Exploring metabolic pathway reconstruction and genome-wide expression profiling in Lactobacillus reuteri to define functional probiotic features - PubMed (original) (raw)
Exploring metabolic pathway reconstruction and genome-wide expression profiling in Lactobacillus reuteri to define functional probiotic features
Delphine M Saulnier et al. PLoS One. 2011.
Abstract
The genomes of four Lactobacillus reuteri strains isolated from human breast milk and the gastrointestinal tract have been recently sequenced as part of the Human Microbiome Project. Preliminary genome comparisons suggested that these strains belong to two different clades, previously shown to differ with respect to antimicrobial production, biofilm formation, and immunomodulation. To explain possible mechanisms of survival in the host and probiosis, we completed a detailed genomic comparison of two breast milk-derived isolates representative of each group: an established probiotic strain (L. reuteri ATCC 55730) and a strain with promising probiotic features (L. reuteri ATCC PTA 6475). Transcriptomes of L. reuteri strains in different growth phases were monitored using strain-specific microarrays, and compared using a pan-metabolic model representing all known metabolic reactions present in these strains. Both strains contained candidate genes involved in the survival and persistence in the gut such as mucus-binding proteins and enzymes scavenging reactive oxygen species. A large operon predicted to encode the synthesis of an exopolysaccharide was identified in strain 55730. Both strains were predicted to produce health-promoting factors, including antimicrobial agents and vitamins (folate, vitamin B(12)). Additionally, a complete pathway for thiamine biosynthesis was predicted in strain 55730 for the first time in this species. Candidate genes responsible for immunomodulatory properties of each strain were identified by transcriptomic comparisons. The production of bioactive metabolites by human-derived probiotics may be predicted using metabolic modeling and transcriptomics. Such strategies may facilitate selection and optimization of probiotics for health promotion, disease prevention and amelioration.
Conflict of interest statement
Competing Interests: JV receives financial research support (no salary) and holds a patent with Biogaia since 2004, Anti-Inflammatory Activity from Lactic Acid Bacteria (WO/2004/069178). JV and DMS have a patent pending with Biogaia but not related to the work and research presented in this manuscript. JV serves as a scientific advisor for Danone. However, this does not alter the authors' adherence to all the PLoS ONE policies on sharing data and materials, as detailed online in PLoS ONE's guide for authors. All bacterial genome sequences presented in this manuscript are publicly available on National Center for Biotechnology Information (NCBI). All microarray data have also been deposited into a public database (GEO). All bacterial strains are deposited in public strain collections (ATCC, BEI), and available to everyone.
Figures
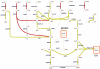
Figure 1. (Poly)saccharide metabolism in L. reuteri ATCC 55730 and ATCC PTA 6475.
Metabolic reactions present in both strains are highlighted in dark grey. Metabolic reactions only present in L. reuteri ATCC 55730 are highlighted in black. Abbreviations are indicated for reactions present at least in one strain. [e]: extracellular; 6pgg: 6-phospho-beta-D-glucosyl-(1,4)-D-glucose; acgal6p: N-acetylgalactosamine 6-phosphate; acgala: N-Acetyl-D-galactosamine; ACGALpts: N-acetylgalactosamine PTS; a-gal-D: alpha-D galactose; CELBpts: cellobiose transport via PEP:Pyr PTS; cellb: cellobiose; CPS_LPL2: capsular polysaccharide linkage unit, LPL specific; CPSS_LPL2: CPS synthase complex, LPL specific; dtdp6dm: dTDP-6-deoxy-L-mannose; dtdpddg: dTDP-4-dehydro-6-deoxy-D-glucose; dtdpddm: dTDP-4-dehydro-6-deoxy-L-mannose; dtdpglc: dTDPglucose; dttp: dTTP; f6p: D-Fructose 6-phosphate; fru: D-Fructose; FRUK: fructokinase; g1p: D-Glucose 1-phosphate; G1PTMT: glucose-1-phosphate thymidylyltransferase; g6p: D-glucose 6-phosphate; g6p-B: beta-D-glucose 6-phosphate; G6P: Glucose-6-phosphate isomerase; gal: D-galactose; gal1p: alpha-D-Galactose 1-phosphate: GALK2: galactokinase; GALM: aldose 1-epimerase; GALS3: a-galactosidase (melibiose); GALSZ: beta-galactosidase; GALT: galactose-1-phosphate uridylyltransferase; GALU: UTP-glucose-1-phosphate uridylyltransferase; glc-D: D-glucose; lcts: lactose; LCTSt6: lactose transport in/out via proton symport; MALP: maltose phosphorylase; malt: maltose; MALT: alpha-glucosidase; man: D-mannose; man1p: D-mannose 1-phosphate; man6p: D-mannose 6-phosphate; MAN6PI: mannose-6-phosphate isomerase; MANpts: D-mannose transport via PEP:Pyr PTS; melib: melibiose; PGGH: 6-phospho-beta-glucosidase; PGI: glucose-6-phosphate isomerase; PGMT: phosphoglucomutase; PGMT_B: b-phosphoglucomutase; PMANM: phosphomannomutase; raffin: raffinose; RAFGH; raffinose galactohydrolase; suc6p: sucrose 6-phosphate: SUCpts: sucrose transport via PEP:Pyr PTS; sucr: sucrose; TDPDRE: dTDP-4-dehydrorhamnose 3,5-epimerase; TDPDRR: dTDP-4-dehydrorhamnose reductase; TDPGDH: dTDPglucose 4,6-dehydratase; tre: trehalose; tre6p: alpha'-trehalose 6-phosphate; TREpts: trehalose transport via PEP:Pyr PTS; udpg; UDPglucose; UDPG4E: UDPglucose 4-epimerase; udpgal: UDPgalactose. The red boxes depict compound that can be incorporated in biomass.
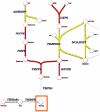
Figure 2. Exopolysaccharide gene cluster comparison in L. reuteri strain 6475, 55730, and L. rhamnosus GG.
Genes encoding similar functions in EPS biosynthesis have a similar color. Genes indicated in dark blue encode proteins putatively involved in the regulation of EPS production and polymerization. Genes indicated in white encode putative glycosyltransferases. Genes indicated in light blue encode proteins involved in the biosynthesis of the dTDP-rhamnose precursor. Genes indicated in plain blue encode hypothetical proteins. Genes indicated in dark grey encode the putative polysaccharide transporter and polymerase. Transposases are indicated by a triangle. Intergenic regions are not represented at this scale. cpsA: cell envelope related transcriptional regulator; cps1C: polysaccharide biosynthesis protein; GFT: galactofuranosyltransferase; GT: glycosyltransferase; HEP: hypothetical extracellular protein; HP: hypothetical protein; P: permease; PS: polysaccharide synthesis protein; recX: regulatory protein; rfbA: glucose-1-phosphate thymidylyltransferase; rfbB: dTDP-glucose 4,6-dehydratase; rfbC: dTDP-4-dehydrorhamnose 3,5-epimerase; rfbD: dTDP-4-dehydrorhamnose reductase; rfbF: dTDP-rhamnosyl transferase; rfbP: undecaprenyl-phosphate galactose phosphotransferase; rgbP: group 2 glycosyl transferase; r_mL_A1, r_mL_A2: glucose-1-phosphate thymidyl transferase; r_mL_B: dTDP glucose 4,6-dehydratase; r_mL_D: dTDP-4-dehydrorhamnose reductase; welE: priming glycosyltransferase; welF, welG: glycosyltransferase; welH: α-L-rhamnose α-1,3-L-rhamnosyltransferase; welI, welJ: glycosyltransferase; wzx: flippase; wzx; non-specific protein-tyrosine kinase; wzx: polysaccharide transporter; wzy: polymerase.
Figure 3. Thiamine biosynthesis pathway in L. reuteri ATCC 55730 and ATCC PTA 6475.
The thiamine synthesis pathway is predicted to be complete in strain 55730 only. Metabolic reactions present in both strains 55730 and 6475 are highlighted in dark grey. Metabolic reactions present only in L. reuteri ATCC 55730 are highlighted in black. 2mahmp: 2-methyl-4-amino-5-hydroxymethylpyrimidine diphosphate; 4ahmmp: 4-amino-5-hydroxymethyl-2-methylpyrimidine; 4ampm: 4-amino-2-methyl-5-phosphomethylpyrimidine; 4mhetz: 4-methyl-5-(2-hydroxyethyl)-thiazole; 4mpetz: 4-methyl-5-(2-phosphoethyl)-thiazole; AHMMPS: 4-amino-5-hydroxymethyl-2-methylpyrimidine synthetase; air: 5-amino-1-(5-phospho-D-ribosyl)imidazole; DXPS: 1-deoxy-D-xylulose 5-phosphate synthase; dxyl5p: 1-deoxy-D-xylulose 5-phosphate; g3p: glyceraldehyde 3-phosphate; gcald: glycolaldehyde; gly: glycine; HETZK: hydroxyethylthiazole kinase; HMPK1: hydroxymethylpyrimidine kinase (ATP); MCOOH: MPT synthase small subunit MoaD; MCOSH: MPT synthase sulfurylated small subunit (MoaD-SH); MOADCST: MoaD:cysteine sulfur transferase; PMPK: phosphomethylpyrimidine kinase; pyr; pyruvate; thm: thiamine; thm[e]: thiamine (extracellular); THMabc: thiamine transport via ABC system; thmmp: thiamine monophosphate; thmpp: thiamine diphosphate; THZPSN2: thiazole phosphate synthesis; TMDPK: thiamine diphosphokinase; TMPKr: thiamine-phosphate kinase; TMPPP: thiamine-phosphate diphosphorylase. The red boxes depict compound that can be incorporated in biomass.
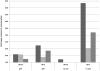
Figure 4. Transcription comparisons of genes involved in reuterin production and conversion in L. reuteri strains overtime.
12 h versus 8 h (dark grey), 16 h versus 8 h (light grey), and 24 h versus 8 h (medium grey). Strains were grown in a semi-defined medium in anoxic conditions at 37°C.
Similar articles
- Genome-scale insights into the metabolic versatility of Limosilactobacillus reuteri.
Luo H, Li P, Wang H, Roos S, Ji B, Nielsen J. Luo H, et al. BMC Biotechnol. 2021 Jul 30;21(1):46. doi: 10.1186/s12896-021-00702-w. BMC Biotechnol. 2021. PMID: 34330235 Free PMC article. - The pan-genome of Lactobacillus reuteri strains originating from the pig gastrointestinal tract.
Wegmann U, MacKenzie DA, Zheng J, Goesmann A, Roos S, Swarbreck D, Walter J, Crossman LC, Juge N. Wegmann U, et al. BMC Genomics. 2015 Dec 1;16:1023. doi: 10.1186/s12864-015-2216-7. BMC Genomics. 2015. PMID: 26626322 Free PMC article. - Pan-Genomic Approaches in Lactobacillus reuteri as a Porcine Probiotic: Investigation of Host Adaptation and Antipathogenic Activity.
Lee JY, Han GG, Choi J, Jin GD, Kang SK, Chae BJ, Kim EB, Choi YJ. Lee JY, et al. Microb Ecol. 2017 Oct;74(3):709-721. doi: 10.1007/s00248-017-0977-z. Epub 2017 Apr 24. Microb Ecol. 2017. PMID: 28439658 - Metabolic Engineering for Probiotics and their Genome-Wide Expression Profiling.
Yadav R, Singh PK, Shukla P. Yadav R, et al. Curr Protein Pept Sci. 2018;19(1):68-74. doi: 10.2174/1389203718666161111130157. Curr Protein Pept Sci. 2018. PMID: 27834122 Review. - Next-Generation Probiotic Therapy to Protect the Intestines From Injury.
Ragan MV, Wala SJ, Goodman SD, Bailey MT, Besner GE. Ragan MV, et al. Front Cell Infect Microbiol. 2022 Jun 28;12:863949. doi: 10.3389/fcimb.2022.863949. eCollection 2022. Front Cell Infect Microbiol. 2022. PMID: 35837474 Free PMC article. Review.
Cited by
- Probiotic microbes sustain youthful serum testosterone levels and testicular size in aging mice.
Poutahidis T, Springer A, Levkovich T, Qi P, Varian BJ, Lakritz JR, Ibrahim YM, Chatzigiagkos A, Alm EJ, Erdman SE. Poutahidis T, et al. PLoS One. 2014 Jan 2;9(1):e84877. doi: 10.1371/journal.pone.0084877. eCollection 2014. PLoS One. 2014. PMID: 24392159 Free PMC article. - Limosilactobacillus reuteri promotes the expression and secretion of enteroendocrine- and enterocyte-derived hormones.
Rienzi SCD, Danhof HA, Forshee MD, Roberts A, Britton RA. Rienzi SCD, et al. bioRxiv [Preprint]. 2024 Aug 31:2024.08.30.610555. doi: 10.1101/2024.08.30.610555. bioRxiv. 2024. PMID: 39257733 Free PMC article. Updated. Preprint. - Lactobacilli and other gastrointestinal microbiota of Peromyscus leucopus, reservoir host for agents of Lyme disease and other zoonoses in North America.
Milovic A, Bassam K, Shao H, Chatzistamou I, Tufts DM, Diuk-Wasser M, Barbour AG. Milovic A, et al. PLoS One. 2020 Aug 20;15(8):e0231801. doi: 10.1371/journal.pone.0231801. eCollection 2020. PLoS One. 2020. PMID: 32817657 Free PMC article. - ClC transporter activity modulates histidine catabolism in Lactobacillus reuteri by altering intracellular pH and membrane potential.
Hall AE, Engevik MA, Oezguen N, Haag A, Versalovic J. Hall AE, et al. Microb Cell Fact. 2019 Dec 12;18(1):212. doi: 10.1186/s12934-019-1264-0. Microb Cell Fact. 2019. PMID: 31830990 Free PMC article. - Genome-scale reconstruction of metabolic networks of Lactobacillus casei ATCC 334 and 12A.
Vinay-Lara E, Hamilton JJ, Stahl B, Broadbent JR, Reed JL, Steele JL. Vinay-Lara E, et al. PLoS One. 2014 Nov 3;9(11):e110785. doi: 10.1371/journal.pone.0110785. eCollection 2014. PLoS One. 2014. PMID: 25365062 Free PMC article.
References
- Reuter G. The Lactobacillus and Bifidobacterium microflora of the human intestine: composition and succession. Curr Issues Intest Microbiol. 2001;2:43–53. - PubMed
- Abrahamsson TR, Sinkiewicz G, Jakobsson T, Fredrikson M, Bjorksten B. Probiotic lactobacilli in breast milk and infant stool in relation to oral intake during the first year of life. J Pediatr Gastroenterol Nutr. 2009;49:349–354. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- P30 DK56338/DK/NIDDK NIH HHS/United States
- R01 DK065075/DK/NIDDK NIH HHS/United States
- P30 DK056338/DK/NIDDK NIH HHS/United States
- R21 AT003482/AT/NCCIH NIH HHS/United States
- R01 AT004326/AT/NCCIH NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases