Normal and neoplastic nonstem cells can spontaneously convert to a stem-like state - PubMed (original) (raw)
. 2011 May 10;108(19):7950-5.
doi: 10.1073/pnas.1102454108. Epub 2011 Apr 15.
Ines Brueckmann, Christina Scheel, Alicia J Kaestli, Paul A Wiggins, Leonardo O Rodrigues, Mary Brooks, Ferenc Reinhardt, Ying Su, Kornelia Polyak, Lisa M Arendt, Charlotte Kuperwasser, Brian Bierie, Robert A Weinberg
Affiliations
- PMID: 21498687
- PMCID: PMC3093533
- DOI: 10.1073/pnas.1102454108
Normal and neoplastic nonstem cells can spontaneously convert to a stem-like state
Christine L Chaffer et al. Proc Natl Acad Sci U S A. 2011.
Abstract
Current models of stem cell biology assume that normal and neoplastic stem cells reside at the apices of hierarchies and differentiate into nonstem progeny in a unidirectional manner. Here we identify a subpopulation of basal-like human mammary epithelial cells that departs from that assumption, spontaneously dedifferentiating into stem-like cells. Moreover, oncogenic transformation enhances the spontaneous conversion, so that nonstem cancer cells give rise to cancer stem cell (CSC)-like cells in vitro and in vivo. We further show that the differentiation state of normal cells-of-origin is a strong determinant of posttransformation behavior. These findings demonstrate that normal and CSC-like cells can arise de novo from more differentiated cell types and that hierarchical models of mammary stem cell biology should encompass bidirectional interconversions between stem and nonstem compartments. The observed plasticity may allow derivation of patient-specific adult stem cells without genetic manipulation and holds important implications for therapeutic strategies to eradicate cancer.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
Fig. 1.
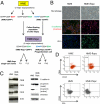
Enrichment of a rare population of floating cells from cultured mammary epithelial cells. (A) Schematic illustrating the derivation of HME-flopcs and single-cell clones (SCCs), as well as the antigen profiles of the major cell subpopulations described in this paper. (B) Phase contrast micrographs (20×) of HME and HME-flopc populations in 2D culture. Immunofluorescence of HME and HME-flopc cells is shown, for E-cadherin (green), β-catenin (red), and ZO-1 [nuclei stained blue with 4,6-diamidino-2-phenylindole (DAPI)]. (C) Immunoblot with antibodies to cytokeratins 14, 18 and 19, vimentin, E-cadherin and β-catenin (total and active forms). (D) Flow cytometry plots for CD44, CD24, and ESA on bulk HME and HME-flopc cells. Gating is set to unstained control cells.
Fig. 2.
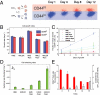
Normal and transformed HME-flopc-CD44lo cells spontaneously convert into CD44hi cells. (A) Population dynamics modeled by a simple growth model in which CD44lo cells either divide (_k_lo) or spontaneously switch (_k_s) to a CD44hi state. CD44hi cells divide (_k_hi) but cannot switch to the CD44lo state. Cell populations are quantified by flow cytometry. (B) Growth rates of purified CD44lo and CD44hi cell populations analyzed in C, n = 6, Promega CellTiter 96 AQueous Assay. (C) Quantification of the spontaneous dedifferentiation of purified CD44lo cells isolated from bulk HME cells, HME-flopcs, and HME-flopcs expressing SV40 early region only (HMLE-flopc), as well as fully transformed HME cells (HMLER) and HME-flopc cells (HMLER-flopcs) expressing the SV40 early region and Ras oncoproteins (n = 3–6, results are mean ± SEM). (D) Switching rate (_k_s) of various CD44lo cell types per population doubling time. (E) HME-flopc-CD44lo cells purified from two independent single-cell clones (no color) were mixed with fluorescent (pLV-Tomato) HME-flopc-CD44hi (CD44hi-Tom) cells to recreate a heterogeneous population. The percentages of CD44hi-Tom cells (red bars) and of de novo_-_derived CD44hi cells arising from HME-flopc-CD44lo cells expressing no fluorescent protein (gray bars) were measured over a time course of 12 d. Results are mean ± SEM.
Fig. 3.
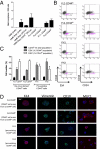
CD44hi cells display stem-like properties in vitro. (A) Mammosphere-forming ability of bulk HME or HME-flopc cells, HME-flopc-CD44lo single-cell clones (SCCs, n = 5), HME-flopc-CD44hi SCCs (n = 4), and CD44hi and CD44lo cells purified by FACS from two independent HME-flopc SCCs (FL1 and FL2). Results are mean ± SEM. (B) Representative flow cytometry plots for the markers CD44, CD24, and ESA of cells isolated from dissociated mammospheres from de novo-derived CD44hi cells (FL1-CD44hi and FL2-CD44hi) and two stable HME-flopc-CD44hi SCCs (FH1 and FH2). (C) Quantification of total CD44lo cells, CD44loESA− and CD44loESA+ populations [from (B) box P7; n = 3]. Results are mean ± SEM. (D) Frozen sections of mammospheres (5 μm) stained by immunofluorescence for myoepithelial [CD10, vimentin, and cytokeratin-14 (K14)] and luminal (K14 and MUC1) differentiation markers.
Fig. 4.
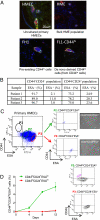
CD44loCD24+ESAlo cells isolated from primary HMECs convert into the CD44hiCD24loESAlo mammary stem-like state. (A) Single-cell suspensions of (i) uncultured primary human mammary epithelial cells (HMECs), (ii) bulk HME cells (expressing hTERT), (iii) FH1 [preexisting HME-flopc CD44hi single-cell clone (SCC)], and (iv) spontaneously arising CD44hi cells (purified from HME-flopc CD44lo SCC-FL1) were injected into the humanized mouse mammary fat pad. After 12 wk, mammary glands were sectioned and stained by immunofluorescence for basal (keratin 14, green) and luminal (pan-cytokeratin, red) markers. (B) Quantification of subpopulations of uncultured HMECs following direct isolation from primary tissue (
SI Materials and Methods
). (C) Flow cytometry plot illustrating the gating strategy used to isolate subpopulations of HMECs. P1, proposed mammary stem cell-enriched fraction; P2, CD44loCD24+ESAlo basal cells; P3, CD44loCD24+ESA+ luminal fraction with phase contrast micrographs (20×) of purified P2 and P3 cell populations. (D) Purified (via flow cytometry) primary CD44loCD24+ESAlo and CD44loCD24+ESA+ HMECs were monitored for 12 d in 2D culture for their ability to spontaneously convert into the CD44hiCD24+ESAlo cell state. Results are mean ± SEM. Flow cytometry plots of the purified CD44loCD24+ESAlo and CD44loCD24+ESA+ cell populations at day 12 are shown.
Fig. 5.
CSCs are created de novo in vivo. (A) Representative FACS profiles of tumor cell populations before injection into NOD/SCID mice and of digested tumor cell populations following 8–10 wk in vivo. (B) Quantitation of in vivo de novo-derived CD44hi cells isolated from various tumor populations, including tumor incidence, tumor weight, and a FACS analysis of immune cells recruited to the different tumor types (n = 6–9 animals/group, P < 0.001, two-way ANOVA).
Similar articles
- The epithelial-mesenchymal transition generates cells with properties of stem cells.
Mani SA, Guo W, Liao MJ, Eaton EN, Ayyanan A, Zhou AY, Brooks M, Reinhard F, Zhang CC, Shipitsin M, Campbell LL, Polyak K, Brisken C, Yang J, Weinberg RA. Mani SA, et al. Cell. 2008 May 16;133(4):704-15. doi: 10.1016/j.cell.2008.03.027. Cell. 2008. PMID: 18485877 Free PMC article. - Functional heterogeneity within the CD44 high human breast cancer stem cell-like compartment reveals a gene signature predictive of distant metastasis.
Leth-Larsen R, Terp MG, Christensen AG, Elias D, Kühlwein T, Jensen ON, Petersen OW, Ditzel HJ. Leth-Larsen R, et al. Mol Med. 2012 Sep 25;18(1):1109-21. doi: 10.2119/molmed.2012.00091. Mol Med. 2012. PMID: 22692575 Free PMC article. - Ganglioside GD2 identifies breast cancer stem cells and promotes tumorigenesis.
Battula VL, Shi Y, Evans KW, Wang RY, Spaeth EL, Jacamo RO, Guerra R, Sahin AA, Marini FC, Hortobagyi G, Mani SA, Andreeff M. Battula VL, et al. J Clin Invest. 2012 Jun;122(6):2066-78. doi: 10.1172/JCI59735. Epub 2012 May 15. J Clin Invest. 2012. PMID: 22585577 Free PMC article. Clinical Trial. - Epithelial mesenchymal transition traits in human breast cancer cell lines parallel the CD44(hi/)CD24 (lo/-) stem cell phenotype in human breast cancer.
Blick T, Hugo H, Widodo E, Waltham M, Pinto C, Mani SA, Weinberg RA, Neve RM, Lenburg ME, Thompson EW. Blick T, et al. J Mammary Gland Biol Neoplasia. 2010 Jun;15(2):235-52. doi: 10.1007/s10911-010-9175-z. Epub 2010 Jun 4. J Mammary Gland Biol Neoplasia. 2010. PMID: 20521089 Review. - Role of epithelial stem/progenitor cells in mammary cancer.
Bruno RD, Smith GH. Bruno RD, et al. Gene Expr. 2011;15(3):133-40. doi: 10.3727/105221611x13176664479368. Gene Expr. 2011. PMID: 22268295 Free PMC article. Review.
Cited by
- Inflammation and Hras signaling control epithelial-mesenchymal transition during skin tumor progression.
Wong CE, Yu JS, Quigley DA, To MD, Jen KY, Huang PY, Del Rosario R, Balmain A. Wong CE, et al. Genes Dev. 2013 Mar 15;27(6):670-82. doi: 10.1101/gad.210427.112. Genes Dev. 2013. PMID: 23512660 Free PMC article. - Poised chromatin at the ZEB1 promoter enables breast cancer cell plasticity and enhances tumorigenicity.
Chaffer CL, Marjanovic ND, Lee T, Bell G, Kleer CG, Reinhardt F, D'Alessio AC, Young RA, Weinberg RA. Chaffer CL, et al. Cell. 2013 Jul 3;154(1):61-74. doi: 10.1016/j.cell.2013.06.005. Cell. 2013. PMID: 23827675 Free PMC article. - Stem cells. Asymmetric apportioning of aged mitochondria between daughter cells is required for stemness.
Katajisto P, Döhla J, Chaffer CL, Pentinmikko N, Marjanovic N, Iqbal S, Zoncu R, Chen W, Weinberg RA, Sabatini DM. Katajisto P, et al. Science. 2015 Apr 17;348(6232):340-3. doi: 10.1126/science.1260384. Epub 2015 Apr 2. Science. 2015. PMID: 25837514 Free PMC article. - Histone deacetylase inhibitors stimulate dedifferentiation of human breast cancer cells through WNT/β-catenin signaling.
Debeb BG, Lacerda L, Xu W, Larson R, Solley T, Atkinson R, Sulman EP, Ueno NT, Krishnamurthy S, Reuben JM, Buchholz TA, Woodward WA. Debeb BG, et al. Stem Cells. 2012 Nov;30(11):2366-77. doi: 10.1002/stem.1219. Stem Cells. 2012. PMID: 22961641 Free PMC article. - Drug Delivery Using Nanoparticles for Cancer Stem-Like Cell Targeting.
Lu B, Huang X, Mo J, Zhao W. Lu B, et al. Front Pharmacol. 2016 Apr 12;7:84. doi: 10.3389/fphar.2016.00084. eCollection 2016. Front Pharmacol. 2016. PMID: 27148051 Free PMC article. Review.
References
- Teo AK, Vallier L. Emerging use of stem cells in regenerative medicine. Biochem J. 2010;428:11–23. - PubMed
- Alison MR, Islam S, Wright NA. Stem cells in cancer: Instigators and propagators? J Cell Sci. 2010;123:2357–2368. - PubMed
- Ailles LE, Weissman IL. Cancer stem cells in solid tumors. Curr Opin Biotechnol. 2007;18:460–466. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous