Inefficient exogenous loading of a tapasin-dependent peptide onto HLA-B*44:02 can be improved by acid treatment or fixation of target cells - PubMed (original) (raw)
. 2012 Jun;42(6):1417-28.
doi: 10.1002/eji.201141954.
Nathalie Demotte, Rosalie M Luiten, Ralf M Leonhardt, Peter Cresswell, Aude Bonehill, Alexandre Michaux, Wenbin Ma, Arend Mulder, Benoît J Van den Eynde, Pierre van der Bruggen, Nathalie Vigneron
Affiliations
- PMID: 22678898
- PMCID: PMC3766947
- DOI: 10.1002/eji.201141954
Inefficient exogenous loading of a tapasin-dependent peptide onto HLA-B*44:02 can be improved by acid treatment or fixation of target cells
Vincent Stroobant et al. Eur J Immunol. 2012 Jun.
Abstract
Antitumor cytolytic T lymphocytes (CTLs) recognize peptides derived from cellular proteins and presented on MHC class I. One category of peptides recognized by these CTLs is derived from proteins encoded by "cancer-germline" genes, which are specifically expressed in tumors, and therefore represent optimal targets for cancer immunotherapy. Here, we identify an antigenic peptide, which is derived from the MAGE-A1-encoded protein (160-169) and presented to CTLs by HLA-B*44:02. Although this peptide is encoded by MAGE-A1, processed endogenously and presented by tumor cells, the corresponding synthetic peptide is hardly able to sensitize target cells to CTL recognition when pulsed exogenously. Endogenous processing and presentation of this peptide is strictly dependent on the presence of tapasin, which is believed to help peptide loading by stabilizing a peptide-receptive form of HLA-B*44:02. Exogenous loading of the peptide can be dramatically improved by paraformaldehyde fixation of surface molecules or by peptide loading at acidic pH. Either strategy allows efficient exogenous loading of the peptide, presumably by generating or stabilizing a peptide-receptive, empty conformation of the HLA. Altogether, our results indicate a potential drawback of short peptide-based vaccination strategies and offer possible solutions regarding the use of problematic epitopes such as the one described here.
© 2012 WILEY-VCH Verlag GmbH & Co. KGaA, Weinheim.
Conflict of interest statement
Conflict of interest
The Ludwig Institute for Cancer Research has filed a patent application on the peptide described in this manuscript.
Figures
Figure 1. CTL 4 recognizes a peptide derived from MAGE-A1 and restricted by HLA-B*44:02
(A) Lysis of melanoma (LB373-MEL) and EBV-B cells expressing MAGE-A1 by CTL 4 (at the indicated effector-target ratios) was measured by chromium release assay. Two hours after transduction of autologous EBV-B cells with the vaccinia constructs, cells were labeled with 51Cr and incubated with CTL 4. Chromium release was then measured after 4 hours. Data are representative of eight experiments. (B) The antigenic peptide recognized by CTL 4 is derived from MAGE-A1 and restricted to HLA-B*44:02. COS-7 cells were transfected with constructs encoding MAGE-A1 and the indicated HLA alleles. Transfected cells were then tested for their ability to stimulate TNF production by CTL 4. Data are shown as mean of three replicates and are representative of 3 experiments. (C) Representation of the MAGE-A1 coding sequence, and the minigenes used to identify the region coding for the antigenic peptide. These constructs were transfected in COS-7 together with the cDNA encoding HLA-B*44:02, and the cells were then tested for their ability to stimulate TNF production by CTL 4. Data are shown as mean of two replicates and are representative of 3 experiments. (D) CTL 4 does not recognize MAGE-A1 peptides after loading on LB1801-EBV. 51Cr-labeled target cells were loaded using 1 μM of the indicated peptide. After loading, CTL cells were added at an effector-to-target ratio of 10 to 1. As control, the CTL MZ2-82/30 was used to verify the integrity of the peptide solutions loaded on HLA-A*01:01+ LB1972-EBV. NT : not tested. Data are shown as mean of three replicates.
Figure 2. The peptide recognized by CTL 4 requires endogenous loading
(A) Activation of CTL 4 is improved when peptide KEADPTGHSY is pulsed on PFA-fixed target cells. B*44:02+ target cells – HLA-B*44:02+ 721.220 cells expressing tapasin (top) or autologous LB1801-EBV (bottom) – were fixed (black squares) or not (empty squares) prior to pulsing with the indicated concentration of peptide at 37°C for 1 hour. Cells were then washed and tested for their ability to activate CTL 4 in an IFN-γ release assay (ELISA). Data are the mean of two duplicates and are representative of three experiments. (B) Peptide KEADPTGHSY is present on the surface of cells expressing HLA-B*44:02 and MAGE-A1. Total peptides were eluted by mild acid treatment from B*44:02+ target cells (HLA-B* 44:02+ 721.220 cells expressing tapasin) transduced with a MAGE-A1 retroviral construct. The acid eluate was fractionated by HPLC, and the fractions were tested for recognition by CTL 4 after pulsing on PFA-fixed target cells, by IFN-γ production. To rule out contamination of the HPLC system, buffer was run on the column before the eluted sample, and the fractions were tested similarly (top). Synthetic peptide was injected under the same HPLC conditions, and the fractions were also tested for CTL recognition (bottom). Data are shown as mean of two replicates. (C) CTL activation is improved when peptide is loaded inside the target cells. HLA-B*44:02+ targets cells (HLA-B*44:02+ 721.220 cells expressing tapasin) were electroporated with the indicated peptides and tested for recognition by CTL 4 (black bars). As a control, peptide was pulsed in the same conditions, except that electroporation was omitted (white bars). NT : not tested. Data are shown as mean +/- SD of four replicates and are representative of two experiments. (D) Presentation of the peptide KEADPTGHSY requires expression of tapasin. HLA-B*44:02+ 721.220 cells expressing (or not) wild type tapasin (WT-TPN) were transfected with constructs encoding MAGE-A1 and cultured with the CTL 4. After 20 h, IFN-γ was measured in the supernatants. Data are shown as mean +/- SD of four replicates and are representative of 4 experiments.
Figure 3. Peptide KEADPTGHSY does not bind surface HLA-B*44:02 when pulsed exogenously
HLA-B*44:02+ target cells 721.220.B*44:02.tapasin expressing (or not) MAGE-A1 were pulsed (or not) with the relevant peptide. Recognition by CTL 4 prior to peptide elution was determined by IFN-γ production (left). Total peptides were acid-eluted from surface HLA molecules and purified on a C18 column. Concentrated (1:1, black bars) or two-fold diluted eluates (1:2, white bars) were then loaded onto PFA-fixed, HLA-B*44:02+ target cells for recognition by CTL 4 as measured by IFN-γ production (right). As a control, peptides were also eluted from HLA-B*44:02+ target cells, which were fixed with paraformaldehyde prior to pulsing of the peptide. Data are shown as mean +/- SD of four (left) and two (right) replicates and are representative of three experiments.
Figure 4. Conditions favoring peptide exchange on surface HLA-B*44:02
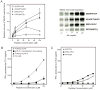
(A) In vitro peptide binding experiment. Tapasin-negative, HLA-B*44:02+ 721.220 cells were radio-labeled with 35S-methionine/cysteine for 30 min and lysed in 1% Triton-X100 in the presence of various concentration of the indicated peptides. As a negative control peptide RRYQNSTEL was used. Stabilized HLA molecules were immunoprecipitated using the W6/32 antibody. The samples were then separated by SDS-PAGE (right), and the intensity of the bands was analyzed using phospho-rimaging (left). The amount of W6/32-reactive complexes recovered was plotted as a percentage of that obtained with the highest concentration of peptide KEADPTGHSY. Data are shown as mean +/- SD of two experiments. (B) Low pH improves exogenous loading of the peptide on HLA-B*44:02. HLA-B*44:02+ 721.220.B*44:02.tapasin cells were incubated for 20 min at 37°C in the presence of the indicated concentration of peptide KEADPTGHSY at pH 5.5 or 7. After neutralization and washing, pulsed cells were tested for their ability to stimulate IFN-γ production by CTL 4. Additionally, cells were incubated at pH 5.5 for 20 min, neutralized then pulsed with peptide, washed and incubated with the CTL. Data are shown as mean +/- SD of two replicates and are representative of four experiments. (C) Cells expressing a lower affinity repertoire are more efficiently recognized by the CTL after pulsing. HLA-B*44:02+ 721.220 cells expressing WT or C95A tapasin mutant were pulsed with peptide KEADPTGHSY at pH 7 for 1h at 37°C, washed, and cultured with CTL 4. After 20 h, IFN-γ produced by the CTL was measured. Data are shown as mean +/- SD of two replicates and are representative of four experiments.
Figure 5. Competition for the endogenous loading of the MAGE-A1 peptide on HLA-A*01:01 (tapasin-independent) and HLA-B*44:02 (tapasin dependent)
(A) Presentation of the peptide EADPTGHSY does not require tapasin. HLA-A*01:01+ 721.220 cells expressing (or not) wild-type tapasin (WT-TPN) were transfected with constructs encoding MAGE-A1 and cultured with the HLA-A*01:01-restricted CTL MZ2-82/30. After 20 h, IFN-γ was measured in the supernatants. Data are shown as mean +/- SD of three replicates and are representative of 3 experiments. (B) Endogenous presentation of peptide (K)EADPTGHSY by HLA-A*01:01 is favored over HLA-B*44:02. HEK-293-EBNA were transiently transfected with HLA-B*44:02 and/or HLA-A*01:01 cDNA in the presence of the indicated amount of MAGE-A1 cDNA. The next day the HLA-B*44:02-restricted CTL 4 or the HLA-A*01:01-rectricted CTL MZ2-82/30 were added. After 20 h, IFN-γ was measured in the supernatants. Data are shown as mean of two replicates and are representative of two experiments.
Similar articles
- A new tumor-specific antigen encoded by MAGE-C2 and presented to cytolytic T lymphocytes by HLA-B44.
Godelaine D, Carrasco J, Brasseur F, Neyns B, Thielemans K, Boon T, Van Pel A. Godelaine D, et al. Cancer Immunol Immunother. 2007 Jun;56(6):753-9. doi: 10.1007/s00262-006-0244-5. Epub 2006 Nov 10. Cancer Immunol Immunother. 2007. PMID: 17096150 Free PMC article. - Differences in the recognition by CTL of peptides presented by the HLA-B*4402 and the HLA-B*4403 molecules which differ by a single amino acid.
Herman J, Jongeneel V, Kuznetsov D, Coulie PG. Herman J, et al. Tissue Antigens. 1999 Feb;53(2):111-21. doi: 10.1034/j.1399-0039.1999.530201.x. Tissue Antigens. 1999. PMID: 10090611 - A MAGE-A1 peptide presented to cytolytic T lymphocytes by both HLA-B35 and HLA-A1 molecules.
Luiten RM, Demotte N, Tine J, van der Bruggen P. Luiten RM, et al. Tissue Antigens. 2000 Jul;56(1):77-81. doi: 10.1034/j.1399-0039.2000.560110.x. Tissue Antigens. 2000. PMID: 10958359 - HER-2, gp100, and MAGE-1 are expressed in human glioblastoma and recognized by cytotoxic T cells.
Liu G, Ying H, Zeng G, Wheeler CJ, Black KL, Yu JS. Liu G, et al. Cancer Res. 2004 Jul 15;64(14):4980-6. doi: 10.1158/0008-5472.CAN-03-3504. Cancer Res. 2004. PMID: 15256472
Cited by
- Three tapasin docking sites in TAP cooperate to facilitate transporter stabilization and heterodimerization.
Leonhardt RM, Abrahimi P, Mitchell SM, Cresswell P. Leonhardt RM, et al. J Immunol. 2014 Mar 1;192(5):2480-94. doi: 10.4049/jimmunol.1302637. Epub 2014 Feb 5. J Immunol. 2014. PMID: 24501197 Free PMC article. - Therapeutic Cancer Vaccines-Antigen Discovery and Adjuvant Delivery Platforms.
Alarcon NO, Jaramillo M, Mansour HM, Sun B. Alarcon NO, et al. Pharmaceutics. 2022 Jul 11;14(7):1448. doi: 10.3390/pharmaceutics14071448. Pharmaceutics. 2022. PMID: 35890342 Free PMC article. Review. - Database of T cell-defined human tumor antigens: the 2013 update.
Vigneron N, Stroobant V, Van den Eynde BJ, van der Bruggen P. Vigneron N, et al. Cancer Immun. 2013 Jul 15;13:15. Print 2013. Cancer Immun. 2013. PMID: 23882160 Free PMC article. Review. - Cytosolic Processing Governs TAP-Independent Presentation of a Critical Melanoma Antigen.
Vigneron N, Ferrari V, Van den Eynde BJ, Cresswell P, Leonhardt RM. Vigneron N, et al. J Immunol. 2018 Oct 1;201(7):1875-1888. doi: 10.4049/jimmunol.1701479. Epub 2018 Aug 22. J Immunol. 2018. PMID: 30135181 Free PMC article. - Neoepitopes of Cancers: Looking Back, Looking Ahead.
Srivastava PK. Srivastava PK. Cancer Immunol Res. 2015 Sep;3(9):969-77. doi: 10.1158/2326-6066.CIR-15-0134. Cancer Immunol Res. 2015. PMID: 26342008 Free PMC article. Review.
References
- Kawakami Y, Eliyahu S, Jennings C, Sakaguchi K, Kang X, Southwood S, Robbins PF, et al. Recognition of multiple epitopes in the human melanoma antigen gp100 by tumor-infiltrating T lymphocytes associated with in vivo tumor regression. J Immunol. 1995;154:3961–3968. - PubMed
- Zarour H, De Smet C, Lehmann F, Marchand M, Lethé B, Romero P, Boon T, et al. The majority of autologous cytolytic T-lymphocyte clones derived from peripheral blood lymphocytes of a melanoma patient recognize an antigenic peptide derived from gene Pmel17/gp100. J Invest Dermatol. 1996;107:63–67. - PubMed
- Rock KL, Gramm C, Rothstein L, Clark K, Stein R, Dick L, Hwang D, et al. Inhibitors of the proteasome block the degradation of most cell proteins and the generation of peptides presented on MHC class I molecules. Cell. 1994;78:761–771. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials