COX5B regulates MAVS-mediated antiviral signaling through interaction with ATG5 and repressing ROS production - PubMed (original) (raw)
COX5B regulates MAVS-mediated antiviral signaling through interaction with ATG5 and repressing ROS production
Yuanyuan Zhao et al. PLoS Pathog. 2012 Dec.
Abstract
Innate antiviral immunity is the first line of the host defense system that rapidly detects invading viruses. Mitochondria function as platforms for innate antiviral signal transduction in mammals through the adaptor protein, MAVS. Excessive activation of MAVS-mediated antiviral signaling leads to dysfunction of mitochondria and cell apoptosis that likely causes the pathogenesis of autoimmunity. However, the mechanism of how MAVS is regulated at mitochondria remains unknown. Here we show that the Cytochrome c Oxidase (CcO) complex subunit COX5B physically interacts with MAVS and negatively regulates the MAVS-mediated antiviral pathway. Mechanistically, we find that while activation of MAVS leads to increased ROS production and COX5B expression, COX5B down-regulated MAVS signaling by repressing ROS production. Importantly, our study reveals that COX5B coordinates with the autophagy pathway to control MAVS aggregation, thereby balancing the antiviral signaling activity. Thus, our study provides novel insights into the link between mitochondrial electron transport system and the autophagy pathway in regulating innate antiviral immunity.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
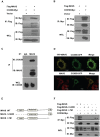
Figure 1. COX5B interacts with MAVS.
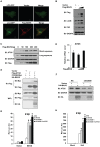
(A and B) HEK293 cells were transfected with combinations of DNA constructs as indicated. Twenty-four hours after transfection, cell lysates were prepared, immunoprecipitated with anti-Flag beads (A) or with anti-Myc antibody (B), followed by immunoblot analysis with the indicated antibodies. WCL (bottom), expression of transfected proteins in whole-cell lysates. (C) HEK293 cell lysates were prepared, immunoprecipitated with anti-MAVS antibody or control IgG, followed by immunoblot analysis. (D) Hela cells were transfected with COX5B-GFP and control vector or YFP-MAVS for 36 h. Cells were fixed, then stained with anti-MAVS antibody (Bottom) or mounted onto slides directly (Top), and imaged by confocal microscopy. (E) Schematic diagram of MAVS and truncated mutants. (F) HEK293 cells were transfected with the indicated plasmids, cell lysates were immunoprecipitated with anti-Flag beads, followed by immunoblot analysis.
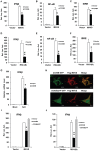
Figure 2. Overexpression of COX5B suppresses MAVS-mediated antiviral signaling.
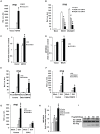
(A–C) MAVS and COX5B, or empty expression vectors were transfected in HEK293 cells together with luciferase reporter constructs driven by promoters of genes encoding IFNβ (A), NF-κB (B) or ISRE(C), as well as pRSV/LacZ as an internal control. Twenty-four hours after transfection, the luciferase activity was measured and normalized based on β-galactosidase activity. Results are presented relative to the luciferase activity in control cells. (D–F) RIG-I(N) and COX5B or empty expression vectors were transfected in HEK293 cells together with IFNβ-luc (D), NF-κB-luc (E) or IRSE-luc (F) along with pRSV/LacZ. Subsequently, cells were lysed for luciferase assays. (G) HEK293 cells were transfected with empty expression vector or COX5B for 24 h. The cells were then untreated or infected with Sendai virus (50HA units/ml) for 12 h, total RNA was isolated to check the expression of IFNβ mRNA by real-time PCR. (H) Hela cells were transfected with COX5B-GFP or COX5BΔTP-GFP and Flag-MAVS, the cells were then stained with the anti-Flag antibody and imaged by confocal microscopy. (I) HEK293 cells were transfected with the indicated constructs together with IFNβ reporter plasmids. Cells were lysed for luciferase assays after 24 h transfection. (J) HEK293 cells were transfected similarly as in (I), except that RIG-I(N) construct was used instead of MAVS. Data from A–G, I and J are representative of at least three independent experiments (mean and s.d. of duplicate or triplicate assays). *, P<0.05; **, P<0.01 versus the control groups.
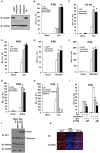
Figure 3. COX5B negatively regulates MAVS-mediated antiviral signaling.
(A) Oligos targeting three different regions of COX5B, NC (control) or siGFP (control) were transfected into HEK293 cells, 48 h after transfection, cell lysates were analyzed by immunoblotting with the indicated antibodies. (B–D) HEK293 cells were first transfected with NC, GFP or COX5B RNAi oligos (either NC or GFP RNAi as negative control). After 48 h transfection, IFNβ (B), NF-κB (C), or ISRE (D) reporter plasmids were transfected into the RNAi cells respectively, followed by the Sendai virus (50HA unit/ml) infection for 16 h, and subsequently cells were lysed for luciferase assays. (E–G) HEK293 cells were transfected with GFP or COX5B RNAi oligos. After 48 h transfection, and the indicated expression vectors RIG-I(N) (E), MDA5(N) (F), or MAVS (G) along with reporter plasmids were transfected into the RNAi cells, followed by luciferase assay after the second transfection for 24 h. (H) HEK293 cells were transfected with 20 nM oligos as indicated, NC oligos were used to balance the equal amount of RNAi oligos, second transfection and virus infection were performed as in (B). (I) HEK293 cells were first transfected with COX5B 3′UTR RNAi oligo, 12 h after transfection, reporter plasmids and the increasing amounts of COX5B expression vectors were then transfected into the RNAi cells, 24 h after the second transfection, the cells were then infected with Sendai virus (50HA unit/ml) for 16 h. Finally, cells were lysed for luciferase assays. (J–K) Knockdown of COX5B enhances dimerization and nuclear translocation of IRF3 induced by Sendai virus infection. (J) HEK293 cells were transfected with COX5B RNAi or control oligos. Forty-eight hours after transfection, cells were infected with SeV or left uninfected for 10 h. Cell lysates were resolved by native gel electrophoresis (upper panel) or SDS-PAGE (lower panels) and analyzed with the indicated antibodies. (K) HEK293 cells were transfected with COX5B RNAi oligos as in (J), subsequently cells were infected with SeV for 9 h, stained with an IRF3 antibody and DAPI, then imaged by confocal microscopy. Data from B–I are representative of at least three independent experiments (mean and s.d. of duplicate assays). *, P<0.05; **, P<0.01 versus the control groups.
Figure 4. COX5B negatively controls the virus amplification.
(A–C) HEK293 cells were transfected with COX5B RNAi oligos or NC. After 48 h transfection, cells were infected by Sendai virus for 12 h, and then lysed to isolate RNA to check the transcription levels of IFNβ (A), RANTES (B) and Viperin (C) by real-time PCR. (D–F) The transfection were performed as in (A–C), except that cells were infected by VSVΔM51-GFP (MOI = 0.1) for 9 h. (G) The RNAi oligo transfection was carried out as in (A), RNAi cells were infected by Sendai virus for 16 h, and the supernatant was collected for measurement of IFNβ by ELISA. (H–J) HEK293 cells were transfected with COX5B RNAi oligos for 48 h, then cells were infected by VSV-GFP (H) or VSVΔM51-GFP (I) at the MOI of 0.01 for 12 h, subsequently the culture supernatants were collected to measure virus titer by plaque assay, or cells were imaged by fluorescence microscopy (J). Data from A–I are representative of at least three independent experiments, (A–F, mean and s.d. of triplicate assays, G–I using duplicate assays). *, P<0.05; **, P<0.01 versus the control groups.
Figure 5. COX5B mediates MAVS signaling by repressing ROS production.
(A) HEK293 cells were pretreated with 10 ug/ml antimycin A for 2 h, then transfected with the indicated plasmids. Twenty-four hours after transfection, cells were lysed to measure the IFNβ induction. (B) HEK293 cells were pretreated with 250 or 500 µM Mito-TEMPO for 4 h, and then transfected with IFNβ reporter and pRSV/LacZ vectors together with empty vector or MAVS, or infected with VSVΔM51-GFP (MOI = 0.1). Cells were lysed to measure IFNβ activity after 24 h transfection or 10 h infection. (C and D) COX5B RNAi oligos or NC were transfected into HEK293 cells, after 20 h transfection, empty vector or MAVS plasmids were transfected again, 30 h after the second transfection, cells were collected for FACS analysis to check cellular or mitochondrial ROS production by staining with DCF (C) or MitoSOX (D), respectively. Results are presented relative to the FACS mean fluorescence intensity over control cells. (E) HEK293 cells were transfected with COX5B RNAi oligo or NC, 36 h after transfection, cells were pretreated with 250 µM Mito-TEMPO for 4 h, and transfected again with indicated plasmids. Twenty-four hours after second transfection, cells were lysed to measure the IFNβ induction. (F) After first transfection and treatment as described in (E), cells were transfected with IFNβ reporter and pRSV/LacZ vectors, followed by 16 h Sendai virus infection, cells were harvested for luciferase assays. (G) The experiments were carried out as in (F) except that cells were pretreated with 100 µM PDTC. (H) HEK293 cells were transfected with the indicated plasmids for 30 h, and then stained with MitoSOX followed by FACS analysis. Cells were treated with 0.1 µM H2O2 for 30 min as positive control. Results are presented relative to the FACS mean fluorescence intensity in control cells. (I) HEK293 cells were transfected with increasing amounts of MAVS expression plasmids, and empty vector was used to balance the total DNA amount. Total protein was prepared and subjected to immunoblot analysis after 24 h transfection. Data from A–B, E–G are representative of at least three independent experiments, (mean and s.d. of duplicate assays), and data from C, D and H are presented as mean ± SD from at least three independent experiments. *, P<0.05; **, P<0.01; *** P<0.001 versus control groups.
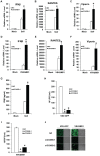
Figure 6. COX5B coordinates with ATG5 to regulate MAVS signaling.
(A) Hela cells were transfected with LC3-GFP (Green) and control vector or Flag-MAVS (Red) for 24 h. Cells were fixed, then stained with anti-Flag antibody, and imaged by confocal microscopy. (B) Hela cells were transfected with increasing amounts of MAVS expression plasmids, and empty vector was used to balance the total DNA amount. Total protein was prepared and subjected to immunoblot analysis after 36 h transfection. (C) HEK293 cells were transfected with increasing amounts of MAVS expression plasmids, and empty vector was used to balance the total DNA amount. Twenty-four hours after transfection, total protein was extracted and subjected to immunoblot analysis. (D) Cell transfection described as seen in (C), after transfection, the total RNAs were prepared for real-time PCR analysis. (E) HEK293 cells were transfected as indicated expression plasmids. Twenty-four hours after transfection, cell lysates were prepared, immunoprecipitated with anti-Flag beads, followed by immunoblot analysis with the indicated antibodies. (F) HEK293 cells were transfected COX5B oligo or NC, after 30 h, transfected again with MAVS or empty vector plasmids, 24 h later, cells were lysed for western blotting. (G) ATG5 and COX5B RNAi oligos (final concentration 20 nM) were transfected into HEK293 cells, NC oligos were used to balance the equal amount of RNAi oligos. Thirty-six hours after transfection, cells were transfected again with the expression plasmids for MAVS or empty vector along with the reporter plasmids followed by measurement of luciferase assays after 24 h transfection. (H) ATG5 and COX5B RNAi oligo transfection was performed as described in (G), and then cells were transfected again with reporter plasmids. Eight hours after second transfection, cells were infected with Sendai virus for 16 h followed by measurement of luciferase assays. Data from D, G and H are representative of at least three independent experiments (mean and s.d. of duplicate assays). *, P<0.05; **, P<0.01 versus control groups.
Figure 7. COX5B and ATG5 suppress MAVS aggregation.
(A) HEK293 cells were transfected with the indicated constructs for 36 h, and crude mitochondrial P5 extracts were isolated, then aliquots of the P5 extracts were subjected to SDD-AGE and SDS-PAGE assays using indicated antibodies respectively. (B) HEK293 cells were transfected with NC or ATG5 RNAi oligos. Thirty-six hours after transfection, Flag-MAVS or empty vectors were transfected into the RNAi cells for 24 h. Crude mitochondrial P5 extracts were prepared, followed by SDD-AGE and SDS-PAGE assays using MAVS or Flag antibody. (C) HEK293 cells were transfected with NC or different doses of ATG5 RNAi oligos as indicated. Thirty-six hours after transfection, knockdown cells were then infected with Sendai virus, 9 h after infection, crude mitochondrial P5 extracts were analyzed by SDD-AGE or SDS-PAGE assays using a MAVS antibody. (D) Cell transfection was performed with indicated plasmids, Thirty-six hours after transfection, crude mitochondrial P5 extracts were isolated, and then aliquots of the P5 extracts were subjected to SDD-AGE and SDS-PAGE assays using indicated antibodies respectively. (E) The experiments were performed as in (B), except that COX5B RNAi oligos were used in lieu of ATG5 RNAi oligos. (F) HEK293 cells were transfected with NC or COX5B RNAi oligos as indicated. Thirty-six hours after transfection, knockdown cells were then infected with Sendai virus, six or nine hours after infection, cell lysates were analyzed by SDD-AGE or SDS-PAGE assays using a MAVS antibody.
Similar articles
- Influenza M2 protein regulates MAVS-mediated signaling pathway through interacting with MAVS and increasing ROS production.
Wang R, Zhu Y, Lin X, Ren C, Zhao J, Wang F, Gao X, Xiao R, Zhao L, Chen H, Jin M, Ma W, Zhou H. Wang R, et al. Autophagy. 2019 Jul;15(7):1163-1181. doi: 10.1080/15548627.2019.1580089. Epub 2019 Feb 20. Autophagy. 2019. PMID: 30741586 Free PMC article. - Dengue Virus Subverts Host Innate Immunity by Targeting Adaptor Protein MAVS.
He Z, Zhu X, Wen W, Yuan J, Hu Y, Chen J, An S, Dong X, Lin C, Yu J, Wu J, Yang Y, Cai J, Li J, Li M. He Z, et al. J Virol. 2016 Jul 27;90(16):7219-7230. doi: 10.1128/JVI.00221-16. Print 2016 Aug 15. J Virol. 2016. PMID: 27252539 Free PMC article. - STAT6 promotes innate immunity against BEFV and VSV by inhibiting STUB1 and NIX-mediated MAVS degradation.
Zhang F, Wang H, He H, Hou P. Zhang F, et al. Vet Microbiol. 2024 Nov;298:110290. doi: 10.1016/j.vetmic.2024.110290. Epub 2024 Oct 25. Vet Microbiol. 2024. PMID: 39471658 - Mechanisms of MAVS regulation at the mitochondrial membrane.
Jacobs JL, Coyne CB. Jacobs JL, et al. J Mol Biol. 2013 Dec 13;425(24):5009-19. doi: 10.1016/j.jmb.2013.10.007. Epub 2013 Oct 9. J Mol Biol. 2013. PMID: 24120683 Free PMC article. Review. - Orchestrating the interferon antiviral response through the mitochondrial antiviral signaling (MAVS) adapter.
Belgnaoui SM, Paz S, Hiscott J. Belgnaoui SM, et al. Curr Opin Immunol. 2011 Oct;23(5):564-72. doi: 10.1016/j.coi.2011.08.001. Epub 2011 Aug 22. Curr Opin Immunol. 2011. PMID: 21865020 Review.
Cited by
- Induction and Evasion of Type-I Interferon Responses during Influenza A Virus Infection.
Muñoz-Moreno R, Martínez-Romero C, García-Sastre A. Muñoz-Moreno R, et al. Cold Spring Harb Perspect Med. 2021 Oct 1;11(10):a038414. doi: 10.1101/cshperspect.a038414. Cold Spring Harb Perspect Med. 2021. PMID: 32661015 Free PMC article. Review. - PTK2B promotes TBK1 and STING oligomerization and enhances the STING-TBK1 signaling.
Lin Y, Yang J, Yang Q, Zeng S, Zhang J, Zhu Y, Tong Y, Li L, Tan W, Chen D, Sun Q. Lin Y, et al. Nat Commun. 2023 Nov 21;14(1):7567. doi: 10.1038/s41467-023-43419-4. Nat Commun. 2023. PMID: 37989995 Free PMC article. - Inflammasome-Independent Role of NLRP3 Mediates Mitochondrial Regulation in Renal Injury.
Kim SM, Kim YG, Kim DJ, Park SH, Jeong KH, Lee YH, Lim SJ, Lee SH, Moon JY. Kim SM, et al. Front Immunol. 2018 Nov 12;9:2563. doi: 10.3389/fimmu.2018.02563. eCollection 2018. Front Immunol. 2018. PMID: 30483252 Free PMC article. - PCBP2 maintains antiviral signaling homeostasis by regulating cGAS enzymatic activity via antagonizing its condensation.
Gu H, Yang J, Zhang J, Song Y, Zhang Y, Xu P, Zhu Y, Wang L, Zhang P, Li L, Chen D, Sun Q. Gu H, et al. Nat Commun. 2022 Mar 23;13(1):1564. doi: 10.1038/s41467-022-29266-9. Nat Commun. 2022. PMID: 35322803 Free PMC article. - Loss of COX5B inhibits proliferation and promotes senescence via mitochondrial dysfunction in breast cancer.
Gao SP, Sun HF, Jiang HL, Li LD, Hu X, Xu XE, Jin W. Gao SP, et al. Oncotarget. 2015 Dec 22;6(41):43363-74. doi: 10.18632/oncotarget.6222. Oncotarget. 2015. PMID: 26506233 Free PMC article.
References
- Akira S, Uematsu S, Takeuchi O (2006) Pathogen recognition and innate immunity. Cell 124: 783–801. - PubMed
- Yoneyama M, Fujita T (2009) RNA recognition and signal transduction by RIG-I-like receptors. Immunol Rev 227: 54–65. - PubMed
- Baccala R, Kono DH, Theofilopoulos AN (2005) Interferons as pathogenic effectors in autoimmunity. Immunol Rev 204: 9–26. - PubMed
- Theofilopoulos AN, Baccala R, Beutler B, Kono DH (2005) Type I interferons (alpha/beta) in immunity and autoimmunity. Annu Rev Immunol 23: 307–336. - PubMed
- Onoguchi K, Yoneyama M, Takemura A, Akira S, Taniguchi T, et al. (2007) Viral infections activate types I and III interferon genes through a common mechanism. J Biol Chem 282: 7576–7581. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
This work was supported by grants from the National Basic Research Program of China (2010CB945300), National Natural Science Foundation of China (30872349), the Ministry of Agriculture of China for Transgenic Research (2009ZX08009154-006) and the Chinese Academy of Sciences (one hundred talents program). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
LinkOut - more resources
Full Text Sources
Miscellaneous