A metabolic network approach for the identification and prioritization of antimicrobial drug targets - PubMed (original) (raw)
Review
A metabolic network approach for the identification and prioritization of antimicrobial drug targets
Arvind K Chavali et al. Trends Microbiol. 2012 Mar.
Abstract
For many infectious diseases, novel treatment options are needed in order to address problems with cost, toxicity and resistance to current drugs. Systems biology tools can be used to gain valuable insight into pathogenic processes and aid in expediting drug discovery. In the past decade, constraint-based modeling of genome-scale metabolic networks has become widely used. Focusing on pathogen metabolic networks, we review in silico strategies used to identify effective drug targets and highlight recent successes as well as limitations associated with such computational analyses. We further discuss how accounting for the host environment and even targeting the host may offer new therapeutic options. These systems-level approaches are beginning to provide novel avenues for drug targeting against infectious agents.
Copyright © 2011 Elsevier Ltd. All rights reserved.
Figures
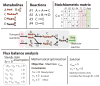
Figure I
A generic metabolic model
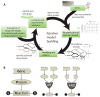
Figure 1. Iterative process of model building and refinement
(a) A functionally annotated genome along with data from biochemical literature is used to assemble a network reconstruction. Flux balance analysis (FBA) allows for modeling and simulation of the reconstructed network. Advanced network analyses (such as gene essentiality or flux variability) allow for identifying potential anti-microbial drug targets. These targets can then be associated with drugs using bioinformatics approaches and obtaining target-drug information from a variety of publicly available databases (e.g. STITCH or DrugBank). Predictions involving targets and drugs can be experimentally validated. Any discrepancies between computational predictions and experimental validation can be informative to improving upon and refining the original reconstruction and modeling platform. (b) Gene-protein-reaction (GPR) relationships, central to the assembly of a metabolic reconstruction, define the genes and gene products needed for each enzymatic reaction. Isozymes can be represented with ‘OR’ statements, while enzyme subunits required to function together to catalyze a particular reaction can be represented with ‘AND’ statements.
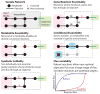
Figure 2. Drug targeting in metabolic networks
Various strategies for identifying drug targets from performing FBA on metabolic reconstructions are illustrated. The sample network shows an input media that represents the environment and exchange reactions, intracellular reactions, and an objective reaction that drains metabolites out of the system. An essential reaction and metabolite that, when removed, block any flux through the objective are highlighted in red. In the conditionally essential panel, the absence of the metabolite highlighted in blue causes the highlighted reaction to become essential under the selected media. One of the network’s synthetic lethal pairs is denoted with “SL”. The dashed line in the flux variability illustration may represent ‘near optimal’ objective flux. A robust reaction maintains near optimal objective flux over a larger range of reaction fluxes.
References
- Segovia-Juarez JL, et al. Identifying control mechanisms of granuloma formation during M. tuberculosis infection using an agent-based model. J Theor Biol. 2004;231:357–376. -PubMed
Publication types
MeSH terms
Substances
Grants and funding
- R01 GM088244/GM/NIGMS NIH HHS/United States
- 5U54 AI057168/AI/NIAID NIH HHS/United States
- T32 AI007046/AI/NIAID NIH HHS/United States
- U54 AI057168/AI/NIAID NIH HHS/United States
- 5T32 AI0070465/AI/NIAID NIH HHS/United States
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous