ClinVar: public archive of relationships among sequence variation and human phenotype - PubMed (original) (raw)
. 2014 Jan;42(Database issue):D980-5.
doi: 10.1093/nar/gkt1113. Epub 2013 Nov 14.
Affiliations
- PMID: 24234437
- PMCID: PMC3965032
- DOI: 10.1093/nar/gkt1113
ClinVar: public archive of relationships among sequence variation and human phenotype
Melissa J Landrum et al. Nucleic Acids Res. 2014 Jan.
Abstract
ClinVar (http://www.ncbi.nlm.nih.gov/clinvar/) provides a freely available archive of reports of relationships among medically important variants and phenotypes. ClinVar accessions submissions reporting human variation, interpretations of the relationship of that variation to human health and the evidence supporting each interpretation. The database is tightly coupled with dbSNP and dbVar, which maintain information about the location of variation on human assemblies. ClinVar is also based on the phenotypic descriptions maintained in MedGen (http://www.ncbi.nlm.nih.gov/medgen). Each ClinVar record represents the submitter, the variation and the phenotype, i.e. the unit that is assigned an accession of the format SCV000000000.0. The submitter can update the submission at any time, in which case a new version is assigned. To facilitate evaluation of the medical importance of each variant, ClinVar aggregates submissions with the same variation/phenotype combination, adds value from other NCBI databases, assigns a distinct accession of the format RCV000000000.0 and reports if there are conflicting clinical interpretations. Data in ClinVar are available in multiple formats, including html, download as XML, VCF or tab-delimited subsets. Data from ClinVar are provided as annotation tracks on genomic RefSeqs and are used in tools such as Variation Reporter (http://www.ncbi.nlm.nih.gov/variation/tools/reporter), which reports what is known about variation based on user-supplied locations.
Figures
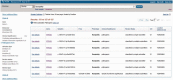
Figure 1.
Tabular display of query results with filters applied. In this example
http://www.ncbi.nlm.nih.gov/clinvar?term=rasopathy&cmd=DetailsSearch
), the query ‘rasopathy’ returned more than 225 results, but when the selection of ‘Pathogenic’ was applied, the number of results was reduced to 127. The number of records in each category is reported. The check mark to the left of the filter name as well as the report at the top of the page are used to remind the user about the filters that have been applied. These filters can be removed using the ‘Clear’ option. To see the details of any record that was found, click on ‘See details’. To alter the display and change the sorting order, open the ‘Display Settings’ menu above the table.
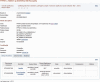
Figure 2.
Representative display of an RCV accession RCV000033555.3 generated by aggregating information in two submissions (SCV000057460 and SCV000058286). The information above the tabbed section includes information from the submitters and values added by NCBI (e.g_._ GeneIDs, Mendelian Inheritance in Man (MIM) numbers, rs numbers and links to MedGen). The information in the ‘Clinical Assertions’ and ‘Evidence’ tabs are organized by what each submitter contributed and are extracted from the SCV records.
Similar articles
- ClinVar: public archive of interpretations of clinically relevant variants.
Landrum MJ, Lee JM, Benson M, Brown G, Chao C, Chitipiralla S, Gu B, Hart J, Hoffman D, Hoover J, Jang W, Katz K, Ovetsky M, Riley G, Sethi A, Tully R, Villamarin-Salomon R, Rubinstein W, Maglott DR. Landrum MJ, et al. Nucleic Acids Res. 2016 Jan 4;44(D1):D862-8. doi: 10.1093/nar/gkv1222. Epub 2015 Nov 17. Nucleic Acids Res. 2016. PMID: 26582918 Free PMC article. - ClinVar: improvements to accessing data.
Landrum MJ, Chitipiralla S, Brown GR, Chen C, Gu B, Hart J, Hoffman D, Jang W, Kaur K, Liu C, Lyoshin V, Maddipatla Z, Maiti R, Mitchell J, O'Leary N, Riley GR, Shi W, Zhou G, Schneider V, Maglott D, Holmes JB, Kattman BL. Landrum MJ, et al. Nucleic Acids Res. 2020 Jan 8;48(D1):D835-D844. doi: 10.1093/nar/gkz972. Nucleic Acids Res. 2020. PMID: 31777943 Free PMC article. - ClinVar: improving access to variant interpretations and supporting evidence.
Landrum MJ, Lee JM, Benson M, Brown GR, Chao C, Chitipiralla S, Gu B, Hart J, Hoffman D, Jang W, Karapetyan K, Katz K, Liu C, Maddipatla Z, Malheiro A, McDaniel K, Ovetsky M, Riley G, Zhou G, Holmes JB, Kattman BL, Maglott DR. Landrum MJ, et al. Nucleic Acids Res. 2018 Jan 4;46(D1):D1062-D1067. doi: 10.1093/nar/gkx1153. Nucleic Acids Res. 2018. PMID: 29165669 Free PMC article. - Database resources of the National Center for Biotechnology Information.
Sayers EW, Beck J, Bolton EE, Bourexis D, Brister JR, Canese K, Comeau DC, Funk K, Kim S, Klimke W, Marchler-Bauer A, Landrum M, Lathrop S, Lu Z, Madden TL, O'Leary N, Phan L, Rangwala SH, Schneider VA, Skripchenko Y, Wang J, Ye J, Trawick BW, Pruitt KD, Sherry ST. Sayers EW, et al. Nucleic Acids Res. 2021 Jan 8;49(D1):D10-D17. doi: 10.1093/nar/gkaa892. Nucleic Acids Res. 2021. PMID: 33095870 Free PMC article. Review. - Mucopolysaccharidosis type VI (MPS VI) and molecular analysis: Review and classification of published variants in the ARSB gene.
Tomanin R, Karageorgos L, Zanetti A, Al-Sayed M, Bailey M, Miller N, Sakuraba H, Hopwood JJ. Tomanin R, et al. Hum Mutat. 2018 Dec;39(12):1788-1802. doi: 10.1002/humu.23613. Epub 2018 Sep 17. Hum Mutat. 2018. PMID: 30118150 Free PMC article. Review.
Cited by
- Genomic and transcriptomic analyses identify distinctive features of triple-negative inflammatory breast cancer.
Wang X, Zhao L, Song X, Wu X, Krishnamurthy S, Semba T, Shao S, Knafl M, Coffer LW 2nd, Alexander A, Vines A, Bopparaju S, Woodward WA, Chu R, Zhang J, Yam C, Loo LWM, Nasrazadani A, Huong LP, Woodman SE, Futreal A; Rare Tumor Initiative Team; Tripathy D, Ueno NT. Wang X, et al. NPJ Precis Oncol. 2024 Nov 18;8(1):265. doi: 10.1038/s41698-024-00729-0. NPJ Precis Oncol. 2024. PMID: 39558017 - Reclassification of Two MLH1 Variants of Uncertain Significance Utilizing Clinical and Functional Data.
Frederiksen JH, Birkedal U, Bachmann S, Eliesen EV, Rasmussen LJ, Pedersen KV, Al-Zehhawi L, Boonen SE, Krogh L, Rønlund K, Graversen L, Assenholt J, Schmiegelow K, Wadt K, Gerdes AM, Hansen TVO. Frederiksen JH, et al. Mol Genet Genomic Med. 2024 Nov;12(11):e70026. doi: 10.1002/mgg3.70026. Mol Genet Genomic Med. 2024. PMID: 39548353 Free PMC article. - BRCAIndica: a resource for ACMG/AMP classified BRCA1 and BRCA2 variants.
Vatsyayan A, Anu RI, Mathur P, Uchil D, Joshi A, Dwivedi A, Sirohi B, Mathew A, Damodaran D, Panda SS, Kolluri S, Ayillath SK, Amalnath D, Shankar G, Pandhare K, Scaria V. Vatsyayan A, et al. Fam Cancer. 2024 Nov 15;24(1):4. doi: 10.1007/s10689-024-00429-5. Fam Cancer. 2024. PMID: 39546087 - Defects in meiosis I contribute to the genesis of androgenetic hydatidiform moles.
Rezaei M, Liang M, Yalcin Z, Martin JH, Kazemi P, Bareke E, Ge ZJ, Fardaei M, Benadiva C, Hemida R, Hassan A, Maher GJ, Abdalla E, Buckett W, Bolze PA, Sandhu I, Duman O, Agrawal S, Qian J, Vallian Broojeni J, Bhati L, Miron P, Allias F, Selim A, Fisher RA, Seckl MJ, Sauthier P, Touitou I, Tan SL, Majewski J, Taketo T, Slim R. Rezaei M, et al. J Clin Invest. 2024 Nov 15;134(22):e170669. doi: 10.1172/JCI170669. J Clin Invest. 2024. PMID: 39545410 Free PMC article. - A novel functional IKBKE variant activating NFAT in a patient with polyarthritis and a remittent fever.
Yamada S, Nagafuchi Y, Yamada M, Suzuki H, Natsumoto B, Ota M, Takazawa I, Hatano H, Kono M, Harada H, Shoda H, Okamura T, Kosaki K, Fujio K. Yamada S, et al. Front Immunol. 2024 Oct 25;15:1475179. doi: 10.3389/fimmu.2024.1475179. eCollection 2024. Front Immunol. 2024. PMID: 39524436 Free PMC article.
References
- Rubinstein WS, Maglott DR, Lee JM, Kattman BL, Malheiro AJ, Ovetsky M, Hem V, Gorelenkov V, Song G, Wallin C, et al. The NIH genetic testing registry: a new, centralized database of genetic tests to enable access to comprehensive information and improve transparency. Nucleic Acids Res. 2013;41:D925–D935. - PMC - PubMed
- den Dunnen JT, Antonarakis SE. Mutation nomenclature extensions and suggestions to describe complex mutations: a discussion. Hum. Mutation. 2000;15:7–12. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources