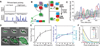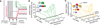Stochastic pulse regulation in bacterial stress response - PubMed (original) (raw)
Stochastic pulse regulation in bacterial stress response
James C W Locke et al. Science. 2011.
Abstract
Gene regulatory circuits can use dynamic, and even stochastic, strategies to respond to environmental conditions. We examined activation of the general stress response mediated by the alternative sigma factor, σ(B), in individual Bacillus subtilis cells. We observed that energy stress activates σ(B) in discrete stochastic pulses, with increasing levels of stress leading to higher pulse frequencies. By perturbing and rewiring the endogenous system, we found that this behavior results from three key features of the σ(B) circuit: an ultrasensitive phosphorylation switch; stochasticity ("noise"), which activates that switch; and a mixed (positive and negative) transcriptional feedback, which can both amplify a pulse and switch it off. Together, these results show how prokaryotes encode signals using stochastic pulse frequency modulation through a compact regulatory architecture.
Figures
Fig. 1
Energy stress modulates the frequency of stochastic pulses of σB activation. (A) Schematic of FM pulse regulation. The input signal (black line) controls the frequency of stochastic pulses (blue line, schematic). (B) Schematic diagram of σB regulatory interactions and states (7). When RsbV (V) is phosphorylated (OFF state), σB is sequestered by RsbW (W) and inactive (28). Under energy stresses such as MPA, RsbV is dephosphorylated by the RsbQP phosphatase complex (QP) (29). Other stress inputs are mediated by the RsbTU phosphatase complex (not shown; see SOM text for discussion). Dephosphorylated RsbV can bind to RsbW, releasing σB to activate target genes, including its own operon (30), and the yfp reporter (yellow). (C) Promoter activity of the PsigB -YFP reporter pulses in individual lineages (colored solid lines), and its mean and standard deviation across all lineages in four data sets (dashed line and shaded area, respectively). (D) Filmstrip of σB activation at 60 µg/mL MPA. Heterogeneous expression levels of PsigB -YFP reflect pulsing activity. (E) MPA concentration strongly modulates the mean frequency, while more weakly modulating the mean amplitude and duration, of pulses. Error bars, mean ± SEM. (F) Pulse amplitude histograms for varying levels of MPA. In (E) and (F), each data point represents data from four microcolonies, acquired on two different days.
Fig. 2
Pulsing is noise-dependent and involves an ultrasensitive phosphoswitch. (A) Pulse frequency in long cells (green) is strongly reduced compared with short cells (gray; data replotted from Fig. 1E). Error bars, mean ± SEM. (B) Variability in PsigB -YFP expression decreases with increasing cell length (see fig. S8). Equal numbers of cells (represented by dots) are plotted in each log-spaced bin (delimited by gray vertical lines). (Inset) Overlay of phase contrast and PsigB -YFP expression (green) at different cell lengths. Note greater σB variability in short cells. (C) σB expression is ultrasensitive to RsbQP phosphatase levels. Each dot represents the mean RsbQP-YFP level and PsigB -CFP level of one cell, using the strain shown schematically (table S1). The red line is a Hill function with Hill coefficient _n_H = 2.12 (95%CI, _n_H = 2.09 to 2.15).
Fig. 3
A mixed transcriptional feedback loop amplifies and terminates pulses. (A) Schematic diagram of supra- and subthreshold protocols. Before time-lapse acquisition (gray region), phosphatase is induced to a constant level by addition of xylose. After the start of acquisition, isopropyl-β-
d
-thiogalactopyranoside (IPTG) is added to induce rsbVWB to levels greater than (solid red line) or less than (dashed red line) the level of phosphatase. This results in pulsed (solid green line) or sustained (dashed green line) σB activity dynamics. (B) σB promoter activity exhibits a transition between sub- and suprathreshold behaviors. Each trace shows the mean PsigB promoter activity averaged over four colonies. The promoter activity of the IPTG-inducible σB operon (x axis) was estimated using a separate strain containing a similar IPTG-inducible yfp reporter. Two repeat movies showed similar behaviors. (Inset) Schematic diagram of strain used for this experiment (table S1). (C) A minimal mathematical model of the open-loop σB network reproduces the main features of the experimental data. (Inset) In this model, the unphosphorylated activator, A, directly activates target genes (see SOM).
Fig. 4
Mechanism of FM pulse control. (A) Schematic time course of phosphatase RsbQP (denoted P, purple), free σB (σB, green), and kinase (W, red) during a pulse cycle. Circled numbers indicate specific steps in (B). (B) Schematic diagram of pulse control. The relative concentration of each component is indicated by size. (1) Initial state: System components are at low levels, kinase activities exceed phosphatase activities, and therefore RsbV is mostly phosphorylated. A threshold-crossing upward fluctuation in RsbQP level dephosphorylates VP, leading to (2) Pulse Initiation. Activation of σB (indicated by glowing halo) leads to up-regulation of operon components (operon feedback). (3) Pulse peak: σB activity peaks just before RsbW kinase activity exceeds phosphatase activity. (4) Termination: Rephosphorylation of RsbV shuts the system off. (5) Dilution: Component levels reset to the original state. (C)Mechanism of frequency modulation. Fluctuations in phosphatase level (purple arrow from state 1 to 2) can cross the kinase threshold (red line) to initiate a pulse, with amplitude determined by the size of fluctuation (dashed line). Increased stress shifts the distribution of phosphatase levels from lower to higher values (dark and light gray, respectively), increasing the frequency of threshold-crossing events and thereby increasing pulse frequency (inset). (D) Tuning of phosphatase expression by IPTG (strain indicated schematically in inset) can regulate pulse frequency. Gray dashed lines show a similar behavior for the mathematical model (fit to data). Each data point represents statistics from two colonies. Two repeat data sets showed similar trends.
Similar articles
- Protein-protein interactions that regulate the energy stress activation of sigma(B) in Bacillus subtilis.
Delumeau O, Lewis RJ, Yudkin MD. Delumeau O, et al. J Bacteriol. 2002 Oct;184(20):5583-9. doi: 10.1128/JB.184.20.5583-5589.2002. J Bacteriol. 2002. PMID: 12270815 Free PMC article. - Stochastic pulsing of gene expression enables the generation of spatial patterns in Bacillus subtilis biofilms.
Nadezhdin E, Murphy N, Dalchau N, Phillips A, Locke JCW. Nadezhdin E, et al. Nat Commun. 2020 Feb 19;11(1):950. doi: 10.1038/s41467-020-14431-9. Nat Commun. 2020. PMID: 32075967 Free PMC article. - Fluoro-phenyl-styrene-sulfonamide, a novel inhibitor of σB activity, prevents the activation of σB by environmental and energy stresses in Bacillus subtilis.
Ringus DL, Gaballa A, Helmann JD, Wiedmann M, Boor KJ. Ringus DL, et al. J Bacteriol. 2013 Jun;195(11):2509-17. doi: 10.1128/JB.00107-13. Epub 2013 Mar 22. J Bacteriol. 2013. PMID: 23524614 Free PMC article. - Extra cytoplasmic function σ factor activation.
Ho TD, Ellermeier CD. Ho TD, et al. Curr Opin Microbiol. 2012 Apr;15(2):182-8. doi: 10.1016/j.mib.2012.01.001. Epub 2012 Feb 28. Curr Opin Microbiol. 2012. PMID: 22381678 Free PMC article. Review. - The RsbRST stress module in bacteria: a signalling system that may interact with different output modules.
Pané-Farré J, Lewis RJ, Stülke J. Pané-Farré J, et al. J Mol Microbiol Biotechnol. 2005;9(2):65-76. doi: 10.1159/000088837. J Mol Microbiol Biotechnol. 2005. PMID: 16319496 Review.
Cited by
- Filtering input fluctuations in intensity and in time underlies stochastic transcriptional pulses without feedback.
Sassi AS, Garcia-Alcala M, Kim MJ, Cluzel P, Tu Y. Sassi AS, et al. Proc Natl Acad Sci U S A. 2020 Oct 27;117(43):26608-26615. doi: 10.1073/pnas.2010849117. Epub 2020 Oct 12. Proc Natl Acad Sci U S A. 2020. PMID: 33046652 Free PMC article. - Optimal transcriptional regulation of dynamic bacterial responses to sudden drug exposures.
Schultz D, Stevanovic M, Tsimring LS. Schultz D, et al. Biophys J. 2022 Nov 1;121(21):4137-4152. doi: 10.1016/j.bpj.2022.09.028. Epub 2022 Sep 26. Biophys J. 2022. PMID: 36168291 Free PMC article. - Resonant learning in scale-free networks.
Goldman S, Aldana M, Cluzel P. Goldman S, et al. PLoS Comput Biol. 2023 Feb 21;19(2):e1010894. doi: 10.1371/journal.pcbi.1010894. eCollection 2023 Feb. PLoS Comput Biol. 2023. PMID: 36809235 Free PMC article. - Functional roles of pulsing in genetic circuits.
Levine JH, Lin Y, Elowitz MB. Levine JH, et al. Science. 2013 Dec 6;342(6163):1193-200. doi: 10.1126/science.1239999. Science. 2013. PMID: 24311681 Free PMC article. Review. - Microfluidics for long-term single-cell time-lapse microscopy: Advances and applications.
Allard P, Papazotos F, Potvin-Trottier L. Allard P, et al. Front Bioeng Biotechnol. 2022 Oct 12;10:968342. doi: 10.3389/fbioe.2022.968342. eCollection 2022. Front Bioeng Biotechnol. 2022. PMID: 36312536 Free PMC article. Review.
References
- Rosenfeld N, Young JW, Alon U, Swain PS, Elowitz MB. Science. 2005;307:1962. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- P50 GM068763/GM/NIGMS NIH HHS/United States
- R01 GM079771/GM/NIGMS NIH HHS/United States
- HHMI/Howard Hughes Medical Institute/United States
- R01GM079771/GM/NIGMS NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases