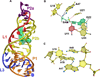Riboswitch structure in the ligand-free state - PubMed (original) (raw)
Review
. 2012 May-Jun;3(3):369-84.
doi: 10.1002/wrna.114. Epub 2011 Sep 28.
Affiliations
- PMID: 21957061
- PMCID: PMC3252462
- DOI: 10.1002/wrna.114
Review
Riboswitch structure in the ligand-free state
Joseph A Liberman et al. Wiley Interdiscip Rev RNA. 2012 May-Jun.
Abstract
Molecular investigations of riboswitches bound to small-molecule effectors have produced a wealth of information on how these molecules achieve high affinity and specificity for a target ligand. X-ray crystal structures have been determined for the ligand-free state for representatives of the preQ₁-I, S-adenosylmethionine I, lysine, and glycine aptamer classes. These structures in conjunction with complimentary techniques, such as in-line probing, NMR spectroscopy, Förster resonance energy transfer, small-angle scattering, and computational simulations, have demonstrated that riboswitches adopt multiple conformations in the absence of ligand. Despite a number of investigations that support ligand-dependent folding, mounting evidence suggests that free-state riboswitches interact with their effectors in the sub-populations of largely prefolded states as embodied by the principle of conformational selection, which has been documented extensively for protein-mediated ligand interactions. Fundamental riboswitch investigations of the bound and free states have advanced our understanding of RNA folding, ligand recognition, and how these factors culminate in communication between an aptamer and its expression platform. An understanding of these topics is essential to comprehend riboswitch gene regulation at the molecular level, which has already provided a basis to understand the mechanism of action of natural antimicrobials.
Copyright © 2011 John Wiley & Sons, Ltd.
Figures
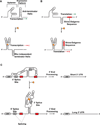
Figure 1
Representative mechanisms of action by which riboswitches regulate gene expression. (a) Schematic depiction of transcriptional repression by a bacterial riboswitch. In the ligand-free state the aptamer is not stably folded and favors anti-terminator helix formation, thus allowing transcription. Upon binding ligand (star) the folded structure predominates and a rho-independent terminator helix forms, thus attenuating transcription. (b) A schematic depiction of translational regulation by a bacterial riboswitch. In the ligand-free state the aptamer is not stably folded and favors an exposed Shine-Dalgarno (SD) sequence favoring translation. Upon ligand binding the folded structure predominates and the SD sequence is sequestered, thus attenuating translation. (c) A schematic depiction of splicing regulation by the THIC TPP riboswitch from Arabidopsis thaliana. When TPP levels are low the aptamer base pairs with a sequence used in 5´-splice-site selection. This bypasses splicing and necessitates use of a 3´-end processing signal (red rectangle) that yields a short 3’-UTR associated with high protein expression levels. When TPP levels are high the aptamer binds TPP during transcription and exposes the 5´-splice site. The ligand-bound aptamer results in splicing whereby the upstream 3´-end processing site is removed, which necessitates the use of an inferior downstream processing site. This results in a longer, unstable mRNA that produces lower quantities of THIC protein.
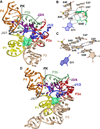
Figure 2
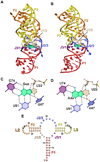
Cartoon and ball-and-stick diagrams of the SAM-I riboswitch aptamer domains. (a) Cartoon depiction of the T. tengcongensis SAM-I riboswitch structure in the ligand-bound state. SAM is depicted as space-filling surface (green); PK is the abbreviation for pseudoknot. A common color code is used in all figures that corresponds to the pairing (P) and joining (J) regions. Dashed black lines show putative hydrogen bonds. The coordinates are from PDB entry 2GIS. (b) The SAM ligand binding site in the bound state derived from (a). (c) The SAM binding site in the ligand-free state. The coordinates are from PDB entry 3IQP. (d) Cartoon diagram of the tertiary structure of the B. subtilis SAM-I riboswitch from PDB entry 3NPB.
Figure 3
Cartoon and ball-and-stick diagrams of the metX SAM-II riboswitch from the Sargasso Sea metagenome. (a) Cartoon depicting the tertiary structure of the SAM-II riboswitch in the bound state. The coordinates are from PDB entry 2QWY. (b) Ball-and-stick diagram of the structure in (a) showing interactions with SAM; red dashed lines indicate putative electrostatic interactions. (c) Close up view of the interaction between A19 and A47 showing the potential for chemical modification. (See the text for details).
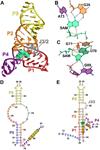
Figure 4
Cartoon, ball-and-stick, and secondary structure diagrams of the SAM-III (SMK) riboswitch from E. faecalis. (a) Cartoon diagram showing the tertiary structure of the SMK riboswitch in the ligand-bound state. The coordinates are from PDB entry 3E5C. (b) Ball-and-stick diagram of the structure in (a) showing interactions between SAM and nucleotides A73 and G26 involved in SAM binding. (c) Additional interactions for the structure in (a) between SAM and nucleotides that stabilize ligand binding. (d) Secondary structure of the E. faecalis SMK riboswitch when SAM is absent. Nucleotides in blue are present in SMK59 and absent in SMK51; the RBS is highlighted in yellow. (e) Secondary structure of the E. faecalis SMK riboswitch when ligand is present. Nucleotides in blue are present in SMK59 and absent in SMK51.
Figure 5
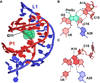
Cartoon, ball-and-stick, and secondary structure diagrams of the adenine and guanine riboswitches. (a) Cartoon diagram showing the tertiary structure of the B. subtilis xpt guanine binding riboswitch. The coordinates are from PDB entry 1Y27. (b) Cartoon diagram showing the tertiary structure of the V. vulnificus add adenine riboswitch. The coordinates are from PDB entry 1Y26. (c) Ball-and-stick diagram of the structure in (a) depicting interactions between guanine and the riboswitch. (d) Ball-and-stick diagram of the structure in (b) depicting interactions between adenine and the riboswitch. (e) Secondary structure diagram of the pbuE adenine riboswitch in the bound state. Nucleotides that were mutated to disrupt loop-loop interactions are highlighted in green as described in the text and ref..
Figure 6
Cartoon and ball-and-stick diagrams of the T. tengcongensis preQ1-I riboswitch. (a) Cartoon diagram showing the tertiary structure of the riboswitch. The A-rich pseudoknotted tail is depicted in blue. The coordinates are from PDB entry 3Q50. (b) Ball-and-stick diagram of the structure in (a) depicting interactions between preQ1 and the aptamer. (c) The preQ1-binding site in the ligand-free state from PDB entry 3Q51.
Figure 7
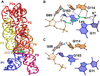
Cartoon and ball-and-stick diagrams of the T. maritima lysine riboswitch. (a) Cartoon diagram depicting the tertiary structure of the lysine riboswitch in the bound state. The coordinates are from PDB entry 3DIL. (b) Ball-and-stick diagram of the structure in (a) depicting hydrogen bonds with the riboswitch and coordination with a potassium ion; water is shown as a red sphere. The coordinates were derived from PDB entry 3DIL (c) The lysine binding site in the ligand-free state riboswitch. No significant conformational changes are observed compared to the structure in (b). The coordinates were derived from PDB entry 3DIS.
Figure 8
Cartoon and ball-and-stick diagrams of the V. cholerae VCII glycine riboswitch. (a) Cartoon diagram depicting the tertiary structure of the glycine VCII riboswitch in the ligand-bound state. The coordinates are from PDB entry 3OWW. (b) Ball-and-stick diagram of the structure in (a) depicting hydrogen bonds between the riboswitch and glycine. (c) The VCII glycine-binding site from a ligand-free crystal structure of the VCII glycine riboswitch. There are no substantial conformational changes relative to the bound structure. The coordinates were derived from PDB entry 3OX0.
SIDEBAR 1
A cartoon illustrating conformational selection by a riboswitch, which is a prominent mode of ligand recognition by proteins. In state 1 only secondary structures are formed. In states 2–5, tertiary structure has formed but the ligand is not bound. The aptamer exists as an equilibrium of conformational states with both local changes to the ligand-binding pocket, as well as long-range tertiary interactions between different pairing regions. Only conformations with a fully formed, unblocked binding sites are receptive to ligand binding. Upon binding the aptamer adopts a stable conformation (state 6) that effects expression of the associated mRNA.
Similar articles
- The dynamic nature of RNA as key to understanding riboswitch mechanisms.
Haller A, Soulière MF, Micura R. Haller A, et al. Acc Chem Res. 2011 Dec 20;44(12):1339-48. doi: 10.1021/ar200035g. Epub 2011 Jun 16. Acc Chem Res. 2011. PMID: 21678902 - Synthetic ligands for PreQ1 riboswitches provide structural and mechanistic insights into targeting RNA tertiary structure.
Connelly CM, Numata T, Boer RE, Moon MH, Sinniah RS, Barchi JJ, Ferré-D'Amaré AR, Schneekloth JS Jr. Connelly CM, et al. Nat Commun. 2019 Apr 2;10(1):1501. doi: 10.1038/s41467-019-09493-3. Nat Commun. 2019. PMID: 30940810 Free PMC article. - ITC analysis of ligand binding to preQ₁ riboswitches.
Liberman JA, Bogue JT, Jenkins JL, Salim M, Wedekind JE. Liberman JA, et al. Methods Enzymol. 2014;549:435-50. doi: 10.1016/B978-0-12-801122-5.00018-0. Methods Enzymol. 2014. PMID: 25432759 Free PMC article. - Linking aptamer-ligand binding and expression platform folding in riboswitches: prospects for mechanistic modeling and design.
Aboul-ela F, Huang W, Abd Elrahman M, Boyapati V, Li P. Aboul-ela F, et al. Wiley Interdiscip Rev RNA. 2015 Nov-Dec;6(6):631-50. doi: 10.1002/wrna.1300. Epub 2015 Sep 11. Wiley Interdiscip Rev RNA. 2015. PMID: 26361734 Free PMC article. Review. - Structural studies of the purine and SAM binding riboswitches.
Gilbert SD, Montange RK, Stoddard CD, Batey RT. Gilbert SD, et al. Cold Spring Harb Symp Quant Biol. 2006;71:259-68. doi: 10.1101/sqb.2006.71.015. Cold Spring Harb Symp Quant Biol. 2006. PMID: 17381305 Review.
Cited by
- Observation of preQ1-II riboswitch dynamics using single-molecule FRET.
Warnasooriya C, Ling C, Belashov IA, Salim M, Wedekind JE, Ermolenko DN. Warnasooriya C, et al. RNA Biol. 2019 Sep;16(9):1086-1092. doi: 10.1080/15476286.2018.1536591. Epub 2018 Oct 30. RNA Biol. 2019. PMID: 30328747 Free PMC article. - Riboswitch structure and dynamics by smFRET microscopy.
Suddala KC, Walter NG. Suddala KC, et al. Methods Enzymol. 2014;549:343-73. doi: 10.1016/B978-0-12-801122-5.00015-5. Methods Enzymol. 2014. PMID: 25432756 Free PMC article. - Ligand-mediated and tertiary interactions cooperatively stabilize the P1 region in the guanine-sensing riboswitch.
Hanke CA, Gohlke H. Hanke CA, et al. PLoS One. 2017 Jun 22;12(6):e0179271. doi: 10.1371/journal.pone.0179271. eCollection 2017. PLoS One. 2017. PMID: 28640851 Free PMC article. - Characterizing slow chemical exchange in nucleic acids by carbon CEST and low spin-lock field R(1ρ) NMR spectroscopy.
Zhao B, Hansen AL, Zhang Q. Zhao B, et al. J Am Chem Soc. 2014 Jan 8;136(1):20-3. doi: 10.1021/ja409835y. Epub 2013 Dec 18. J Am Chem Soc. 2014. PMID: 24299272 Free PMC article. - Computational Methods for Modeling Aptamers and Designing Riboswitches.
Gong S, Wang Y, Wang Z, Zhang W. Gong S, et al. Int J Mol Sci. 2017 Nov 17;18(11):2442. doi: 10.3390/ijms18112442. Int J Mol Sci. 2017. PMID: 29149090 Free PMC article. Review.
References
- Mironov AS, Gusarov I, Rafikov R, Lopez LE, Shatalin K, Kreneva RA, Perumov DA, Nudler E. Sensing small molecules by nascent RNA: a mechanism to control transcription in bacteria. Cell. 2002;111:747–756. - PubMed
- Winkler W, Nahvi A, Breaker RR. Thiamine derivatives bind messenger RNAs directly to regulate bacterial gene expression. Nature. 2002;419:952–956. - PubMed
- Wachter A. Riboswitch-mediated control of gene expression in eukaryotes. RNA Biol. 2010;7:67–76. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources