Orthogonal Cas9 proteins for RNA-guided gene regulation and editing - PubMed (original) (raw)
Orthogonal Cas9 proteins for RNA-guided gene regulation and editing
Kevin M Esvelt et al. Nat Methods. 2013 Nov.
Abstract
The Cas9 protein from the Streptococcus pyogenes CRISPR-Cas acquired immune system has been adapted for both RNA-guided genome editing and gene regulation in a variety of organisms, but it can mediate only a single activity at a time within any given cell. Here we characterize a set of fully orthogonal Cas9 proteins and demonstrate their ability to mediate simultaneous and independently targeted gene regulation and editing in bacteria and in human cells. We find that Cas9 orthologs display consistent patterns in their recognition of target sequences, and we identify an unexpectedly versatile Cas9 protein from Neisseria meningitidis. We provide a basal set of orthogonal RNA-guided proteins for controlling biological systems and establish a general methodology for characterizing additional proteins.
Conflict of interest statement
Competing Financial Interests
The authors have filed for patents concerning the use of Cas9 proteins for gene targeting and regulation.
Figures
Fig. 1
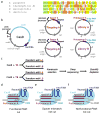
Comparison and characterization of putatively orthogonal Cas9 proteins. (a) Repeat sequences of SP, ST1, NM, and TD. Bases are colored to indicate the degree of conservation. (b) Plasmids used for characterization of Cas9 proteins in E. coli. All carry compatible replication origins and antibiotic resistance genes. (c) Selection scheme to identify PAMs. Cells expressing a Cas9 protein and one of two spacer-containing targeting plasmids were transformed with one of two PAM libraries with corresponding protospacers and subjected to antibiotic selection. Surviving uncleaved plasmids were subjected to deep sequencing. Cas9-mediated PAM depletion was quantified by comparing the relative abundance of each sequence within the matched versus the mismatched protospacer libraries. (d) Functional PAMs are depleted from the library by Cas9 when the targeting plasmid spacer matches the library plasmid protospacer. (e) Cas9 does not cut when the spacer and protospacer do not match. (f) Nonfunctional PAMs are never cut or depleted.
Fig. 2
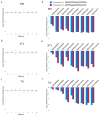
Depletion of functional protospacer-adjacent motifs (PAMs) from libraries by Cas9 proteins. The log frequency of each base at every position for matched spacer-protospacer pairs is plotted relative to control conditions in which spacer and protospacer did not match. Results reflect the mean depletion of libraries by NM (a), ST1 (b), and TD (c) based on two distinct protospacer sequences (d). Depletion of specific sequences for each protospacer are plotted separately for each Cas9 protein (d–f).
Fig. 3
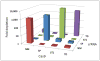
Orthogonal recognition of crRNAs in E. coli. Cells with all 16 combinations of Cas9 and crRNA were challenged with a plasmid bearing a matched or mismatched protospacer and appropriate PAM. Sufficient cells were plated to reliably obtain colonies from matching spacer and protospacer pairings. Total colony counts on the resulting 32 plates were used to calculate the fold depletion. Values less than one were set to one for clarity.
Fig. 4
Transcriptional repression simultaneous with nuclease activity in bacteria. (a) Reporter plasmids for quantification of Cas9 repression contained protospacer B and a suitable PAM in the non-template strand after the YFP start codon. Normalized cellular fluorescence is shown for mismatched and matched spacer-protospacer pairs. PAMs for each Cas9 are shown. (b) Cas9 ortholog repression was verified with a second reporter plasmid containing protospacer A and a PAM in the non-template strand within the 5′ UTR. Error bars represent the standard deviation of eight independently picked cultures for all repression experiments. (c) Cells containing the plasmids used for NM-mediated repression (Fig. 3a) were transformed with a compatible plasmid encoding SP, its tracrRNA, and a 5-spacer CRISPR locus designed to cleave filamentous phage gene III at multiple sites and challenged with M13mp18. The phage defense plasmid completely prevented plaque formation while preserving YFP repression. (d) Cells were transformed with a compatible plasmid encoding carbenicillin resistance and either wild-type gene III or a recoded version lacking protospacers and plated. Plasmids encoding wild-type gene III were perfectly excluded from cells encoding the SP phage defense, which did not interfere with YFP repression. Scale bars, 10 mm.
Fig. 5
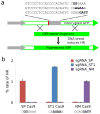
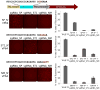
Cas9-mediated gene editing in human cells. (a) A homologous recombination assay was used to quantify gene editing efficiency. Cas9-mediated double-strand breaks within the protospacer stimulated repair of the interrupted GFP cassette using the donor template, yielding cells with intact GFP. Three different templates were used in order to provide the correct PAM for each Cas9. Fluorescent cells were quantified by flow cytometry. (b) Homologous recombination efficiencies for NM, ST1, and TD in combination with each of their respective sgRNAs. Substrate PAMs are displayed below each Cas9. Data represent mean values ± s.e.m. (n=3).
Fig. 6
Transcriptional activation in human cells. (a) Reporter constructs for transcriptional activation featured a minimal promoter driving tdTomato. Nuclease-null Cas9-VP64 fusion proteins binding to upstream protospacers resulted in transcriptional activation and enhanced fluorescence. (b) Cells were transfected with all combinations of Cas9 activators and sgRNAs and tdTomato fluorescence visualized by microscopy. Transcriptional activation occurred only when each Cas9 was paired with its own sgRNA. Scale bars, 100 μm. (c) Activation was quantified by flow cytometry along with a TAL-VP64 effector targeting an upstream sequence for comparison. Data represent mean values ± s.e.m. (n=3).
Similar articles
- Bacterial CRISPR/Cas DNA endonucleases: A revolutionary technology that could dramatically impact viral research and treatment.
Kennedy EM, Cullen BR. Kennedy EM, et al. Virology. 2015 May;479-480:213-20. doi: 10.1016/j.virol.2015.02.024. Epub 2015 Mar 7. Virology. 2015. PMID: 25759096 Free PMC article. Review. - One-step high-efficiency CRISPR/Cas9-mediated genome editing in Streptomyces.
Huang H, Zheng G, Jiang W, Hu H, Lu Y. Huang H, et al. Acta Biochim Biophys Sin (Shanghai). 2015 Apr;47(4):231-43. doi: 10.1093/abbs/gmv007. Epub 2015 Mar 3. Acta Biochim Biophys Sin (Shanghai). 2015. PMID: 25739462 - Orthogonal Cas9-Cas9 chimeras provide a versatile platform for genome editing.
Bolukbasi MF, Liu P, Luk K, Kwok SF, Gupta A, Amrani N, Sontheimer EJ, Zhu LJ, Wolfe SA. Bolukbasi MF, et al. Nat Commun. 2018 Nov 19;9(1):4856. doi: 10.1038/s41467-018-07310-x. Nat Commun. 2018. PMID: 30451839 Free PMC article. - Editing of the Bacillus subtilis Genome by the CRISPR-Cas9 System.
Altenbuchner J. Altenbuchner J. Appl Environ Microbiol. 2016 Aug 15;82(17):5421-7. doi: 10.1128/AEM.01453-16. Print 2016 Sep 1. Appl Environ Microbiol. 2016. PMID: 27342565 Free PMC article. - Applications of Cas9 as an RNA-programmed RNA-binding protein.
Nelles DA, Fang MY, Aigner S, Yeo GW. Nelles DA, et al. Bioessays. 2015 Jul;37(7):732-9. doi: 10.1002/bies.201500001. Epub 2015 Apr 16. Bioessays. 2015. PMID: 25880497 Free PMC article. Review.
Cited by
- Making gene editing accessible in resource limited environments: recommendations to guide a first-time user.
Goolab S, Scholefield J. Goolab S, et al. Front Genome Ed. 2024 Sep 25;6:1464531. doi: 10.3389/fgeed.2024.1464531. eCollection 2024. Front Genome Ed. 2024. PMID: 39386178 Free PMC article. Review. - Multiplex, single-cell CRISPRa screening for cell type specific regulatory elements.
Chardon FM, McDiarmid TA, Page NF, Daza RM, Martin BK, Domcke S, Regalado SG, Lalanne JB, Calderon D, Li X, Starita LM, Sanders SJ, Ahituv N, Shendure J. Chardon FM, et al. Nat Commun. 2024 Sep 18;15(1):8209. doi: 10.1038/s41467-024-52490-4. Nat Commun. 2024. PMID: 39294132 Free PMC article. - Gene editing of angiotensin for blood pressure management.
Masi S, Dalpiaz H, Borghi C. Masi S, et al. Int J Cardiol Cardiovasc Risk Prev. 2024 Aug 20;23:200323. doi: 10.1016/j.ijcrp.2024.200323. eCollection 2024 Dec. Int J Cardiol Cardiovasc Risk Prev. 2024. PMID: 39258007 Free PMC article. - Advancing CRISPR base editing technology through innovative strategies and ideas.
Fan X, Lei Y, Wang L, Wu X, Li D. Fan X, et al. Sci China Life Sci. 2024 Sep 2. doi: 10.1007/s11427-024-2699-5. Online ahead of print. Sci China Life Sci. 2024. PMID: 39231901 Review. - Editing microbes to mitigate enteric methane emissions in livestock.
Khan FA, Ali A, Wu D, Huang C, Zulfiqar H, Ali M, Ahmed B, Yousaf MR, Putri EM, Negara W, Imran M, Pandupuspitasari NS. Khan FA, et al. World J Microbiol Biotechnol. 2024 Aug 13;40(10):300. doi: 10.1007/s11274-024-04103-x. World J Microbiol Biotechnol. 2024. PMID: 39134917 Review.
References
- Bhaya D, Davison M, Barrangou R. CRISPR-Cas systems in bacteria and archaea: versatile small RNAs for adaptive defense and regulation. Annual review of genetics. 2011;45:273–297. - PubMed
- Wiedenheft B, Sternberg SH, Doudna JA. RNA-guided genetic silencing systems in bacteria and archaea. Nature. 2012;482:331–338. - PubMed
- Cho SW, Kim S, Kim JM, Kim JS. Targeted genome engineering in human cells with the Cas9 RNA-guided endonuclease. Nature biotechnology. 2013;31:230–232. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials