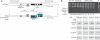High-frequency genome editing using ssDNA oligonucleotides with zinc-finger nucleases - PubMed (original) (raw)
High-frequency genome editing using ssDNA oligonucleotides with zinc-finger nucleases
Fuqiang Chen et al. Nat Methods. 2011.
Abstract
Zinc-finger nucleases (ZFNs) have enabled highly efficient gene targeting in multiple cell types and organisms. Here we describe methods for using simple ssDNA oligonucleotides in tandem with ZFNs to efficiently produce human cell lines with three distinct genetic outcomes: (i) targeted point mutation, (ii) targeted genomic deletion of up to 100 kb and (iii) targeted insertion of small genetic elements concomitant with large genomic deletions.
Figures
Figure 1
ssODN design and genome editing at the human RSK2 locus. (a) The schematic shows a 125-mer ssODN (RSK2-125) donor DNA used to incorporate three mutation types into the RSK2 locus: a silent cytosine to adenine (C to A) mutation to create a silent BamHI site, a codon conversion (TGC to GTT) to create the desired cysteine to valine change in the resulting protein and a ZFN-blocking mutation (ZBM) for each ZFN arm (ZFN-L and ZFN-R). (b) Acrylamide gel separation of amplified and BamHI-digested genomic DNA from pooled K562 cells transfected with the indicated constructs or encoding the indicated proteins and collected 2 d after nucleofection. The frequency of BamHI cleavage was quantified by densitometry. Each lane represents pooled cells from an independent transfection event. M, DNA marker (Sigma). (c) Immunoblots probing the kinase activity of the RSK2 C436V mutant in K562 cells with wild-type or ssODN-mutated RSK2 C436V (three clones) treated with fmk and phorbol 12-myristate 13-acetate (PMA) to stimulate ERK that activates RSK2 as indicated. Cell extracts were immunoprecipitated with antibodies to RSK2 and the precipitates immunoblotted with antibodies to active phosphorylated hydrophobic motif of RSK2 (anti-pHM RSK2) that detect active RSK2 or to total RSK2 (anti-RSK2). Pre-precipitation cell extracts were immunoblotted with antibodies to phosphoERK (anti-pERK) that detect active ERK or to total ERK (anti-ERK).
Figure 2
Deletion of chromosomal segments using ssODNs and ZFNs at the human AAVS1 locus. (a) General ssODN design rules for deletion of chromosomal segments relative to the ZFN cut site. Sequence distal to the ZFN cleavage site (purple) and DNA sequence containing the ZFN half-site farthest from the distal deletion sequence (green) are shown. (b) ssODN sequence used to delete 5 kb upstream of the AAVS1 ZFN cut site. The ZFN binding half-site is underlined. (c) Agarose gel separation of amplified genomic DNA from K562 cells transfected with the following constructs 2 d after nucleofection (1, ssODN plus construct encoding ZFN; 2, ssODN only; and 3, construct encoding ZFN only; see supplementary note 2 for ssODN sequence). The expected fragment sizes of the wild-type and deletion alleles are indicated. PCR fragments greater than 1.5 kb were not detected under the experimental conditions. M, DNA marker (Sigma); *3′ deletion from the ZFN cut site; **5′ and 3′ deletion off the ZFN cut site by transfecting both d5-AAVS1-0.1kb and d3-AAVS1-0.1kb deletion ssODNs; supplementary note 2); all other lanes are 5′ deletions from the cut site.
Comment in
- The author file: Greg Davis.
Baker M. Baker M. Nat Methods. 2011 Sep;8(9):699. doi: 10.1038/nmeth.1677. Nat Methods. 2011. PMID: 21984998 No abstract available.
Similar articles
- Gene editing using ssODNs with engineered endonucleases.
Chen F, Pruett-Miller SM, Davis GD. Chen F, et al. Methods Mol Biol. 2015;1239:251-65. doi: 10.1007/978-1-4939-1862-1_14. Methods Mol Biol. 2015. PMID: 25408411 - Rapid mutation of endogenous zebrafish genes using zinc finger nucleases made by Oligomerized Pool ENgineering (OPEN).
Foley JE, Yeh JR, Maeder ML, Reyon D, Sander JD, Peterson RT, Joung JK. Foley JE, et al. PLoS One. 2009;4(2):e4348. doi: 10.1371/journal.pone.0004348. Epub 2009 Feb 9. PLoS One. 2009. PMID: 19198653 Free PMC article. - Codon swapping of zinc finger nucleases confers expression in primary cells and in vivo from a single lentiviral vector.
Abarrategui-Pontes C, Créneguy A, Thinard R, Fine EJ, Thepenier V, Fournier le RL, Cradick TJ, Bao G, Tesson L, Podevin G, Anegon I, Nguyen TH. Abarrategui-Pontes C, et al. Curr Gene Ther. 2014;14(5):365-76. doi: 10.2174/156652321405140926161748. Curr Gene Ther. 2014. PMID: 25687502 - Advances in genetic modification of farm animals using zinc-finger nucleases (ZFN).
Petersen B, Niemann H. Petersen B, et al. Chromosome Res. 2015 Feb;23(1):7-15. doi: 10.1007/s10577-014-9451-7. Chromosome Res. 2015. PMID: 25596823 Review. - Genome engineering with zinc-finger nucleases.
Carroll D. Carroll D. Genetics. 2011 Aug;188(4):773-82. doi: 10.1534/genetics.111.131433. Genetics. 2011. PMID: 21828278 Free PMC article. Review.
Cited by
- Direct reprogramming of urine-derived cells with inducible MyoD for modeling human muscle disease.
Kim EY, Page P, Dellefave-Castillo LM, McNally EM, Wyatt EJ. Kim EY, et al. Skelet Muscle. 2016 Sep 15;6:32. doi: 10.1186/s13395-016-0103-9. eCollection 2016. Skelet Muscle. 2016. PMID: 27651888 Free PMC article. - Making gene editing accessible in resource limited environments: recommendations to guide a first-time user.
Goolab S, Scholefield J. Goolab S, et al. Front Genome Ed. 2024 Sep 25;6:1464531. doi: 10.3389/fgeed.2024.1464531. eCollection 2024. Front Genome Ed. 2024. PMID: 39386178 Free PMC article. Review. - Functional testing of a human PBX3 variant in zebrafish reveals a potential modifier role in congenital heart defects.
Farr GH 3rd, Imani K, Pouv D, Maves L. Farr GH 3rd, et al. Dis Model Mech. 2018 Oct 18;11(10):dmm035972. doi: 10.1242/dmm.035972. Dis Model Mech. 2018. PMID: 30355621 Free PMC article. - Cell and Gene Therapies for Mucopolysaccharidoses: Base Editing and Therapeutic Delivery to the CNS.
Christensen CL, Ashmead RE, Choy FYM. Christensen CL, et al. Diseases. 2019 Jun 26;7(3):47. doi: 10.3390/diseases7030047. Diseases. 2019. PMID: 31248000 Free PMC article. Review. - Seamless genome editing in human pluripotent stem cells using custom endonuclease-based gene targeting and the piggyBac transposon.
Yusa K. Yusa K. Nat Protoc. 2013 Oct;8(10):2061-78. doi: 10.1038/nprot.2013.126. Epub 2013 Sep 26. Nat Protoc. 2013. PMID: 24071911
References
- Urnov FD, et al. Nature. 2005;435:646–651. - PubMed
- Bibikova M, et al. Science. 2003;300:764. - PubMed
- Porteus MH, Baltimore D. Science. 2003;300:763. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources