Cytotoxic chromosomal targeting by CRISPR/Cas systems can reshape bacterial genomes and expel or remodel pathogenicity islands - PubMed (original) (raw)
Cytotoxic chromosomal targeting by CRISPR/Cas systems can reshape bacterial genomes and expel or remodel pathogenicity islands
Reuben B Vercoe et al. PLoS Genet. 2013 Apr.
Abstract
In prokaryotes, clustered regularly interspaced short palindromic repeats (CRISPRs) and their associated (Cas) proteins constitute a defence system against bacteriophages and plasmids. CRISPR/Cas systems acquire short spacer sequences from foreign genetic elements and incorporate these into their CRISPR arrays, generating a memory of past invaders. Defence is provided by short non-coding RNAs that guide Cas proteins to cleave complementary nucleic acids. While most spacers are acquired from phages and plasmids, there are examples of spacers that match genes elsewhere in the host bacterial chromosome. In Pectobacterium atrosepticum the type I-F CRISPR/Cas system has acquired a self-complementary spacer that perfectly matches a protospacer target in a horizontally acquired island (HAI2) involved in plant pathogenicity. Given the paucity of experimental data about CRISPR/Cas-mediated chromosomal targeting, we examined this process by developing a tightly controlled system. Chromosomal targeting was highly toxic via targeting of DNA and resulted in growth inhibition and cellular filamentation. The toxic phenotype was avoided by mutations in the cas operon, the CRISPR repeats, the protospacer target, and protospacer-adjacent motif (PAM) beside the target. Indeed, the natural self-targeting spacer was non-toxic due to a single nucleotide mutation adjacent to the target in the PAM sequence. Furthermore, we show that chromosomal targeting can result in large-scale genomic alterations, including the remodelling or deletion of entire pre-existing pathogenicity islands. These features can be engineered for the targeted deletion of large regions of bacterial chromosomes. In conclusion, in DNA-targeting CRISPR/Cas systems, chromosomal interference is deleterious by causing DNA damage and providing a strong selective pressure for genome alterations, which may have consequences for bacterial evolution and pathogenicity.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
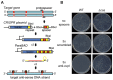
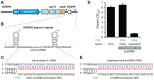
Figure 1. An engineered CRISPR plasmid with spacers targeting the chromosome displays Cas–dependent toxicity.
(A) Strategy for chromosomal targeting (see Materials and Methods). Protospacers were PCR-amplified from target DNA with full or partial repeats and BbsI sites on the primers. PCR products were digested with BbsI and cloned into BbsI-digested plasmids containing a leader sequence and one repeat. This process was repeated to generate multiple spacer inserts. These arrays generate crRNAs that target the template DNA strand but not the mRNA. (B) Transformation plates of P. atrosepticum WT or a Δ_cas_ mutant (PCF80) after growth for 36 h on LBA containing Ap. Plasmids transformed contained no spacers (pC1-780), 3 scrambled spacers (pS3-780) and 3 anti-expI spacers (pE3-780). Representatives are shown from experiments performed at least in triplicate.
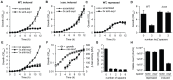
Figure 2. Controlled expression of single anti-expI or anti- lacZ spacers causes cas –dependent growth inhibition and a reduction in viable count.
Plasmids containing either 3 scrambled spacers (pS3-16) or 3 spacers targeting expI (pE3-16) were induced in either (A) WT or (B) Δ_cas_ (PCF80) strains or (C) repressed in the WT. (D) Plasmids containing either no spacers (pC1-16) or 3 spacers targeting lacZ (pL3-16) were induced in either WT or Δ_cas_ strains and measured after 12 h. (E) Plasmids containing no spacers (pC1-16) or one spacer targeting expI (pE1-16) or lacZ (pL1-16) were induced in the WT. (F) Expression of the cas operon measured from a chromosomal transcriptional/translational lacZ fusion (PCF79). (G) Plasmids containing no spacers (pC1-16) or one, two, three or eight identical spacers (pTraG1-16, pTraG2-16, pTraG3-16 and pTraG8-16) display similar toxicity. (H) Viable count (cfu/ml) after 12 h with either repression or induction of plasmids encoding no spacers (none) or 1× anti-expI (expI). Limit of detection was 1×102. Data shown are the mean ± the SD of three experiments.
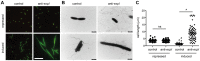
Figure 3. Chromosomal targeting results in cell elongation indicative of DNA damage.
Plasmids containing no spacers (control; pC1-16) or one spacer targeting expI (anti-expI; pE1-16) were repressed or induced in the WT and then visualised by (A) LIVE/DEAD staining and fluorescence microscopy (white scale bar; 20 µm; all images to scale) or (B) transmission electron microscopy (TEM) (black scale bars; 2 µm) (C) Quantification of cell lengths of 60 cells from each treatment as assessed by TEM. ns, not significant; *, p-value of <0.0001 when assessed by unpaired two-tailed t-test (PRISM).
Figure 4. Protospacer, PAM, and repeat mutants can escape toxicity.
(A) Predicted pairing between an expI crRNA and the expI protospacer (PAM is green, spacer is blue, protospacer is red and seed sequence is bold and underlined). (B) Left; protospacer sequences in WT, Δ_expI_ (PCF81), Δ_expI_ with a single WT (RBV01), C3T (RBV04), C6T (RBV03) or G-1T (PAM) (RBV02) expI protospacer. Right; toxicity assays of a single anti-expI spacer (expI; pE1-16; white bars) or a plasmid with no spacers (none; pC1-16; black bar) when expressed in the backgrounds shown on the left. (C) Predicted folds and position of mutations for WT, G20A, C18A and C18A/G8U single CRISPR repeats. The black triangle represents the site of cleavage by Cas6f. (D) Toxicity assays of plasmids expressing a single anti-expI spacer flanked by either WT (pE1-16), G20A (pE1-16 G20A), C18A (pE1-16 C18A) and C18A/G8U repeats (pE1-16 C18A/G8U) in WT P. atrosepticum.
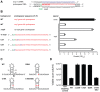
Figure 5. A single nucleotide PAM mutation enables escape from native CRISPR/Cas targeting.
(A) P. atrosepticum CRISPR/Cas genomic organisation. The cas1 and the _cas2_-cas3 hybrid genes are shown in blue, _csy1_-3 in pale blue, cas6f in orange and the CRISPRs as grey arrows in the direction of transcription. CRISPR2 and 3 are separated by a toxin/antitoxin system (white arrows). (B) Spacer 6 of CRISPR2 (from leader) contains deviations from the repeat consensus (shown in blue) and (C) has a 100% match to a protospacer (red) within eca0560 in HAI2 in the P. atrosepticum genome. The protospacer matching spacer 6 contains a non-consensus 5′-protospacer-TG-3′ PAM (green). (D) Toxicity assays in the WT with plasmids containing consensus repeats and either no spacer (pC1-16), a single native spacer 6 (5′-protospacer-TG-3′ PAM; pTraGS6-16) or an engineered spacer (5′-protospacer-GG-3′ PAM; pTraG1-16) that targets eca0560. The protospacer locations in eca0560 of the native (pTraGS6-16) and engineered (pTraG1-16) spacers are shown below in red. (E) Sequence and pairing of the engineered anti-eca0560 spacer (in pTraG1-16) with the consensus 5′-protospacer-GG-3′ PAM (green).
Figure 6. CRISPR/Cas–mediated chromosomal targeting causes rapid genome evolution.
(A) Schematic of HAI2 inserted in the P. atrosepticum genome. The attL and attR sites are indicated by black and white boxes respectively, eca0560 and eca0573 are shown as white arrows and the protospacer in eca0560 is indicated in grey and the KmR marker in eca0573 is depicted in black. Excision of HAI2 results in the circularised form, pHAI2, which contains the attP site and results in the generation of the attB site located within the phe-tRNA gene in the genome. In the absence of the circularised pHAI2 form, the strain is designated ΔHAI2. Primers used for strain confirmation in (B) are shown as black arrows and the cas1 gene was used as a positive control. (B) PCR results for representative class I and class II mutants and the WT. PCR was performed for cas1, eca0560, attR, attL, attP and attB and with primers shown in part (A). Schematic representations of (C) the class I mutants (designated ΔHAI2) that have precisely lost the 97,875 bp island and (D) the class II mutants 2, 5 and 14 containing a 40,227 bp deletion between TTGGCAC sequences in both eca0522 and internal to the KmR insertion in eca0573. (E) Scale genetic map of the 7 classified HAI2 class II mutants defining the deleted regions. Black bars indicate the presence of the gene as specified. The gray vertical line represents the CRISPR-targeted eca0560 gene. The star represents three accurately sequenced junctions. The blue arrow depicts the site of Km insertion within the chromosome. Class II mutants are numbered as in Figure S4 and their PCR profiles are shown in Figure S5.
Similar articles
- Chromosomal targeting by CRISPR-Cas systems can contribute to genome plasticity in bacteria.
Dy RL, Pitman AR, Fineran PC. Dy RL, et al. Mob Genet Elements. 2013 Sep 1;3(5):e26831. doi: 10.4161/mge.26831. Epub 2013 Oct 25. Mob Genet Elements. 2013. PMID: 24251073 Free PMC article. - Priming in the Type I-F CRISPR-Cas system triggers strand-independent spacer acquisition, bi-directionally from the primed protospacer.
Richter C, Dy RL, McKenzie RE, Watson BN, Taylor C, Chang JT, McNeil MB, Staals RH, Fineran PC. Richter C, et al. Nucleic Acids Res. 2014 Jul;42(13):8516-26. doi: 10.1093/nar/gku527. Epub 2014 Jul 2. Nucleic Acids Res. 2014. PMID: 24990370 Free PMC article. - Protospacer-Adjacent Motif Specificity during Clostridioides difficile Type I-B CRISPR-Cas Interference and Adaptation.
Maikova A, Boudry P, Shiriaeva A, Vasileva A, Boutserin A, Medvedeva S, Semenova E, Severinov K, Soutourina O. Maikova A, et al. mBio. 2021 Aug 31;12(4):e0213621. doi: 10.1128/mBio.02136-21. Epub 2021 Aug 24. mBio. 2021. PMID: 34425703 Free PMC article. - The subtype I-F CRISPR-Cas system influences pathogenicity island retention in Pectobacterium atrosepticum via crRNA generation and Csy complex formation.
Richter C, Fineran PC. Richter C, et al. Biochem Soc Trans. 2013 Dec;41(6):1468-74. doi: 10.1042/BST20130151. Biochem Soc Trans. 2013. PMID: 24256239 Review. - Friendly Fire: Biological Functions and Consequences of Chromosomal Targeting by CRISPR-Cas Systems.
Heussler GE, O'Toole GA. Heussler GE, et al. J Bacteriol. 2016 Apr 28;198(10):1481-6. doi: 10.1128/JB.00086-16. Print 2016 May 15. J Bacteriol. 2016. PMID: 26929301 Free PMC article. Review.
Cited by
- Pectobacterium atrosepticum and Pectobacterium carotovorum Harbor Distinct, Independently Acquired Integrative and Conjugative Elements Encoding Coronafacic Acid that Enhance Virulence on Potato Stems.
Panda P, Vanga BR, Lu A, Fiers M, Fineran PC, Butler R, Armstrong K, Ronson CW, Pitman AR. Panda P, et al. Front Microbiol. 2016 Mar 31;7:397. doi: 10.3389/fmicb.2016.00397. eCollection 2016. Front Microbiol. 2016. PMID: 27065965 Free PMC article. - Major and minor crRNA annealing sites facilitate low stringency DNA protospacer binding prior to Type I-A CRISPR-Cas interference in Sulfolobus.
Mousaei M, Deng L, She Q, Garrett RA. Mousaei M, et al. RNA Biol. 2016 Nov;13(11):1166-1173. doi: 10.1080/15476286.2016.1229735. Epub 2016 Sep 12. RNA Biol. 2016. PMID: 27618562 Free PMC article. - Efficient Cas9-based genome editing of Rhodobacter sphaeroides for metabolic engineering.
Mougiakos I, Orsi E, Ghiffary MR, Post W, de Maria A, Adiego-Perez B, Kengen SWM, Weusthuis RA, van der Oost J. Mougiakos I, et al. Microb Cell Fact. 2019 Nov 25;18(1):204. doi: 10.1186/s12934-019-1255-1. Microb Cell Fact. 2019. PMID: 31767004 Free PMC article. - Adaptation by Type V-A and V-B CRISPR-Cas Systems Demonstrates Conserved Protospacer Selection Mechanisms Between Diverse CRISPR-Cas Types.
Wu WY, Jackson SA, Almendros C, Haagsma AC, Yilmaz S, Gort G, van der Oost J, Brouns SJJ, Staals RHJ. Wu WY, et al. CRISPR J. 2022 Aug;5(4):536-547. doi: 10.1089/crispr.2021.0150. Epub 2022 Jul 12. CRISPR J. 2022. PMID: 35833800 Free PMC article. - Resistance and tolerance to foreign elements by prokaryotic immune systems - curating the genome.
Goldberg GW, Marraffini LA. Goldberg GW, et al. Nat Rev Immunol. 2015 Nov;15(11):717-24. doi: 10.1038/nri3910. Nat Rev Immunol. 2015. PMID: 26494050 Free PMC article. Review.
References
- Petty NK, Evans TJ, Fineran PC, Salmond GP (2007) Biotechnological exploitation of bacteriophage research. Trends Biotechnol 25: 7–15. - PubMed
- Hendrix RW (2003) Bacteriophage genomics. Curr Opin Microbiol 6: 506–511. - PubMed
- Weinbauer MG (2004) Ecology of prokaryotic viruses. FEMS Microbiol Rev 28: 127–181. - PubMed
- Labrie SJ, Samson JE, Moineau S (2010) Bacteriophage resistance mechanisms. Nat Rev Microbiol 8: 317–327. - PubMed
Publication types
MeSH terms
Grants and funding
This work was supported by University of Otago Research Grants, a Priming Partnerships Grant, Tertiary Education Commission NZ, the Marsden Fund, and a Rutherford Discovery Fellowship (PCF), both from the Royal Society of NZ. CR was supported by a University of Otago Postgraduate Scholarship and a DAAD Doktorandenstipendium. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous