The power of AAA-ATPases on the road of pre-60S ribosome maturation--molecular machines that strip pre-ribosomal particles - PubMed (original) (raw)
Review
The power of AAA-ATPases on the road of pre-60S ribosome maturation--molecular machines that strip pre-ribosomal particles
Dieter Kressler et al. Biochim Biophys Acta. 2012 Jan.
Abstract
The biogenesis of ribosomes is a fundamental cellular process, which provides the molecular machines that synthesize all cellular proteins. The assembly of eukaryotic ribosomes is a highly complex multi-step process that requires more than 200 ribosome biogenesis factors, which mediate a broad spectrum of maturation reactions. The participation of many energy-consuming enzymes (e.g. AAA-type ATPases, RNA helicases, and GTPases) in this process indicates that the expenditure of energy is required to drive ribosome assembly. While the precise function of many of these enzymes remains elusive, recent progress has revealed that the three AAA-type ATPases involved in 60S subunit biogenesis are specifically dedicated to the release and recycling of distinct biogenesis factors. In this review, we will highlight how the molecular power of yeast Drg1, Rix7, and Rea1 is harnessed to promote the release of their substrate proteins from evolving pre-60S particles and, where appropriate, discuss possible catalytic mechanisms.
Copyright © 2011 Elsevier B.V. All rights reserved.
Figures
Fig. 1
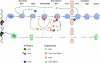
Biogenesis of the large (blue) and small (green) ribosomal subunit and the contribution of AAA-ATPases. Rix7, Rea1, and Drg1 are dedicated to the release and recycling of distinct biogenesis factors from different pre-60S particles. Major landmark of pre-60S ribosomes are depicted together with their corresponding bait proteins. These factors, namely Ssf1, Nsa1, Rix1, Arx1, and Lsg1, are only associated during a short time window with pre-60S particles and thus co-purify rather distinct pre-60S ribosomes. Ssf1 and Nsa1 purify nucleolar particles, whereas Rix1 purifies a nucleoplasmic intermediate. Arx1 represents an export-competent particle that carries export factors. Finally, Lsg1 is associated with an almost mature, cytoplasmic 60S particle. The three AAA-ATPases and their potential substrates are indicated.
Fig. 2
Schematic domain organization of Drg1 and Rix7. A, Linear representation of a protein monomer with the N-terminal, the D1 and the D2 domain depicted in yellow, orange and red, respectively. Note that the α-helical lid domains of D1 and D2 are shown in the respective dark orange and dark red color. The Walker A (A), Walker B (B) motifs, sensor-I (I) and II (II) and the arginine finger (R) of each domain are indicated. B, Representation of the secondary-structure element distribution within each ATPase domain with the α/β domain depicted in green (β-strands) and red (α-helices) and the α-helical elements of the α-helical lid domain depicted in light blue. The classic clade specific α-helix insertion is shown in dark blue. C and D, Two orthogonal views showing the hexameric organization of these AAA-ATPases. One protomer with its sub-domains is highlighted in the same colors as in A.
Fig. 3
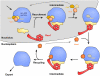
Model of a possible Rea1 cycle during 60S maturation. Rea1 associates with a nucleolar particle and releases the Ytm1-Erb1-Nop7 sub-complex. Then nucleoplasmic recruitment of Rsa4 to the pre-60S particle initiates the second Rea1-dependent reaction, which releases Rsa4 and Rea1. Note that several details of this multistep process are still elusive.
Fig. 4
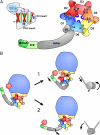
Domain organization of Rea1. A, Rea1 consists of an N-terminal domain, six ATPase modules that form the ring domain, followed by a long α-helical linker, a D/E rich region and a MIDAS domain. The predicted structural organization of an ATPase module: the central α/β domain is shown in red (α-helices) and green (β-strands), including Walker A (A), and Walker B (B) motif, sensor-I (I) and the arginine finger (R) indicated by a purple dot. The lid domain including the sensor-II (II) region is depicted in light blue. The clade-specific helix2 insert and pre-sensor-I insert (PSI) are indicated in dark blue (colors are identical to secondary structure prediction of Figs. S5 and S6.) B, Possible mechanisms of Rea1-mediated release reactions. Mechanism 1 displays a ratchet-hoist model, where energy is utilized to move the Rea1 ATPase ring on the pre-ribosome, thus creating a tension that pulls off the substrate. In mechanism 2 (power-stroke model) the Rea1 ring domain stays firmly attached at the pre-ribosome. Conformational changes within the ATPase ring are causing a power stroke: i.e. an active movement of the linker domain with the attached D/E rich region and the MIDAS domain.
Similar articles
- Rlp24 activates the AAA-ATPase Drg1 to initiate cytoplasmic pre-60S maturation.
Kappel L, Loibl M, Zisser G, Klein I, Fruhmann G, Gruber C, Unterweger S, Rechberger G, Pertschy B, Bergler H. Kappel L, et al. J Cell Biol. 2012 Nov 26;199(5):771-82. doi: 10.1083/jcb.201205021. J Cell Biol. 2012. PMID: 23185031 Free PMC article. - Mak5 and Ebp2 act together on early pre-60S particles and their reduced functionality bypasses the requirement for the essential pre-60S factor Nsa1.
Pratte D, Singh U, Murat G, Kressler D. Pratte D, et al. PLoS One. 2013 Dec 2;8(12):e82741. doi: 10.1371/journal.pone.0082741. eCollection 2013. PLoS One. 2013. PMID: 24312670 Free PMC article. - The AAA-ATPase Rea1 drives removal of biogenesis factors during multiple stages of 60S ribosome assembly.
Bassler J, Kallas M, Pertschy B, Ulbrich C, Thoms M, Hurt E. Bassler J, et al. Mol Cell. 2010 Jun 11;38(5):712-21. doi: 10.1016/j.molcel.2010.05.024. Mol Cell. 2010. PMID: 20542003 Free PMC article. - Principles of 60S ribosomal subunit assembly emerging from recent studies in yeast.
Konikkat S, Woolford JL Jr. Konikkat S, et al. Biochem J. 2017 Jan 15;474(2):195-214. doi: 10.1042/BCJ20160516. Biochem J. 2017. PMID: 28062837 Free PMC article. Review. - Driving ribosome assembly.
Kressler D, Hurt E, Bassler J. Kressler D, et al. Biochim Biophys Acta. 2010 Jun;1803(6):673-83. doi: 10.1016/j.bbamcr.2009.10.009. Epub 2009 Oct 30. Biochim Biophys Acta. 2010. PMID: 19879902 Review.
Cited by
- Long-range intramolecular allostery and regulation in the dynein-like AAA protein Mdn1.
Mickolajczyk KJ, Olinares PDB, Niu Y, Chen N, Warrington SE, Sasaki Y, Walz T, Chait BT, Kapoor TM. Mickolajczyk KJ, et al. Proc Natl Acad Sci U S A. 2020 Aug 4;117(31):18459-18469. doi: 10.1073/pnas.2002792117. Epub 2020 Jul 21. Proc Natl Acad Sci U S A. 2020. PMID: 32694211 Free PMC article. - The N-terminal extension of yeast ribosomal protein L8 is involved in two major remodeling events during late nuclear stages of 60S ribosomal subunit assembly.
Tutuncuoglu B, Jakovljevic J, Wu S, Gao N, Woolford JL Jr. Tutuncuoglu B, et al. RNA. 2016 Sep;22(9):1386-99. doi: 10.1261/rna.055798.115. Epub 2016 Jul 7. RNA. 2016. PMID: 27390266 Free PMC article. - Interrelation between protein synthesis, proteostasis and life span.
Arnsburg K, Kirstein-Miles J. Arnsburg K, et al. Curr Genomics. 2014 Feb;15(1):66-75. doi: 10.2174/1389202915666140210210542. Curr Genomics. 2014. PMID: 24653664 Free PMC article. - Mechanistic insight into eukaryotic 60S ribosomal subunit biogenesis by cryo-electron microscopy.
Greber BJ. Greber BJ. RNA. 2016 Nov;22(11):1643-1662. doi: 10.1261/rna.057927.116. RNA. 2016. PMID: 27875256 Free PMC article. Review. - Eukaryotic Ribosome Biogenesis: The 60S Subunit.
Moraleva AA, Deryabin AS, Rubtsov YP, Rubtsova MP, Dontsova OA. Moraleva AA, et al. Acta Naturae. 2022 Apr-Jun;14(2):39-49. doi: 10.32607/actanaturae.11541. Acta Naturae. 2022. PMID: 35925480 Free PMC article.
References
- Armache J.P., Jarasch A., Anger A.M., Villa E., Becker T., Bhushan S., Jossinet F., Habeck M., Dindar G., Franckenberg S., Marquez V., Mielke T., Thomm M., Berninghausen O., Beatrix B., Soding J., Westhof E., Wilson D.N., Beckmann R. Cryo-EM structure and rRNA model of a translating eukaryotic 80S ribosome at 5.5-A resolution. Proc. Natl. Acad. Sci. U. S. A. 2010;107:19748–19753. - PMC - PubMed
- Rabl J., Leibundgut M., Ataide S.F., Haag A., Ban N. Crystal structure of the eukaryotic 40S ribosomal subunit in complex with initiation factor 1. Science. 2011;331:730–736. - PubMed
- Ben-Shem A., Jenner L., Yusupova G., Yusupov M. Crystal structure of the eukaryotic ribosome. Science. 2010;330:1203–1209. - PubMed
- Kressler D., Hurt E., Baßler J. Driving ribosome assembly. Biochim. Biophys. Acta. 2010;1803:673–683. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases