S. patent application 20090183268 entitled “Methods and systems for medical sequencing analysis” was filed by the National Center for Genome Resources on July 16, 2009. This application has claims related to this work. The other authors declare no competing interests.
Figures
Fig. 1
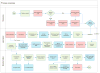
Workflow of the comprehensive carrier screening test. Workflow shows receiving samples and DNA extraction, target enrichment from DNA samples, multiplexed sequencing library preparation, NGS, and bioinformatic analysis. (The bioinformatic decision tree is shown in fig. S4.)
Fig. 2
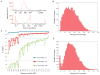
Analytic metrics of multiplexed carrier testing by NGS. (A) Chromatograms of size distributions of sequencing libraries after target enrichment. Top: Target enrichment by hybrid capture. Bottom: Target enrichment by microdroplet PCR. Size markers are shown at 40 and 8000 nt. FU, fluorescence units. (B) Frequency distribution of target coverage after hybrid selection and 1.75 Gb of singleton 50-mer Illumina GAIIx SBS of sample NA13675. Aligned sequences had a quality score of >25. (C) Target coverage as a function of depth of sequencing across 104 samples and six experiments. (D) Frequency distribution of target coverage after microdroplet PCR and 1.49 Gb of singleton 50-mer SBS of sample NA20379. Aligned sequences had a quality score of >25.
Fig. 3
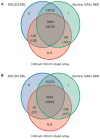
Venn diagrams of specificity of on-target SNP calls and genotypes in six samples. Target nucleotides were enriched by hybrid selection and sequenced by Illumina GAIIx SBS and SOLiD3 SBL at sixfold multiplexing. The samples were also genotyped with Infinium Omni1-Quad SNP arrays. (A) Comparison of SNP calls and genotypes obtained by SBS, SBL, and arrays at nucleotides surveyed by all three methods. SNPs were called if present in >10 uniquely aligning SBS reads, >14% of reads, and with average quality score of >20. Heterozygotes were identified if present in 14 to 86% of reads. Numbers refer to SNP calls. Numbers in brackets refer to SNP genotypes. (B) Comparison of SNP calls and genotypes obtained by SBS, SBL, and arrays. SNPs were called if present in more than four uniquely aligning SBS reads, >14% of reads, and with average quality score of >20. Heterozygotes were identified if present in 14 to 86% of reads.
Fig. 4
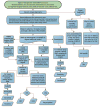
Decision tree to classify sequence variation and evaluate carrier status. After reads were aligned to references, substitution, insertion, and deletion events and their associated quality metrics were recorded. Variants were classified as heterozygous or homozygous and annotated by comparison with mutation databases. Variants not in the mutation databases were evaluated for putative functional consequence and were retained as disease mutations if predicted to result in protein truncation. Variants with a frequency of <5% among all samples and that were known to cause a disease phenotype or loss of protein function and that were only found as homozygous in affected individuals were retained and reported.
Fig. 5
Detection of gross deletion mutations by local reduction in normalized aligned reads. (A) Deletion of CLN3 introns 6 to 8, 966bpdel, exons7-8del and fs, chr16:28405752_28404787del in four known compound heterozygotes (NA20381, NA20382, NA20383, and NA20384; red diamonds) and one undescribed carrier (NA00006; green diamond) among 72 samples sequenced. (B) Heterozygous deletion in HBA1 (chr16:141620_172294del, 30,676-bp deletion from 5′ of ζ2 to 3′ of θ1 in ALU regions) in one known (NA10798; red diamond; normalized coverage, 26; mean normalized coverage, 61.9 ± 15.2) and two undescribed carriers [NA19193 (normalized coverage, 28) and NA01982 (normalized coverage, 31); green diamonds] among 72 samples. Heterozygous deletion in NA10798 was confirmed by array hybridization. (C) Known homozygous deletion of exons 7 and 8 of SMN1 in one of eight samples (NA03813; red diamond). (D) Detection of a gross deletion that is a cause of Duchenne muscular dystrophy (OMIM #310200, DMD exons 51 to 55 del, chrX:31702000_31555711del) by reduction in normalized aligned reads at chrX:31586112. Among 72 samples, one (NA04364; red diamond) was from an affected male, and another (NA18540, a female JPT/HAN HapMap sample) was determined to carry a deletion that extends to at least chrX:31860199 [see (E)]. (E) An undescribed heterozygous deletion of DMD 3′ exon 44–3′ exon 50 (chrX:32144956-31702228del) in NA18540 (green diamond), a JPT/HAN HapMap sample. This deletion extends from at least chrX:31586112 to chrX:31860199 [see (D)]. Sample NA05022 (red diamond) is the uncharacterized mother of an affected son with 3′ exon 44–3′ exon 50 del, chrX:32144956-31702228del. Given the absence of the mutation in the mother, it likely occurred de novo in the son, as observed in one-third of DMD patients (62). (F) Hemizygous deletion in PLP1 exons3_4, c.del349_495del, chrX:102928207_102929424del in one (NA13434; red diamond) of eight samples. (G) Absence of gross deletion CG984340 (ERCC6 exon 9, c.1993_2169del, 665_723del, exon 9 del, chr10:50360915_50360739del) in 72 DNA samples. The sample in red (NA01712) was incorrectly annotated to be a compound heterozygote with CG984340 on the basis of cDNA sequencing.
Fig. 6
Clinical metrics of multiplexed carrier testing by NGS. (A) Comparison of 92,128 SNP genotypes by array hybridization with those obtained by target enrichment, SBS, and a bioinformatic decision tree in 26 samples. SNPs were called if present in >10 uniquely aligning reads, >14% of reads, and average quality score of >20. Heterozygotes were identified if present in 14 to 86% of reads. TP = SNP called and genotyped correctly. TN = reference genotype called correctly. FN = SNP genotype undercall. FP = SNP genotype overcall. Accuracy = (TP + TN)/(TP + FN + TN + FP). Sensitivity = TP/(TP + FN). Specificity = TN/(TN + FP). Positive predictive value (PPV) = TP/(TP + FP). Negative predictive value (NPV) = TN/(TN + FN). (B) Distribution of allele frequencies of SNP calls by hybrid capture and SBS in 26 samples. Light blue, heterozygotes by array hybridization. (C) Receiver operating characteristic (ROC) curve of sensitivity and specificity of SNP genotypes by hybrid capture and SBS in 26 samples (when compared with array-based genotypes). Genomic regions with less than 20× coverage were excluded. Upon varying the number of reads calling the SNP, the area under the curve (AUC) was 0.97. (D) ROC curve of SNP genotypes by hybrid capture and SBS in 26 samples. Genomic regions with less than 20× coverage were excluded. Upon varying the percent reads calling the SNP, AUC was 0.97.
Fig. 7
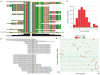
Disease mutations and estimated carrier burden in 104 DNA samples. (A) Sample NA07092, from an affected male with X-linked recessive Lesch-Nyhan syndrome (OMIM #300322), had been characterized as a deletion of HPRT1 exon 8 by cDNA sequencing (19), but has an explanatory splicing mutation (intron 8, IVS8+1_4delGTAA, chrX:133460381_133460384delGTAA). (B) Sample NA09545, from an affected male with X-linked recessive Pelizaeus-Merzbacher disease (PMD; OMIM #312080), characterized as a substitution disease mutation [PLP1 exon 5, c.767C>T, P215S (20)], also featured PLP1 gene duplication [which is reported in 62% of sporadic PMD (21)]. (C) Distribution of carrier burden of severe pediatric diseases among 104 DNA samples. (D) Ward hierarchical clustering of 227 severe pediatric disease mutations in 104 DNA samples.
Fig. 8
Five reads from NA202057 showing AGA exon 4, c.488G>C, C163S, chr4:178596912G>C and exon 4, c.482G>A, R161Q, chr4:178596918G>A (black arrows). One hundred and ninety-three of 400 reads contained these substitution disease mutations (CM910010 and CM910011). The top lines of doublets are Illumina GAIIx 50-nt reads. The bottom lines are NCBI reference genome, build 36.3. Colors represent quality (Q) scores of each nucleotide: red, >30; orange, 20 to 29; green, 10 to 19. Reads aligned uniquely to these coordinates.
Comment in
- Molecular technologies open new clinical genetic vistas.
Jackson L, Pyeritz RE. Jackson L, et al. Sci Transl Med. 2011 Jan 12;3(65):65ps2. doi: 10.1126/scitranslmed.3002064. Sci Transl Med. 2011. PMID: 21228396 - Sequencing: the next generation. Moving beyond population-based recessive disease carrier screening.
Kobelka CE. Kobelka CE. Clin Genet. 2011 Jul;80(1):25-6. doi: 10.1111/j.1399-0004.2011.01677.x. Epub 2011 Apr 25. Clin Genet. 2011. PMID: 21476994
Similar articles
- Comprehensive carrier screening and molecular diagnostic testing for recessive childhood diseases.
Kingsmore S. Kingsmore S. PLoS Curr. 2012 May 2;4:e4f9877ab8ffa9. doi: 10.1371/4f9877ab8ffa9. PLoS Curr. 2012. PMID: 22872815 Free PMC article. - Screening for Tay-Sachs disease carriers by full-exon sequencing with novel variant interpretation outperforms enzyme testing in a pan-ethnic cohort.
Cecchi AC, Vengoechea ES, Kaseniit KE, Hardy MW, Kiger LA, Mehta N, Haque IS, Moyer K, Page PZ, Muzzey D, Grinzaid KA. Cecchi AC, et al. Mol Genet Genomic Med. 2019 Aug;7(8):e836. doi: 10.1002/mgg3.836. Epub 2019 Jul 10. Mol Genet Genomic Med. 2019. PMID: 31293106 Free PMC article. - Comprehensive carrier genetic test using next-generation deoxyribonucleic acid sequencing in infertile couples wishing to conceive through assisted reproductive technology.
Martin J, Asan, Yi Y, Alberola T, Rodríguez-Iglesias B, Jiménez-Almazán J, Li Q, Du H, Alama P, Ruiz A, Bosch E, Garrido N, Simon C. Martin J, et al. Fertil Steril. 2015 Nov;104(5):1286-93. doi: 10.1016/j.fertnstert.2015.07.1166. Epub 2015 Sep 3. Fertil Steril. 2015. PMID: 26354092 - Carrier testing for autosomal-recessive disorders.
Vallance H, Ford J. Vallance H, et al. Crit Rev Clin Lab Sci. 2003 Aug;40(4):473-97. doi: 10.1080/10408360390247832. Crit Rev Clin Lab Sci. 2003. PMID: 14582604 Review. - Variables that underlie cost efficacy of prenatal screening.
Kinzler WL, Morrell K, Vintzileos AM. Kinzler WL, et al. Obstet Gynecol Clin North Am. 2002 Jun;29(2):277-86, v-vi. doi: 10.1016/s0889-8545(01)00007-9. Obstet Gynecol Clin North Am. 2002. PMID: 12108828 Review.
Cited by
- Alström Syndrome: Mutation Spectrum of ALMS1.
Marshall JD, Muller J, Collin GB, Milan G, Kingsmore SF, Dinwiddie D, Farrow EG, Miller NA, Favaretto F, Maffei P, Dollfus H, Vettor R, Naggert JK. Marshall JD, et al. Hum Mutat. 2015 Jul;36(7):660-8. doi: 10.1002/humu.22796. Epub 2015 May 18. Hum Mutat. 2015. PMID: 25846608 Free PMC article. - Challenges in the care for consanguineous couples: an exploratory interview study among general practitioners and midwives.
Teeuw ME, Hagelaar A, ten Kate LP, Cornel MC, Henneman L. Teeuw ME, et al. BMC Fam Pract. 2012 Oct 26;13:105. doi: 10.1186/1471-2296-13-105. BMC Fam Pract. 2012. PMID: 23102514 Free PMC article. - Maps for the world of genomic medicine: the 2011 CSHL Personal Genomes meeting.
Zheng-Bradley X, Flicek P. Zheng-Bradley X, et al. Hum Mutat. 2012 Jun;33(6):1016-9. doi: 10.1002/humu.22024. Epub 2012 Apr 16. Hum Mutat. 2012. PMID: 22253119 Free PMC article. - Implementing comprehensive genetic carrier screening in China-Harnessing the power of genomic medicine for the effective prevention/management of birth defects and rare genetic diseases in China.
Shen Y, Qiu X, Gui B, He S, Huang H, Xue J, Xu X, Zhang X, He L. Shen Y, et al. Pediatr Investig. 2018 May 11;2(1):30-36. doi: 10.1002/ped4.12021. eCollection 2018 Mar. Pediatr Investig. 2018. PMID: 32851227 Free PMC article. Review. - The landscape of autosomal-recessive pathogenic variants in European populations reveals phenotype-specific effects.
Fridman H, Yntema HG, Mägi R, Andreson R, Metspalu A, Mezzavila M, Tyler-Smith C, Xue Y, Carmi S, Levy-Lahad E, Gilissen C, Brunner HG. Fridman H, et al. Am J Hum Genet. 2021 Apr 1;108(4):608-619. doi: 10.1016/j.ajhg.2021.03.004. Epub 2021 Mar 18. Am J Hum Genet. 2021. PMID: 33740458 Free PMC article.
References
- Kaback MM. Hexosaminidase A deficiency. In: Pagon RA, Bird TC, Dolan CR, Stephens K, editors. GeneReviews. University of Washington; Seattle: 1993.
- Kronn D, Jansen V, Ostrer H. Carrier screening for cystic fibrosis, Gaucher disease, and Tay-Sachs disease in the Ashkenazi Jewish population: The first 1000 cases at New York University Medical Center, New York NY. Arch Intern Med. 1998;158:777–781. - PubMed
- Kaback MM. Population-based genetic screening for reproductive counseling: The Tay-Sachs disease model. Eur J Pediatr. 2000;159:S192–S195. - PubMed
Publication types
MeSH terms
Grants and funding
- U19 HD077693/HD/NICHD NIH HHS/United States
- U01 AI066569-05S1/AI/NIAID NIH HHS/United States
- P20 RR016480/RR/NCRR NIH HHS/United States
- U01 AI066569/AI/NIAID NIH HHS/United States
- RR016480/RR/NCRR NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical