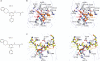A selective jumonji H3K27 demethylase inhibitor modulates the proinflammatory macrophage response - PubMed (original) (raw)
. 2012 Aug 16;488(7411):404-8.
doi: 10.1038/nature11262.
Chun-wa Chung, Zhongjun Cheng, John Liddle, KaHing Che, Gerard Joberty, Marcus Bantscheff, Chas Bountra, Angela Bridges, Hawa Diallo, Dirk Eberhard, Sue Hutchinson, Emma Jones, Roy Katso, Melanie Leveridge, Palwinder K Mander, Julie Mosley, Cesar Ramirez-Molina, Paul Rowland, Christopher J Schofield, Robert J Sheppard, Julia E Smith, Catherine Swales, Robert Tanner, Pamela Thomas, Anthony Tumber, Gerard Drewes, Udo Oppermann, Dinshaw J Patel, Kevin Lee, David M Wilson
Affiliations
- PMID: 22842901
- PMCID: PMC4691848
- DOI: 10.1038/nature11262
A selective jumonji H3K27 demethylase inhibitor modulates the proinflammatory macrophage response
Laurens Kruidenier et al. Nature. 2012.
Abstract
The jumonji (JMJ) family of histone demethylases are Fe2+- and α-ketoglutarate-dependent oxygenases that are essential components of regulatory transcriptional chromatin complexes. These enzymes demethylate lysine residues in histones in a methylation-state and sequence-specific context. Considerable effort has been devoted to gaining a mechanistic understanding of the roles of histone lysine demethylases in eukaryotic transcription, genome integrity and epigenetic inheritance, as well as in development, physiology and disease. However, because of the absence of any selective inhibitors, the relevance of the demethylase activity of JMJ enzymes in regulating cellular responses remains poorly understood. Here we present a structure-guided small-molecule and chemoproteomics approach to elucidating the functional role of the H3K27me3-specific demethylase subfamily (KDM6 subfamily members JMJD3 and UTX). The liganded structures of human and mouse JMJD3 provide novel insight into the specificity determinants for cofactor, substrate and inhibitor recognition by the KDM6 subfamily of demethylases. We exploited these structural features to generate the first small-molecule catalytic site inhibitor that is selective for the H3K27me3-specific JMJ subfamily. We demonstrate that this inhibitor binds in a novel manner and reduces lipopolysaccharide-induced proinflammatory cytokine production by human primary macrophages, a process that depends on both JMJD3 and UTX. Our results resolve the ambiguity associated with the catalytic function of H3K27-specific JMJs in regulating disease-relevant inflammatory responses and provide encouragement for designing small-molecule inhibitors to allow selective pharmacological intervention across the JMJ family.
Figures
Figure 1. Structure of the H3K27me3-derived peptide bound to mouse JMJD3
a, Domain topology and numbering system of JMJD3 construct used for crystallization of the enzyme in the free state. b, Overview of the structure of H3(20–34)K27me3–mouse JMJD3(1157–1641) complex, with the histone peptide (K27me3, yellow) in a stick representation and the engineered enzyme (Supplementary Fig. 2) colour coded by domain as in a: GATAL (green), JmjC (blue) and helical (pink) domains. The heteroatoms in the stick representation are coloured according to atom type: oxygen (red) and nitrogen (blue). The N and C termini of the enzyme are indicated. c, View of the intermolecular interactions between the bound histone peptide (yellow) and the enzyme residues (grey), with hydrogen bonds shown as dashed pink lines. d, View of the catalytic pocket of the enzyme (grey), the base of which is lined by NOG (pink) and Ni2+ (green). The histone peptide is shown in yellow. Details of the hydrogen bonding in the catalytic pocket are outlined in Supplementary Fig. 5a. e, Effect of mutations of the native H3K27me3 substrate peptide (highlighted in red) at R26 and P30 on turnover. Mass spectrometric quantification of the H3K27me2 product was performed after a 6-min reaction with concentrations of peptide spanning 0.6 to 250 μM (Supplementary Fig. 8). Data are shown at approximately the _K_m (Michaelis constant) of the standard, wild-type (WT), H3K27me3 peptide (22 μM) for clarity. Data are presented as the mean ± s.d.
Figure 2. Structure of the inhibitor GSK-J1 bound in the catalytic pocket of human JMJD3
a, Chemical structures of GSK-J1 and GSK-J2. b, Stereo view of the intermolecular interactions between the bound inhibitor GSK-J1 (orange) and the human JMJD3 enzyme residues (grey) that line the catalytic pocket, with hydrogen bonds shown as dashed pink lines. The heteroatoms in the stick representation are coloured according to atom type: oxygen (red) and nitrogen (blue). c, Stereo view of the superposition of the bound H3K27me3 peptide and the bound inhibitor GSK-J1 in their respective complexes with JMJD3. For clarity, the histone peptide (yellow), NOG (pink), Ni2+ (green) and certain JMJD3 side chains (grey) are shown only from the peptide complex, and GSK-J1 (orange) and Co2+ (blue) are shown from the inhibitor complex.
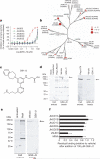
Figure 3. GSK-J1 is selective for H3K27 demethylases of the KDM6 subfamily and specifically binds to endogenous JMJD3
a, Evaluation of the selectivity of GSK-J1 in JMJ mass spectrometric assays. Data are presented as the mean ± s.d. b, Phylogenetic tree of human JMJ enzymes illustrating the selectivity of GSK-J1 for demethylases of the KDM6 subfamily over other KDM subfamilies of methyl-lysine demethylases, as determined by melting point (_T_m) shift screening. Temperature differences (°C) are shown in red circles. Only UTX and JMJD3 show significant stabilization (>1 °C), indicating ligand binding. c–f, Chemoproteomics studies show target engagement in a cellular environment. c, GSK-J3, an amine analogue of GSK-J1 used for bead immobilization. d, Immobilized GSK-J3 was able to affinity-capture Flag-tagged JMJD3 and Flag-tagged UTX from transiently transfected HEK-293 cells. Western blotting analysis with an anti-Flag antibody shows the capture of JMJD3 (left, arrow) and UTX (right, arrow), both of which were competitively inhibited by the addition of 100 μM GSK-J1. e, PMA treatment of HL-60 cells (80 nM for 16 h) induced the expression of JMJD3, as shown by western blotting. Immobilized GSK-J3 captured the endogenous JMJD3 protein (arrow), and this capture was competitively inhibited by 100 μM GSK-J1. f, GSK-J1 is selective for JMJD3, as shown by mass-spectrometry-based quantification of JMJC proteins (captured with GSK-J3 beads) from PMA-stimulated HL-60 cells in the presence of 100 μM GSK-J1. Data are presented as the mean from two experiments, and error bars represent the 95% confidence intervals.
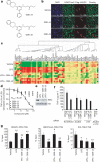
Figure 4. GSK-J1 inhibits TNF-α production by human primary macrophages in an H3K27-dependent manner
a, Chemical structures of the ethyl ester pro-drugs GSK-J4 and GSK-J5. b, Administration of 25 μM GSK-J4, but not GSK-J5, preserved nuclear H3K27me3 staining (green) in Flag–JMJD3-transfected (red, arrows) HeLa cells. Scale bars, 50 μm. c, Heat map representation of cytokine expression by human primary macrophages activated with LPS (for 2 h) in the presence of 30 μM GSK-J4 or GSK-J5 (n = 4 donors; green, low expression; red, high expression). d, TNF-α production by human primary macrophages activated with LPS in the presence of the indicated concentrations of GSK-J4 or GSK-J5 for 6 h (data are presented as the mean ± s.e.m. from n = 5 donors). e, Western blot showing the degradation of inhibitor of NF-κB α (IκBα) in whole cell lysates of LPS-stimulated human primary macrophages (30 min) in the presence of 30 μM GSK-J4 or GSK-J5. f, TNF-α production by LPS-stimulated human primary macrophages (1 h) transfected with scrambled, or JMJD3-directed or UTX-directed siRNA, in the presence of 30 μM GSK-J4 or GSK-J5 (data are presented as the mean ± s.d. from five transfection replicates from one representative experiment). g, ChIP analysis of the association of RNA polymerase II (RNAPII), H3K27me3 and total H3 (tH3) with the TNFA TSS in LPS-stimulated human primary macrophages (1 h) in the presence of 30 μM GSK-J4 or GSK-J5 (data are presented as the mean ± s.e.m. from n = 8 donors). P values were calculated using a two-tailed Student's _t_-test. *, P < 0.05 compared with vehicle; **, P < 0.01 compared with vehicle; #, P < 0.05 compared with LPS; ##, P < 0.01 compared with LPS.
Comment in
- Structure-based drug design: Opening the door to an epigenetic target.
Harrison C. Harrison C. Nat Rev Drug Discov. 2012 Sep;11(9):672. doi: 10.1038/nrd3827. Epub 2012 Aug 20. Nat Rev Drug Discov. 2012. PMID: 22903043 No abstract available. - Inhibition of demethylases by GSK-J1/J4.
Heinemann B, Nielsen JM, Hudlebusch HR, Lees MJ, Larsen DV, Boesen T, Labelle M, Gerlach LO, Birk P, Helin K. Heinemann B, et al. Nature. 2014 Oct 2;514(7520):E1-2. doi: 10.1038/nature13688. Nature. 2014. PMID: 25279926 No abstract available. - Kruidenier et al. reply.
Kruidenier L, Chung CW, Cheng Z, Liddle J, Che K, Joberty G, Bantscheff M, Bountra C, Bridges A, Diallo H, Eberhard D, Hutchinson S, Jones E, Katso R, Leveridge M, Mander PK, Mosley J, Ramirez-Molina C, Rowland P, Schofield CJ, Sheppard RJ, Smith JE, Swales C, Tanner R, Thomas P, Tumber A, Drewes G, Oppermann U, Patel DJ, Lee K, Wilson DM. Kruidenier L, et al. Nature. 2014 Oct 2;514(7520):E2. doi: 10.1038/nature13689. Nature. 2014. PMID: 25279927 No abstract available.
Similar articles
- Structure of the Arabidopsis JMJ14-H3K4me3 Complex Provides Insight into the Substrate Specificity of KDM5 Subfamily Histone Demethylases.
Yang Z, Qiu Q, Chen W, Jia B, Chen X, Hu H, He K, Deng X, Li S, Tao WA, Cao X, Du J. Yang Z, et al. Plant Cell. 2018 Jan;30(1):167-177. doi: 10.1105/tpc.17.00666. Epub 2017 Dec 12. Plant Cell. 2018. PMID: 29233856 Free PMC article. - Identification of JmjC domain-containing UTX and JMJD3 as histone H3 lysine 27 demethylases.
Hong S, Cho YW, Yu LR, Yu H, Veenstra TD, Ge K. Hong S, et al. Proc Natl Acad Sci U S A. 2007 Nov 20;104(47):18439-44. doi: 10.1073/pnas.0707292104. Epub 2007 Nov 14. Proc Natl Acad Sci U S A. 2007. PMID: 18003914 Free PMC article. - Design and discovery of new pyrimidine coupled nitrogen aromatic rings as chelating groups of JMJD3 inhibitors.
Hu J, Wang X, Chen L, Huang M, Tang W, Zuo J, Liu YC, Shi Z, Liu R, Shen J, Xiong B. Hu J, et al. Bioorg Med Chem Lett. 2016 Feb 1;26(3):721-725. doi: 10.1016/j.bmcl.2016.01.006. Epub 2016 Jan 7. Bioorg Med Chem Lett. 2016. PMID: 26776360 - Structural definitions of Jumonji family demethylase selectivity.
Pilka ES, James T, Lisztwan JH. Pilka ES, et al. Drug Discov Today. 2015 Jun;20(6):743-9. doi: 10.1016/j.drudis.2014.12.013. Epub 2014 Dec 31. Drug Discov Today. 2015. PMID: 25555749 Review. - Structural insights into histone lysine demethylation.
Hou H, Yu H. Hou H, et al. Curr Opin Struct Biol. 2010 Dec;20(6):739-48. doi: 10.1016/j.sbi.2010.09.006. Epub 2010 Oct 21. Curr Opin Struct Biol. 2010. PMID: 20970991 Free PMC article. Review.
Cited by
- Mechanisms of epigenetic regulation of leukemia onset and progression.
Ntziachristos P, Mullenders J, Trimarchi T, Aifantis I. Ntziachristos P, et al. Adv Immunol. 2013;117:1-38. doi: 10.1016/B978-0-12-410524-9.00001-3. Adv Immunol. 2013. PMID: 23611284 Free PMC article. Review. - Sustained antidepressant effects of ketamine metabolite involve GABAergic inhibition-mediated molecular dynamics in aPVT glutamatergic neurons.
Kawatake-Kuno A, Li H, Inaba H, Hikosaka M, Ishimori E, Ueki T, Garkun Y, Morishita H, Narumiya S, Oishi N, Ohtsuki G, Murai T, Uchida S. Kawatake-Kuno A, et al. Neuron. 2024 Apr 17;112(8):1265-1285.e10. doi: 10.1016/j.neuron.2024.01.023. Epub 2024 Feb 19. Neuron. 2024. PMID: 38377990 - Epigenetics: the fine-tuner in inflammatory bowel disease?
Stylianou E. Stylianou E. Curr Opin Gastroenterol. 2013 Jul;29(4):370-7. doi: 10.1097/MOG.0b013e328360bd12. Curr Opin Gastroenterol. 2013. PMID: 23743674 Free PMC article. Review. - Inhibition of H3K27me3-specific histone demethylases JMJD3 and UTX blocks reactivation of herpes simplex virus 1 in trigeminal ganglion neurons.
Messer HG, Jacobs D, Dhummakupt A, Bloom DC. Messer HG, et al. J Virol. 2015 Mar;89(6):3417-20. doi: 10.1128/JVI.03052-14. Epub 2014 Dec 31. J Virol. 2015. PMID: 25552720 Free PMC article. - Epigenetic Control of Macrophage Polarisation and Soluble Mediator Gene Expression during Inflammation.
Kapellos TS, Iqbal AJ. Kapellos TS, et al. Mediators Inflamm. 2016;2016:6591703. doi: 10.1155/2016/6591703. Epub 2016 Apr 10. Mediators Inflamm. 2016. PMID: 27143818 Free PMC article. Review.
References
- Mosammaparast N, Shi Y. Reversal of histone methylation: biochemical and molecular mechanisms of histone demethylases. Annu. Rev. Biochem. 2010;79:155–179. - PubMed
- Klose RJ, Zhang Y. Regulation of histone methylation by demethylimination and demethylation. Nature Rev. Mol. Cell Biol. 2007;8:307–318. - PubMed
- Shi Y. Histone lysine demethylases: emerging roles in development, physiology and disease. Nature Rev. Genet. 2007;8:829–833. - PubMed
- Trojer P, Reinberg D. Histone lysine demethylases and their impact on epigenetics. Cell. 2006;125:213–217. - PubMed
- Loenarz C, Schofield CJ. Physiological and biochemical aspects of hydroxylations and demethylations catalyzed by human 2-oxoglutarate oxygenases. Trends Biochem. Sci. 2011;36:7–18. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- 18358/VAC_/Versus Arthritis/United Kingdom
- P30 CA008748/CA/NCI NIH HHS/United States
- 092809/WT_/Wellcome Trust/United Kingdom
- WT_/Wellcome Trust/United Kingdom
- 18358/ARC_/Arthritis Research UK/United Kingdom
- CAPMC/ CIHR/Canada
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases