Collective dynamics differentiates functional divergence in protein evolution - PubMed (original) (raw)
Collective dynamics differentiates functional divergence in protein evolution
Tyler J Glembo et al. PLoS Comput Biol. 2012.
Abstract
Protein evolution is most commonly studied by analyzing related protein sequences and generating ancestral sequences through Bayesian and Maximum Likelihood methods, and/or by resurrecting ancestral proteins in the lab and performing ligand binding studies to determine function. Structural and dynamic evolution have largely been left out of molecular evolution studies. Here we incorporate both structure and dynamics to elucidate the molecular principles behind the divergence in the evolutionary path of the steroid receptor proteins. We determine the likely structure of three evolutionarily diverged ancestral steroid receptor proteins using the Zipping and Assembly Method with FRODA (ZAMF). Our predictions are within ~2.7 Å all-atom RMSD of the respective crystal structures of the ancestral steroid receptors. Beyond static structure prediction, a particular feature of ZAMF is that it generates protein dynamics information. We investigate the differences in conformational dynamics of diverged proteins by obtaining the most collective motion through essential dynamics. Strikingly, our analysis shows that evolutionarily diverged proteins of the same family do not share the same dynamic subspace, while those sharing the same function are simultaneously clustered together and distant from those, that have functionally diverged. Dynamic analysis also enables those mutations that most affect dynamics to be identified. It correctly predicts all mutations (functional and permissive) necessary to evolve new function and ~60% of permissive mutations necessary to recover ancestral function.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
Figure 1. 3D structures of AncCR, AncGR1 and AncGR2.
AncCR was within 2.5 Å all-atom RMSD from the experimentally determined AncCR. AncGR1 was within 2.9 Å all-atom RMSD from the experimentally determined crystal structure. AncGR2 was within 2.8 Å all-atom RMSD from the experimentally determined AncGR2. Included for reference is a cartoon figure with helices labeled for reference and the ligand is bound, represented in blue spheres.
Figure 2. Plot and ribbon diagram of the dynamics of the three ancestral proteins characterized by slowest collective mode.
(A) The first two principal components of AncCR, AncGR1 and AncGR2 plotted against each other. The principal components were found via a Singular Value Decomposition of the G matrix (See Methods). Higher order modes are mostly orthogonal or mixed and therefore not represented here. (B) 3D structures of AncCR, AncGR1 and AncGR2 colored by residue fluctuation. The critical mutations in AncCR and AncGR1 have greater flexibility and thus, higher binding promiscuity. AncGR2 has much lower flexibility in general amongst these residues and therefore more selective binding. The S212Δ mutation also rigidifies the lower loop at the bottom end of h10 by shortening the loop and removing degrees of freedom. This also alters the packing of h10 (the frontmost helix) and decreases flexibility.
Figure 3. The change in fluctuation along the most collective mode between AncCR, AncGR1 and AncGR2.
The X, Y, Z, and Y27R mutation groups necessary to alter function toward cortisol binding specificity are noted in red, and those permissive W mutations necessary to reverse function and recover promiscuous binding are noted in purple. A cutoff of ±0.002 Å2 is applied to differentiate mutations critical to altering dynamics as also used in Fig. 4. The upper left region of the graph indicates mutations that most alter dynamics when comparing the function-altering mutation from AncGR1 (binding promiscuity) to AncGR2 (binding specificity to cortisol) whereas the lower right region of the plot indicates mutations that most alter dynamics when comparing AncCR and AncGR1, which do not diverge functionally.
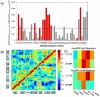
Figure 4. The change in net fluctuations and correlations of the mutated residues for successive evolution of MR to GR proteins.
(A) The change in net fluctuation between successive ancestral proteins, AncCR, AncGR1 and AncGR2 for mutated residues. Those residues identified as critical to alter-function are noted in red. The activation-function (AF) helix contains mutations 224 and 229. A cutoff (solid line) results in all critical mutations identified except for Y91C and L197M. Y27R is noted as critical to function but sites 65, 117, and 158 are false positives. (B) The cross correlation map with AncGR2 on the upper left and AncGR1 on the lower right. Circled in black are changes in the cross correlation associated with critical residues near the binding pocket. Squared in black are the changes in cross correlation due to critical mutation N26T forming a hydrogen bond with the AF-helix. Circles in white are additional changes in cross correlation not associated with critical mutations. (C) The cross correlations between the X and W mutations. The correlation between X and W mutations is higher for AncGR2, whereas AncGR1 X functional mutations are uncorrelated, increasing the flexibility in the binding pocket and allowing for promiscuous binding.
Figure 5. The secondary structure is predicted through multiple sequence alignment with modern day homologs.
These secondary structural elements are then connected with loops in extended conformation to generate hundreds of conformations with high flexibility. Only a few are shown here. These structures all undergo a FRODA simulation which collapses them by adding attractive perturbations between all hydrophobic contact pairs (represented by arrows) into tightly packed structures with hydrophobic cores. A subset of hydrophobic residues are shown as spheres. After scoring, the collapsed structures they are ran in a restrained r-REMD simulation for 5 ns and then an unrestrained REMD simulation for 5 ns or until converged. The 3 ancestral structures are prediction to within 2.7 Å all atom RMSD of a similar experimentally determined structure. The final ensemble of restraint free generated structures are analyzed for dynamics using PCA.
Similar articles
- Rewiring Ancient Residue Interaction Networks Drove the Evolution of Specificity in Steroid Receptors.
Okafor CD, Hercules D, Kell SA, Ortlund EA. Okafor CD, et al. Structure. 2020 Feb 4;28(2):196-205.e3. doi: 10.1016/j.str.2019.11.012. Epub 2019 Dec 9. Structure. 2020. PMID: 31831214 - Vestigialization of an allosteric switch: genetic and structural mechanisms for the evolution of constitutive activity in a steroid hormone receptor.
Bridgham JT, Keay J, Ortlund EA, Thornton JW. Bridgham JT, et al. PLoS Genet. 2014 Jan;10(1):e1004058. doi: 10.1371/journal.pgen.1004058. Epub 2014 Jan 9. PLoS Genet. 2014. PMID: 24415950 Free PMC article. - Biophysical mechanisms for large-effect mutations in the evolution of steroid hormone receptors.
Harms MJ, Eick GN, Goswami D, Colucci JK, Griffin PR, Ortlund EA, Thornton JW. Harms MJ, et al. Proc Natl Acad Sci U S A. 2013 Jul 9;110(28):11475-80. doi: 10.1073/pnas.1303930110. Epub 2013 Jun 24. Proc Natl Acad Sci U S A. 2013. PMID: 23798447 Free PMC article. - Analyzing protein structure and function using ancestral gene reconstruction.
Harms MJ, Thornton JW. Harms MJ, et al. Curr Opin Struct Biol. 2010 Jun;20(3):360-6. doi: 10.1016/j.sbi.2010.03.005. Epub 2010 Apr 21. Curr Opin Struct Biol. 2010. PMID: 20413295 Free PMC article. Review. - Structural divergence and distant relationships in proteins: evolution of the globins.
Lecomte JT, Vuletich DA, Lesk AM. Lecomte JT, et al. Curr Opin Struct Biol. 2005 Jun;15(3):290-301. doi: 10.1016/j.sbi.2005.05.008. Curr Opin Struct Biol. 2005. PMID: 15922591 Review.
Cited by
- The Role of Conformational Dynamics and Allostery in the Disease Development of Human Ferritin.
Kumar A, Glembo TJ, Ozkan SB. Kumar A, et al. Biophys J. 2015 Sep 15;109(6):1273-81. doi: 10.1016/j.bpj.2015.06.060. Epub 2015 Aug 6. Biophys J. 2015. PMID: 26255589 Free PMC article. - A hinge migration mechanism unlocks the evolution of green-to-red photoconversion in GFP-like proteins.
Kim H, Zou T, Modi C, Dörner K, Grunkemeyer TJ, Chen L, Fromme R, Matz MV, Ozkan SB, Wachter RM. Kim H, et al. Structure. 2015 Jan 6;23(1):34-43. doi: 10.1016/j.str.2014.11.011. Structure. 2015. PMID: 25565105 Free PMC article. - Integration of structural dynamics and molecular evolution via protein interaction networks: a new era in genomic medicine.
Kumar A, Butler BM, Kumar S, Ozkan SB. Kumar A, et al. Curr Opin Struct Biol. 2015 Dec;35:135-42. doi: 10.1016/j.sbi.2015.11.002. Epub 2015 Dec 9. Curr Opin Struct Biol. 2015. PMID: 26684487 Free PMC article. Review. - Dynamic coupling of residues within proteins as a mechanistic foundation of many enigmatic pathogenic missense variants.
Ose NJ, Butler BM, Kumar A, Kazan IC, Sanderford M, Kumar S, Ozkan SB. Ose NJ, et al. PLoS Comput Biol. 2022 Apr 7;18(4):e1010006. doi: 10.1371/journal.pcbi.1010006. eCollection 2022 Apr. PLoS Comput Biol. 2022. PMID: 35389981 Free PMC article. - Protein dynamics provide mechanistic insights about epistasis among common missense polymorphisms.
Ose NJ, Campitelli P, Patel R, Kumar S, Ozkan SB. Ose NJ, et al. Biophys J. 2023 Jul 25;122(14):2938-2947. doi: 10.1016/j.bpj.2023.01.037. Epub 2023 Feb 2. Biophys J. 2023. PMID: 36726312 Free PMC article.
References
- James LC, Tawfik DS. Conformational diversity and protein evolution - a 60-year-old hypothesis revisited. Trends Biochem Sci. 2003;28:361–368. - PubMed
- O'Brien PJ, Herschlag D. Catalytic promiscuity and the evolution of new enzymatic activities. Chem Biol. 1999;6:R97–R105. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous