SUMO-targeted ubiquitin ligase, Rad60, and Nse2 SUMO ligase suppress spontaneous Top1-mediated DNA damage and genome instability - PubMed (original) (raw)
SUMO-targeted ubiquitin ligase, Rad60, and Nse2 SUMO ligase suppress spontaneous Top1-mediated DNA damage and genome instability
Johanna Heideker et al. PLoS Genet. 2011 Mar.
Abstract
Through as yet undefined proteins and pathways, the SUMO-targeted ubiquitin ligase (STUbL) suppresses genomic instability by ubiquitinating SUMO conjugated proteins and driving their proteasomal destruction. Here, we identify a critical function for fission yeast STUbL in suppressing spontaneous and chemically induced topoisomerase I (Top1)-mediated DNA damage. Strikingly, cells with reduced STUbL activity are dependent on tyrosyl-DNA phosphodiesterase 1 (Tdp1). This is notable, as cells lacking Tdp1 are largely aphenotypic in the vegetative cell cycle due to the existence of alternative pathways for the removal of covalent Top1-DNA adducts (Top1cc). We further identify Rad60, a SUMO mimetic and STUbL-interacting protein, and the SUMO E3 ligase Nse2 as critical Top1cc repair factors in cells lacking Tdp1. Detection of Top1ccs using chromatin immunoprecipitation and quantitative PCR shows that they are elevated in cells lacking Tdp1 and STUbL, Rad60, or Nse2 SUMO ligase activity. These unrepaired Top1ccs are shown to cause DNA damage, hyper-recombination, and checkpoint-mediated cell cycle arrest. We further determine that Tdp1 and the nucleotide excision repair endonuclease Rad16-Swi10 initiate the major Top1cc repair pathways of fission yeast. Tdp1-based repair is the predominant activity outside S phase, likely acting on transcription-coupled Top1cc. Epistasis analyses suggest that STUbL, Rad60, and Nse2 facilitate the Rad16-Swi10 pathway, parallel to Tdp1. Collectively, these results reveal a unified role for STUbL, Rad60, and Nse2 in protecting genome stability against spontaneous Top1-mediated DNA damage.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
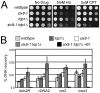
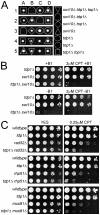
Figure 1. STUbL mutant tdp1 Δ cells are synergistically sensitive to CPT and have increased spontaneous Top1cc levels.
(A) Serial dilutions of the indicated strains were spotted onto media with or without the indicated drugs. (B) ChIP-qPCR assays of an _nmt41_-inducible Top1-FLAG in the indicated strains, at the subtelomeres of Chr 2 (telo2R), the centromeric inner repeats of Chr 2 (cnt2), the rDNA (rDNA2), and upstream of mes1 on Chr 1. The data represents the average DNA recovery compared to the input DNA samples with standard deviations from four independent experiments. ChIP-qPCR data of _nmt41_-Top1-FLAG slx8-1 tdp1Δ cells grown in repressing media (+B1) acts as a negative control. Cells were grown at 25°C.
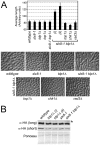
Figure 2. STUbL mutant tdp1 Δ cells activate the DNA damage checkpoint in a Top1-dependent manner.
(A) Upper: graph depicting the average cell length of the indicated strains grown in liquid media at 25°C. The error bars represent 95% confidence intervals between three independent experiments. Lower: representative images of the indicated genotypes are shown. (B) Western blot analysis of whole cell lysates from the indicated strains expressing HA-tagged Chk1 from the endogenous locus (long and short exposures are shown). The uppermost bands (asterisk) in the top panel are phosphorylated Chk1. Ponceau is shown as a loading control.
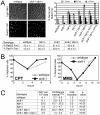
Figure 3. STUbL mutant tdp1 Δ cells exhibit elevated Top1-dependent DNA damage and spontaneous genomic instability.
(A) Upper left: live cell images of the indicated strains expressing Rad22-YFP (Rad52). Upper right: graph depicting the percentage of nuclei containing one, two, or multiple Rad22-YFP foci using live-cell microscopy in the indicated genotypes. Error bars represent the standard deviations from three independent experiments. Base: Table depicting the percentage of nuclei containing one or more Rad22-YFP or Rad11-YFP (RPA) foci in the indicated genotypes. Standard deviations are derived from three independent experiments. (B) Log phase cultures of slx8-1 and wildtype cells were treated with 40 µM CPT or 0.008% MMS. G2 checkpoint arrest and recovery was monitored through determining the percentage of septated cells at the indicated times. (C) Table depicting frequency of spontaneous mitotic recombination between tandem adenine heteroalleles in the indicated strains. Rates represent the mean of the means, between at least three independent assays per strain. All experiments were incubated at 25°C.
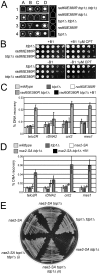
Figure 4. The Rad60∶Ubc9 complex and the Nse2 SUMO E3 ligase are essential to protect tdp1 Δ cells from Top1-induced DNA damage.
(A) A representative tetrad dissection is shown from a cross between rad60E380R and top1Δ tdp1Δ double mutant cells. The key depicts the genotypes present, which are denoted by various shapes placed around each colony. Wildtype cells do not have a shape placed around them. (B) Serial dilutions of the indicated strains expressing Top1 under a thiamine repressible promoter were spotted onto control or CPT containing media with (+B1) or without (-B1) thiamine to repress or induce Top1 expression, respectively. All strains were incubated at 32°C. (C) ChIP-qPCR assays of an _nmt41-_inducible Top1-FLAG in the indicated strains at the subtelomeres of Chr 2 (telo2R), the centromeric inner repeats of Chr 2 (cnt2), the rDNA (rDNA2), and upstream of mes1 on Chr 1. The data represents the average DNA recovery compared to the input DNA samples with standard deviations from at least three independent experiments. ChIP-qPCR data of _nmt41-_Top1-FLAG rad60E380R tdp1Δ grown in repressed media (+B1) is shown as a negative control. Cells were grown at 25°C. (D) ChIP qPCR assays of the indicated strains as in (C). (E) Cells of the indicated genotype were restruck directly from tetrad dissection plates onto YES media. Cells were incubated at 32°C.
Figure 5. Tdp1-deficient cells depend on Swi10, but not on homologous recombination repair factors, to prevent Top1-induced cell death.
(A) A representative tetrad dissection is shown of a cross between swi10Δ and the top1Δ tdp1Δ double mutant. The key depicts the genotypes present, which are denoted by various shapes placed around each colony. Wildtype cells do not have a shape placed around them. (B) Serial dilutions of the indicated strains expressing Top1 under a thiamine repressible promoter were spotted onto control or CPT containing media with (+B1) or without (-B1) thiamine to repress or induce Top1 expression, respectively. (C) Five fold serial dilutions of the indicated strains were spotted onto rich media that contained either no drug (YES), or camptothecin (CPT). Plates were incubated at 32°C.
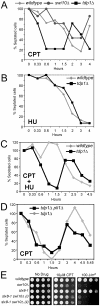
Figure 6. STUbL facilitates Top1cc repair in the Rad16-Swi10 pathway.
(A) Log phase cultures of the indicated strains were treated with 40 µM CPT and checkpoint arrest and recovery was monitored through determining the percentage of septated cells at the indicated times. (B) As for (A), except cultures were treated with 15 mM HU. (C) As for (B), except cultures were co-treated with 15 mM HU and 40 µM CPT. (D) The indicated strains were treated with CPT and checkpoint arrest and recovery were monitored as in (A-C). In all of these graphs, the asynchronous septation index was set at 100% for that observed at time zero in each strain studied. (E) The indicated strains were serially diluted and spotted on drug-free or CPT rich media, or were UV-irradiated at the indicated dose, and incubated at 25°C.
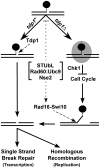
Figure 7. Model depicting the parallel actions of Tdp1 and Rad16-Swi10 in Top1cc removal, potentially facilitated by STUbL, Rad60∶Ubc9, and Nse2.
In wild-type cells, Tdp1 efficiently removes Top1cc (black circle), leaving a substrate for single strand break repair (SSBR) or homologous recombination (HR). In the absence of Tdp1, Top1cc is converted into a checkpoint visible lesion (shaded oval) that arrests cell cycle progression in a Chk1-dependent manner. Top1cc can ultimately be removed by Rad16-Swi10, a process that may be facilitated by STUbL, Rad60∶Ubc9 and Nse2. SSBR or HR can then heal the resulting lesion.
Similar articles
- SUMO-targeted ubiquitin ligase activity can either suppress or promote genome instability, depending on the nature of the DNA lesion.
Nie M, Moser BA, Nakamura TM, Boddy MN. Nie M, et al. PLoS Genet. 2017 May 5;13(5):e1006776. doi: 10.1371/journal.pgen.1006776. eCollection 2017 May. PLoS Genet. 2017. PMID: 28475613 Free PMC article. - Tdp1 processes chromate-induced single-strand DNA breaks that collapse replication forks.
Ganguly A, Guo L, Sun L, Suo F, Du LL, Russell P. Ganguly A, et al. PLoS Genet. 2018 Aug 27;14(8):e1007595. doi: 10.1371/journal.pgen.1007595. eCollection 2018 Aug. PLoS Genet. 2018. PMID: 30148840 Free PMC article. - SUMO orchestrates multiple alternative DNA-protein crosslink repair pathways.
Serbyn N, Bagdiul I, Noireterre A, Michel AH, Suhandynata RT, Zhou H, Kornmann B, Stutz F. Serbyn N, et al. Cell Rep. 2021 Nov 23;37(8):110034. doi: 10.1016/j.celrep.2021.110034. Cell Rep. 2021. PMID: 34818558 Free PMC article. - Cooperativity of the SUMO and Ubiquitin Pathways in Genome Stability.
Nie M, Boddy MN. Nie M, et al. Biomolecules. 2016 Feb 25;6(1):14. doi: 10.3390/biom6010014. Biomolecules. 2016. PMID: 26927199 Free PMC article. Review. - Tyrosyl-DNA-phosphodiesterases (TDP1 and TDP2).
Pommier Y, Huang SY, Gao R, Das BB, Murai J, Marchand C. Pommier Y, et al. DNA Repair (Amst). 2014 Jul;19:114-29. doi: 10.1016/j.dnarep.2014.03.020. Epub 2014 May 22. DNA Repair (Amst). 2014. PMID: 24856239 Free PMC article. Review.
Cited by
- Dual recruitment of Cdc48 (p97)-Ufd1-Npl4 ubiquitin-selective segregase by small ubiquitin-like modifier protein (SUMO) and ubiquitin in SUMO-targeted ubiquitin ligase-mediated genome stability functions.
Nie M, Aslanian A, Prudden J, Heideker J, Vashisht AA, Wohlschlegel JA, Yates JR 3rd, Boddy MN. Nie M, et al. J Biol Chem. 2012 Aug 24;287(35):29610-9. doi: 10.1074/jbc.M112.379768. Epub 2012 Jun 22. J Biol Chem. 2012. PMID: 22730331 Free PMC article. - The yeast homologue of the microtubule-associated protein Lis1 interacts with the sumoylation machinery and a SUMO-targeted ubiquitin ligase.
Alonso A, D'Silva S, Rahman M, Meluh PB, Keeling J, Meednu N, Hoops HJ, Miller RK. Alonso A, et al. Mol Biol Cell. 2012 Dec;23(23):4552-66. doi: 10.1091/mbc.E12-03-0195. Epub 2012 Oct 3. Mol Biol Cell. 2012. PMID: 23034179 Free PMC article. - Implementation and performance of SIBYLS: a dual endstation small-angle X-ray scattering and macromolecular crystallography beamline at the Advanced Light Source.
Classen S, Hura GL, Holton JM, Rambo RP, Rodic I, McGuire PJ, Dyer K, Hammel M, Meigs G, Frankel KA, Tainer JA. Classen S, et al. J Appl Crystallogr. 2013 Feb 1;46(Pt 1):1-13. doi: 10.1107/S0021889812048698. Epub 2013 Jan 17. J Appl Crystallogr. 2013. PMID: 23396808 Free PMC article. - STUbLs in chromatin and genome stability.
Garza R, Pillus L. Garza R, et al. Biopolymers. 2013 Feb;99(2):146-54. doi: 10.1002/bip.22125. Biopolymers. 2013. PMID: 23175389 Free PMC article. Review. - Ctp1-dependent clipping and resection of DNA double-strand breaks by Mre11 endonuclease complex are not genetically separable.
Jensen KL, Russell P. Jensen KL, et al. Nucleic Acids Res. 2016 Sep 30;44(17):8241-9. doi: 10.1093/nar/gkw557. Epub 2016 Jun 20. Nucleic Acids Res. 2016. PMID: 27325741 Free PMC article.
References
- Schleker T, Nagai S, Gasser SM. Posttranslational modifications of repair factors and histones in the cellular response to stalled replication forks. DNA Repair (Amst) 2009;8:1089–1100. - PubMed
- Ulrich HD. Mutual interactions between the SUMO and ubiquitin systems: a plea of no contest. Trends Cell Biol. 2005;15:525–532. - PubMed
- Kerscher O, Felberbaum R, Hochstrasser M. Modification of proteins by ubiquitin and ubiquitin-like proteins. Annu Rev Cell Dev Biol. 2006;22:159–180. - PubMed
- Watts FZ, Skilton A, Ho JC, Boyd LK, Trickey MA, et al. The role of Schizosaccharomyces pombe SUMO ligases in genome stability. Biochem Soc Trans. 2007;35:1379–1384. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials