Evidence supporting a zoonotic origin of human coronavirus strain NL63 - PubMed (original) (raw)
. 2012 Dec;86(23):12816-25.
doi: 10.1128/JVI.00906-12. Epub 2012 Sep 19.
Shimena Li, Boyd Yount, Alexander Smith, Leslie Sturges, John C Olsen, Juliet Nagel, Joshua B Johnson, Sudhakar Agnihothram, J Edward Gates, Matthew B Frieman, Ralph S Baric, Eric F Donaldson
Affiliations
- PMID: 22993147
- PMCID: PMC3497669
- DOI: 10.1128/JVI.00906-12
Evidence supporting a zoonotic origin of human coronavirus strain NL63
Jeremy Huynh et al. J Virol. 2012 Dec.
Abstract
The relationship between bats and coronaviruses (CoVs) has received considerable attention since the severe acute respiratory syndrome (SARS)-like CoV was identified in the Chinese horseshoe bat (Rhinolophidae) in 2005. Since then, several bats throughout the world have been shown to shed CoV sequences, and presumably CoVs, in the feces; however, no bat CoVs have been isolated from nature. Moreover, there are very few bat cell lines or reagents available for investigating CoV replication in bat cells or for isolating bat CoVs adapted to specific bat species. Here, we show by molecular clock analysis that alphacoronavirus (α-CoV) sequences derived from the North American tricolored bat (Perimyotis subflavus) are predicted to share common ancestry with human CoV (HCoV)-NL63, with the most recent common ancestor between these viruses occurring approximately 563 to 822 years ago. Further, we developed immortalized bat cell lines from the lungs of this bat species to determine if these cells were capable of supporting infection with HCoVs. While SARS-CoV, mouse-adapted SARS-CoV (MA15), and chimeric SARS-CoVs bearing the spike genes of early human strains replicated inefficiently, HCoV-NL63 replicated for multiple passages in the immortalized lung cells from this bat species. These observations support the hypothesis that human CoVs are capable of establishing zoonotic-reverse zoonotic transmission cycles that may allow some CoVs to readily circulate and exchange genetic material between strains found in bats and other mammals, including humans.
Figures
Fig 1
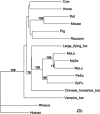
Phylogeny of the mitochondrial cytochrome b gene from several mammalian species. This maximum likelihood tree shows that North American bats of the family Vespertilionidae are more closely related to themselves than to other bat or common mammalian species. EpFu, big brown bat; PeSu, tricolored bat; MyLu, little brown myotis; MyLe, eastern small-footed myotis; MySe, northern long-eared myotis. The scale bar represents nucleotide substitutions. Only nodes with bootstrap support above 70% are labeled.
Fig 2
A novel α-CoV in the tricolored bat is closely related to HCoV-NL63. Novel α-CoV sequences have been found in the fecal samples of several North American bat species. (A) Schematic showing where the two fragments (used for the trees in panels B and C) occur in the NL63 genome. (B) A >2.2-kb fragment of the replicase region (starting at position 16480 in the schematic) was sequenced from three samples, including two from the big brown bat (NECoV and ARCoV.1) and one from the tricolored bat (ARCoV.2). A maximum likelihood tree comparing the nucleotide sequences of these bat CoVs to other known CoVs indicated that NECoV and ARCoV.1 are very closely related while ARCoV.2 is significantly different from NECoV and ARCoV.1. The three novel α-CoV sequences form a novel cluster in the α-CoV group. (C) Molecular clock analysis using a 650- to 800-nt portion of the highly conserved replicase region (starting at position 13810 in panel A) predicted that the MRCA of bat CoVs from the Hipposideros caffer ruber bats and HCoV-229E was likely to have existed 212 to 350 years ago (in agreement with Pfefferle et al., 2009 [36]). The MRCA for HCoV-NL63 and ARCoV.2 was predicted to have existed 563 to 822 years ago. NECOV and ARCoV.1 were identical in this region, so only ARCoV.1 is shown, and it clustered with other bat α-CoVs.
Fig 3
Immortalization of tricolored bat lung cells, PESU-B5L. Primary tricolored bat lung cells were grown in enriched growth medium and passaged four times to generate a suitable quantity for immortalization. Primary cells were immortalized with lentiviral vectors containing hTERT and the Bmi-1 proto-oncogene. (A) Primary tricolored bat lung cells. (B) Primary PESU-B5L lung cells reached senescence by passage 10. (C) Immortalized cells at passage 10, including 4 passages as primary cells and 6 passages postimmortalization. The immortalized PESU-B5L cells were still going strong after more than 30 passages (over 100 doublings).
Fig 4
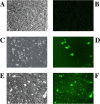
Infection of PESU-B5L cells with human coronavirus NL63 expressing GFP. PESU-B5L cells were infected with mock or NL63gfp at an MOI of 0.25 and monitored by bright-field and fluorescence microscopy. (A and B) Mock-infected cells by bright-field (A) and fluorescence (B) microscopy. (C through F) Cells infected with NL63gfp visualized at day 3 postinfection by bright-field microscopy (C) and by fluorescence microscopy (D) and at day 5 postinfection by bright-field microscopy (E) and by fluorescence microscopy (F).
Fig 5
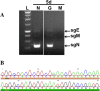
Verification of NL63 replication in PESU-B5L cells. Total RNA was harvested from cells on day 5 postinfection and subjected to RT-PCR using primers designed to detect leader-containing transcripts of the N gene, which would be present only if NL63 replicated. (A) Subgenomic transcripts were detected in the PESU-B5L cells infected with both wtNL63 and NL63gfp. (B) The sgN band for wtNL63 was excised, and the DNA was purified and sequenced to verify that the HCoV-NL63 leader sequence and the 5′ end of the N gene were present. L, ladder; N, wt-NL63; G, NL63gfp; M, mock. The leader sequence has a beige bar beneath it, and the 5′ end of the N gene has a green bar beneath it.
Fig 6
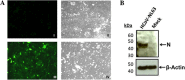
Evidence of HCoV-NL63 infection in PESU-B5L cells. PESU-B5L cells were infected with HCoV-NL63 at an MOI of 0.8. At 72 h postinfection, cells were probed for the nucleocapsid protein of HCoV-NL63 by immunofluorescence assay and Western blotting. (A) I, mock-infected cells, fluorescence microscopy; II, mock-infected cells, bright-field microscopy; III, HCoV-NL63-infected cells, fluorescence microscopy; IV, HCoV-NL63-infected cells, bright-field microscopy. (B) Western blot of HCoV-NL63 and mock-infected PESU-B5L cell lysates with β-actin as a loading control. Nucleocapsid (N) protein appears as a distinct band at ∼45 kDa.
Fig 7
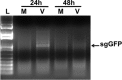
Evidence of SARS-CoV replication in PESU-B5L cells. PESU-B5L cells were infected with SARS-CoV at an MOI of ∼1, and total RNA was harvested from the cells at 24-h intervals and subjected to RT-PCR using primers designed to detect leader-containing transcripts of the subgenomic GFP (sgGFP), indicative of SARS-CoV replication. Subgenomic transcripts were detected at 24 h postinfection but disappeared by 48 h postinfection, suggesting that SARS-CoV replicates in these cells at a low level. L, Promega 1-kb ladder; M, mock-infected cells; V, SARS-CoV-infected cells.
Fig 8
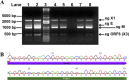
Evidence of SARS-CoV replication in PESU-B5L cells. PESU-B5L cells were infected with the mouse-adapted strain of SARS-CoV, MA15, and with chimeric SARS-CoV viruses bearing the spike genes of GD03 and HC/SZ/61/03 at an MOI of ∼1. Total RNA was harvested from the cells at 48 and 72 h postinfection and subjected to RT-PCR using primers designed to detect leader-containing subgenomic transcripts indicative of replication. (A) Subgenomic (sg) transcripts for X1, envelope (E), membrane (M), and open reading frame 6 [ORF6 (X3)] were detected at both time points for all three strains. Lane 1, MA15 at 48 h; lane 2, MA15 at 72 h; lane 3, Invitrogen 1-kb Plus ladder; lane 4, SARS+GD03 at 48 h; lane 5, SARS+GD03 at 72 h; lane 6, mock-infected cells; lane 7, SARS+HC/SZ/61/03 at 48 h; and lane 8, SARS+HC/SZ/61/03 at 72 h. (B) The sg M bands for all three strains were excised, and the DNA was purified and sequenced to verify that the SARS-CoV leader sequence and the 5′ end of the M gene were present. Shown here is sg M from SARS-CoV strain GD03.
Similar articles
- Surveillance of Bat Coronaviruses in Kenya Identifies Relatives of Human Coronaviruses NL63 and 229E and Their Recombination History.
Tao Y, Shi M, Chommanard C, Queen K, Zhang J, Markotter W, Kuzmin IV, Holmes EC, Tong S. Tao Y, et al. J Virol. 2017 Feb 14;91(5):e01953-16. doi: 10.1128/JVI.01953-16. Print 2017 Mar 1. J Virol. 2017. PMID: 28077633 Free PMC article. - Evidence for an Ancestral Association of Human Coronavirus 229E with Bats.
Corman VM, Baldwin HJ, Tateno AF, Zerbinati RM, Annan A, Owusu M, Nkrumah EE, Maganga GD, Oppong S, Adu-Sarkodie Y, Vallo P, da Silva Filho LV, Leroy EM, Thiel V, van der Hoek L, Poon LL, Tschapka M, Drosten C, Drexler JF. Corman VM, et al. J Virol. 2015 Dec;89(23):11858-70. doi: 10.1128/JVI.01755-15. Epub 2015 Sep 16. J Virol. 2015. PMID: 26378164 Free PMC article. - Severe Acute Respiratory Syndrome (SARS) Coronavirus ORF8 Protein Is Acquired from SARS-Related Coronavirus from Greater Horseshoe Bats through Recombination.
Lau SK, Feng Y, Chen H, Luk HK, Yang WH, Li KS, Zhang YZ, Huang Y, Song ZZ, Chow WN, Fan RY, Ahmed SS, Yeung HC, Lam CS, Cai JP, Wong SS, Chan JF, Yuen KY, Zhang HL, Woo PC. Lau SK, et al. J Virol. 2015 Oct;89(20):10532-47. doi: 10.1128/JVI.01048-15. Epub 2015 Aug 12. J Virol. 2015. PMID: 26269185 Free PMC article. - Ecology, evolution and classification of bat coronaviruses in the aftermath of SARS.
Drexler JF, Corman VM, Drosten C. Drexler JF, et al. Antiviral Res. 2014 Jan;101:45-56. doi: 10.1016/j.antiviral.2013.10.013. Epub 2013 Oct 31. Antiviral Res. 2014. PMID: 24184128 Free PMC article. Review. - The zoonotic potential of bat-borne coronaviruses.
Ravelomanantsoa NAF, Guth S, Andrianiaina A, Andry S, Gentles A, Ranaivoson HC, Brook CE. Ravelomanantsoa NAF, et al. Emerg Top Life Sci. 2020 Dec 11;4(4):353-369. doi: 10.1042/ETLS20200097. Emerg Top Life Sci. 2020. PMID: 33258903 Review.
Cited by
- Human coronavirus EMC does not require the SARS-coronavirus receptor and maintains broad replicative capability in mammalian cell lines.
Müller MA, Raj VS, Muth D, Meyer B, Kallies S, Smits SL, Wollny R, Bestebroer TM, Specht S, Suliman T, Zimmermann K, Binger T, Eckerle I, Tschapka M, Zaki AM, Osterhaus AD, Fouchier RA, Haagmans BL, Drosten C. Müller MA, et al. mBio. 2012 Dec 11;3(6):e00515-12. doi: 10.1128/mBio.00515-12. mBio. 2012. PMID: 23232719 Free PMC article. - Structural Requirements and Plasticity of Receptor-Binding Domain in Human Coronavirus Spike.
Li Y, Zheng P, Liu T, Shi C, Wang B, Xu Y, Jin T. Li Y, et al. Front Mol Biosci. 2022 Jul 12;9:930931. doi: 10.3389/fmolb.2022.930931. eCollection 2022. Front Mol Biosci. 2022. PMID: 35903152 Free PMC article. Review. - Stability of SARS-CoV-2 in Biological Fluids of Animals.
Kwon T, Gaudreault NN, Cool K, McDowell CD, Morozov I, Richt JA. Kwon T, et al. Viruses. 2023 Mar 16;15(3):761. doi: 10.3390/v15030761. Viruses. 2023. PMID: 36992470 Free PMC article. - Predicting the animal hosts of coronaviruses from compositional biases of spike protein and whole genome sequences through machine learning.
Brierley L, Fowler A. Brierley L, et al. PLoS Pathog. 2021 Apr 20;17(4):e1009149. doi: 10.1371/journal.ppat.1009149. eCollection 2021 Apr. PLoS Pathog. 2021. PMID: 33878118 Free PMC article. - Molecular optimization, docking, and dynamic simulation profiling of selective aromatic phytochemical ligands in blocking the SARS-CoV-2 S protein attachment to ACE2 receptor: an in silico approach of targeted drug designing.
Dey D, Paul PK, Al Azad S, Al Mazid MF, Khan AM, Sharif MA, Rahman MH. Dey D, et al. J Adv Vet Anim Res. 2021 Mar 5;8(1):24-35. doi: 10.5455/javar.2021.h481. eCollection 2021 Mar. J Adv Vet Anim Res. 2021. PMID: 33860009 Free PMC article.
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous