Assembling the 20 Gb white spruce (Picea glauca) genome from whole-genome shotgun sequencing data - PubMed (original) (raw)
. 2013 Jun 15;29(12):1492-7.
doi: 10.1093/bioinformatics/btt178. Epub 2013 May 22.
Anthony Raymond, Shaun D Jackman, Stephen Pleasance, Robin Coope, Greg A Taylor, Macaire Man Saint Yuen, Christopher I Keeling, Dana Brand, Benjamin P Vandervalk, Heather Kirk, Pawan Pandoh, Richard A Moore, Yongjun Zhao, Andrew J Mungall, Barry Jaquish, Alvin Yanchuk, Carol Ritland, Brian Boyle, Jean Bousquet, Kermit Ritland, John Mackay, Jörg Bohlmann, Steven J M Jones
Affiliations
- PMID: 23698863
- PMCID: PMC3673215
- DOI: 10.1093/bioinformatics/btt178
Assembling the 20 Gb white spruce (Picea glauca) genome from whole-genome shotgun sequencing data
Inanc Birol et al. Bioinformatics. 2013.
Abstract
White spruce (Picea glauca) is a dominant conifer of the boreal forests of North America, and providing genomics resources for this commercially valuable tree will help improve forest management and conservation efforts. Sequencing and assembling the large and highly repetitive spruce genome though pushes the boundaries of the current technology. Here, we describe a whole-genome shotgun sequencing strategy using two Illumina sequencing platforms and an assembly approach using the ABySS software. We report a 20.8 giga base pairs draft genome in 4.9 million scaffolds, with a scaffold N50 of 20,356 bp. We demonstrate how recent improvements in the sequencing technology, especially increasing read lengths and paired end reads from longer fragments have a major impact on the assembly contiguity. We also note that scalable bioinformatics tools are instrumental in providing rapid draft assemblies.
Availability: The Picea glauca genome sequencing and assembly data are available through NCBI (Accession#: ALWZ0100000000 PID: PRJNA83435). http://www.ncbi.nlm.nih.gov/bioproject/83435.
Figures
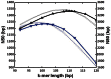
Fig. 1.
Assembly optimization. The de Bruijn graph stage (pre-unitig) of the assembly was used to optimize the overlap parameter, _k_-mer length, and the effect of inclusion of longer reads was assessed. The contiguity metrics N50 (solid curves, left _y_-axis) and N20 (dotted curves, right _y_-axis) are shown for assemblies that use the short reads only (blue) and short and long reads (black). The contiguity of the two datasets peaked for different _k_-mer lengths, with dataset of short and long reads having a maximum N50 and a maximum N20 for the same k = 109 bp. For short reads only, optimization with respect to N20 resulted in a slightly lower _k_-mer length (98 bp) compared with optimization with respect to N50 (101 bp), both of which are lower than the optimum _k_-mer length for the full dataset. Longer _k_-mers were desirable, as they help disambiguate longer repeat motifs
Comment in
- Genomics: Sprucing up forest tree genomics.
Skipper M. Skipper M. Nat Rev Genet. 2013 Jul;14(7):444. doi: 10.1038/nrg3518. Epub 2013 Jun 4. Nat Rev Genet. 2013. PMID: 23732336 No abstract available.
References
- Altschul SF, et al. Basic local alignment search tool. J. Mol. Biol. 1990;215:403–410. -PubMed
- Burrows M, Wheeler D. Technical Report 124. Palo Alto, CA: Digital Equipment Corporation; 1994. A block sorting lossless data compression algorithm.
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases