Genomic sequencing of colorectal adenocarcinomas identifies a recurrent VTI1A-TCF7L2 fusion - PubMed (original) (raw)
. 2011 Sep 4;43(10):964-968.
doi: 10.1038/ng.936.
Adam J Bass # 1 2 3 4, Lear E Brace 1, Alex H Ramos 1 4, Yotam Drier 5, Kristian Cibulskis 4, Carrie Sougnez 4, Douglas Voet 4, Gordon Saksena 4, Andrey Sivachenko 4, Rui Jing 4, Melissa Parkin 4, Trevor Pugh 1 4, Roel G Verhaak 1 4, Nicolas Stransky 4, Adam T Boutin 1, Jordi Barretina 4, David B Solit 6, Evi Vakiani 7, Wenlin Shao 8, Yuji Mishina 8, Markus Warmuth 8, Jose Jimenez 9, Derek Y Chiang 10, Sabina Signoretti 11 12, William G Kaelin Jr 1 2, Nicole Spardy 1, William C Hahn 1 2 3 4, Yujin Hoshida 4, Shuji Ogino 1 11 12 13, Ronald A DePinho 1 2 14 15, Lynda Chin 1 4 15 16, Levi A Garraway 1 2 3 4, Charles S Fuchs 1 2 13, Jose Baselga 9 17, Josep Tabernero 9, Stacey Gabriel 4, Eric S Lander 4 18 19, Gad Getz 4, Matthew Meyerson 1 3 4 11
Affiliations
- PMID: 21892161
- PMCID: PMC3802528
- DOI: 10.1038/ng.936
Genomic sequencing of colorectal adenocarcinomas identifies a recurrent VTI1A-TCF7L2 fusion
Adam J Bass et al. Nat Genet. 2011.
Abstract
Prior studies have identified recurrent oncogenic mutations in colorectal adenocarcinoma and have surveyed exons of protein-coding genes for mutations in 11 affected individuals. Here we report whole-genome sequencing from nine individuals with colorectal cancer, including primary colorectal tumors and matched adjacent non-tumor tissues, at an average of 30.7× and 31.9× coverage, respectively. We identify an average of 75 somatic rearrangements per tumor, including complex networks of translocations between pairs of chromosomes. Eleven rearrangements encode predicted in-frame fusion proteins, including a fusion of VTI1A and TCF7L2 found in 3 out of 97 colorectal cancers. Although TCF7L2 encodes TCF4, which cooperates with β-catenin in colorectal carcinogenesis, the fusion lacks the TCF4 β-catenin-binding domain. We found a colorectal carcinoma cell line harboring the fusion gene to be dependent on VTI1A-TCF7L2 for anchorage-independent growth using RNA interference-mediated knockdown. This study shows previously unidentified levels of genomic rearrangements in colorectal carcinoma that can lead to essential gene fusions and other oncogenic events.
Figures
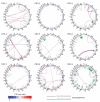
Figure 1
DNA structural rearrangements and copy number alterations detected in the nine colorectal tumors displayed as CIRCOS plots. Chromosomes are arranged circularly end-to-end with each chromosome’s cytobands marked in the outer ring. The inner ring displays copy number data inferred from whole-genome sequencing with blue indicating losses and red indicating gains. Within the circle, rearrangements are shown as arcs with intrachromosomal events in green and interchromosomal translocations in purple.
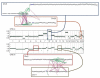
Figure 2
Complex rearrangements between chromosome pairs in two colorectal carcinomas. The central portion of the figure contains copy-number profiles across all chromosomes with the chromosome identity labeled across the x axis and the scale for copy-number ratio (log2) depicted on the y axis of each plot. The upper plot shows the tumor CRC-4, and the lower plot shows the copy-number profile for CRC-6, with the black dots marking the copy-number ratio inferred along each locus across the genome. The upper inset boxes show detailed views of the copy numbers and rearrangements for chromosomes 8 (dark blue) and 20 (ochre) for CRC-4 with the centromere labeled as a purple circle. Rearrangements detected by dRanger are shown in green (intrachromosomal) and purple (interchromosomal). The lower inset boxes show detailed copy-number and rearrangement images for CRC-6, with inset boxes showing chromosome 5 (red) and 11 (gray), with lines marking positions of genomic rearrangements.
Figure 3
Recurrent gene fusion between VTI1A and TCF7L2. (a) The upper schematic depicts the positions of exons (vertical lines) within VTI1A and TCF7L2, which reside adjacent to each other on chromosome 10. The blowup displays the locations of discordant paired-end reads found in tumor CRC-9 for which one read (labeled in blue) is in an intron of VTI1A and the other read (labeled in red) is in an intron of TCF7L2. (b) The upper schematic depicts the structure of the predicted fusion transcript generated by the fusion. The presence of the exact reads spanning the fusion of the two introns (marked by lightning bolt) is depicted in the inset with regions of the reads corresponding to original VTI1A intron in blue and those of TCF7L2 in red. (c) The protein domain structure of native VTI1A and TCF4-TCF7L2, including the two alternate C-terminal tails of TCF4, are shown. Below are the structures of the fusion protein encoded by the fusion of exon 3 of VTI1A to exon 4 of TCF7L2 identified in CRC-9. Two variants of the fusion are shown as data from the NCI-H508 cell line and reveal that variants encoding both the full length (E-tail) and shorter (B-tail) C termini are both expressed (data not shown). (d) Measurement of the relative expression of the VTI1A-TCF7L2 mRNA in NCI-H508 cells infected with one of two short hairpin RNA constructs targeting the fusion gene relative to expression in a cell infected with control vectors targeting GFP. (e) Anchorage-independent growth of the NCIH508 cell line, which expresses VTI1A-TCF7L2, and negative control DLD-1 colorectal adenocarcinoma cells following RNA-interference–mediated knockdown of VTI1A-TCF7L2 compared to control knockdown targeting GFP.
References
- Fearon ER, Vogelstein B. A genetic model for colorectal tumorigenesis. Cell. 1990;61:759–767. - PubMed
- Sjöblom T, et al. The consensus coding sequences of human breast and colorectal cancers. Science. 2006;314:268–274. - PubMed
- Wood LD, et al. The genomic landscapes of human breast and colorectal cancers. Science. 2007;318:1108–1113. - PubMed
- Clevers H. Wnt/B-Catenin signaling in development and disease. Cell. 2006;127:469–480. - PubMed
- Nishisho I, et al. Mutations of chromsome 5q21 genes in FAP and colorectal cancer patients. Science. 1991;253:665–669. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- K08CA134931/CA/NCI NIH HHS/United States
- P50 CA101942/CA/NCI NIH HHS/United States
- K08 CA134931/CA/NCI NIH HHS/United States
- P50CA127003/CA/NCI NIH HHS/United States
- P50 CA127003/CA/NCI NIH HHS/United States
- U54 HG003067/HG/NHGRI NIH HHS/United States
- R01 CA068490/CA/NCI NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases