Differentiation-defective phenotypes revealed by large-scale analyses of human pluripotent stem cells - PubMed (original) (raw)
. 2013 Dec 17;110(51):20569-74.
doi: 10.1073/pnas.1319061110. Epub 2013 Nov 20.
Mari Ohnuki, Kazutoshi Takahashi, Keisuke Okita, Hisashi Noma, Yuka Sawamura, Ito Teramoto, Megumi Narita, Yoshiko Sato, Tomoko Ichisaka, Naoki Amano, Akira Watanabe, Asuka Morizane, Yasuhiro Yamada, Tosiya Sato, Jun Takahashi, Shinya Yamanaka
Affiliations
- PMID: 24259714
- PMCID: PMC3870695
- DOI: 10.1073/pnas.1319061110
Differentiation-defective phenotypes revealed by large-scale analyses of human pluripotent stem cells
Michiyo Koyanagi-Aoi et al. Proc Natl Acad Sci U S A. 2013.
Abstract
We examined the gene expression and DNA methylation of 49 human induced pluripotent stem cells (hiPSCs) and 10 human embryonic stem cells and found overlapped variations in gene expression and DNA methylation in the two types of human pluripotent stem cell lines. Comparisons of the in vitro neural differentiation of 40 hiPSCs and 10 human embryonic stem cells showed that seven hiPSC clones retained a significant number of undifferentiated cells even after neural differentiation culture and formed teratoma when transplanted into mouse brains. These differentiation-defective hiPSC clones were marked by higher expression levels of several genes, including those expressed from long terminal repeats of specific human endogenous retroviruses. These data demonstrated a subset of hiPSC lines that have aberrant gene expression and defective potential in neural differentiation, which need to be identified and eliminated before applications in regenerative medicine.
Conflict of interest statement
Conflict of interest statement: S.Y. is a member without salary of the scientific advisory boards of iPierian, iPS Academia Japan, Megakaryon Corporation, and HEALIOS K. K. Japan.
Figures
Fig. 1.
hiPSCs and hESCs have overlapped variations in RNA expression and DNA methylation. Scatter plots of mRNA expression (A), miRNA expression (B), and DNA methylation (C) data comparing the average of 49 hiPSC lines (y axis) to the average of 10 hESC lines (x axis). The RNA expression value is shown on a log 2 scale. Green lines indicate twofold differences in the RNA expression levels between the clones. Differentially expressed probes (t test, FDR < 0.05) are shown in magenta. (D) The variations in the mRNA expression levels of 61 differentially expressed probes in hESCs (red) and hiPSCs (black) are shown. Probes are arranged in order of the absolute value of the FC between hESCs and hiPSCs. (E) The DNA methylation profiles for CpGs contained in reported hES-hiPS DMRs and overlapping with our platform. Probes are arranged in order of the differences between the average DNA methylation level of hESCs and that of hiPSCs. The heat map represents the DNA methylation levels from completely methylated (1, magenta) to unmethylated (0, white) samples. The methylation status of the upstream region of PON3 (F) and TCERG1L (G) was examined by pyrosequencing.
Fig. 2.
A differentiation-defective phenotype in a subset of hiPSC clones. (A) A schematic diagram of the SFEBq method used for neural differentiation. (B) Neural induction was performed for 2 hESC and 21 hiPSC lines which were established from various origins by retroviral or episomal vector methods. On day 14, we examined the proportion of PSA-NCAM–expressing cells by flow cytometry (n = 2). (C) The proportions of PSA-NCAM- (white), OCT3/4- (gray), and TRA1-60-positive (black) cells 14 d after neural differentiation. (D) The proportions of OCT3/4-positive cells on day 14 after neural differentiation are ranked in order of their maximum value. The numbers in parentheses show the number of trials.
Fig. 3.
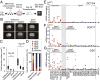
Activation of specific endogenous retroviral LTR7s in defective clones. (A) A scatter plot of the mRNA expression data comparing the average of 38 good clones (y axis) to the average of seven defective clones (x axis). Green lines indicate fivefold differences in expression. A total of 19 differentially expressed probes are colored magenta. (B) The expression levels of LTR7-related genes (HHLA1, ABHD12B, and C4orf51) were examined by microarray. (C) A schematic diagram of three LTR7-related genes. HHLA1 and OC90 are neighboring genes. Dots indicate microarray probes. Magenta dots show probes that are located after LTR7 regions, which were up-regulated in defective clones. (D) A scatter plot of array probes that recognized LTR7-related genes and two other genes [EFR3 homolog A (S. cerevisiae, EFR3A) and potassium voltage-gated channel, KQT-like subfamily, member 3 (KCNQ3)], which are genes neighboring HHLA1 and OC90, respectively. (E) The exon array of the ABHD12B and C4orf51. The average levels of the normalized exon expression are shown. (F) The DNA methylation status of LTR7 and its neighboring regions of HHLA1, ABHD12B, and C4orf51 was examined by pyrosequencing. (n.s., not significant; *P < 0.05, **P < 0.01, Mann–Whitney U test).
Fig. 4.
Transplantation of neural cells derived from hiPSCs and hESCs into mouse brains. (A) A schematic diagram of the SFEBq method used for dopaminergic neural differentiation. On day 29, the cells were transplanted into NOD/SCID mouse brains. (B) Magnetic resonance images of coronal sections of the grafted brains. The section surface of grafted cells indicated as the white shadow in right brain was measured as described in the lower panels. (C) A box-and-whisker plot of the surface sizes of graft sections 30 and 60 d after transplantation. The median, quartile, and range are shown. *P < 0.05 (t test). (D) The proportion of each kind of graft. Grafts were categorized according to their components as determined in H&E sections and were classified by the proportion of neural tissues by a microscopic observation. The expression levels of the undifferentiated cell marker, OCT3/4 (E), the endoderm marker, SOX17 (F), and the mesoderm and endoderm marker, GSC (G), in pretransplantation cultures, undifferentiated hESC lines, and somatic cells (HDF and DP) were examined by qRT-PCR. The colors of the dots were identical to the proportion of neural tissues in D.
Similar articles
- Preventing Pluripotent Cell Teratoma in Regenerative Medicine Applied to Hematology Disorders.
Bedel A, Beliveau F, Lamrissi-Garcia I, Rousseau B, Moranvillier I, Rucheton B, Guyonnet-Dupérat V, Cardinaud B, de Verneuil H, Moreau-Gaudry F, Dabernat S. Bedel A, et al. Stem Cells Transl Med. 2017 Feb;6(2):382-393. doi: 10.5966/sctm.2016-0201. Epub 2016 Sep 7. Stem Cells Transl Med. 2017. PMID: 28191782 Free PMC article. - Reduction of N-glycolylneuraminic acid in human induced pluripotent stem cells generated or cultured under feeder- and serum-free defined conditions.
Hayashi Y, Chan T, Warashina M, Fukuda M, Ariizumi T, Okabayashi K, Takayama N, Otsu M, Eto K, Furue MK, Michiue T, Ohnuma K, Nakauchi H, Asashima M. Hayashi Y, et al. PLoS One. 2010 Nov 23;5(11):e14099. doi: 10.1371/journal.pone.0014099. PLoS One. 2010. PMID: 21124894 Free PMC article. - Teratoma formation of human embryonic stem cells in three-dimensional perfusion culture bioreactors.
Stachelscheid H, Wulf-Goldenberg A, Eckert K, Jensen J, Edsbagge J, Björquist P, Rivero M, Strehl R, Jozefczuk J, Prigione A, Adjaye J, Urbaniak T, Bussmann P, Zeilinger K, Gerlach JC. Stachelscheid H, et al. J Tissue Eng Regen Med. 2013 Sep;7(9):729-41. doi: 10.1002/term.1467. Epub 2012 Mar 21. J Tissue Eng Regen Med. 2013. PMID: 22438087 - Teratoma: from spontaneous tumors to the pluripotency/malignancy assay.
Bulic-Jakus F, Katusic Bojanac A, Juric-Lekic G, Vlahovic M, Sincic N. Bulic-Jakus F, et al. Wiley Interdiscip Rev Dev Biol. 2016 Mar-Apr;5(2):186-209. doi: 10.1002/wdev.219. Epub 2015 Dec 23. Wiley Interdiscip Rev Dev Biol. 2016. PMID: 26698368 Review. - Evaluating Strategies to Assess the Differentiation Potential of Human Pluripotent Stem Cells: A Review, Analysis and Call for Innovation.
Smith L, Quelch-Cliffe R, Liu F, Aguilar AH, Przyborski S. Smith L, et al. Stem Cell Rev Rep. 2025 Jan;21(1):107-125. doi: 10.1007/s12015-024-10793-5. Epub 2024 Sep 28. Stem Cell Rev Rep. 2025. PMID: 39340737 Free PMC article. Review.
Cited by
- Efficient Generation of Hypothalamic Neurons from Human Pluripotent Stem Cells.
Wang L, Egli D, Leibel RL. Wang L, et al. Curr Protoc Hum Genet. 2016 Jul 1;90:21.5.1-21.5.14. doi: 10.1002/cphg.3. Curr Protoc Hum Genet. 2016. PMID: 27367166 Free PMC article. - Reprogramming Urine-Derived Cells using Commercially Available Self-Replicative RNA and a Single Electroporation.
Bouma MJ, Arendzen CH, Mummery CL, Mikkers H, Freund C. Bouma MJ, et al. Curr Protoc Stem Cell Biol. 2020 Dec;55(1):e124. doi: 10.1002/cpsc.124. Curr Protoc Stem Cell Biol. 2020. PMID: 32956580 Free PMC article. - Methods of induced pluripotent stem cells for clinical application.
Seki T, Fukuda K. Seki T, et al. World J Stem Cells. 2015 Jan 26;7(1):116-25. doi: 10.4252/wjsc.v7.i1.116. World J Stem Cells. 2015. PMID: 25621111 Free PMC article. Review. - The Decrease in Human Endogenous Retrovirus-H Activity Runs in Parallel with Improvement in ADHD Symptoms in Patients Undergoing Methylphenidate Therapy.
Cipriani C, Pitzianti MB, Matteucci C, D'Agati E, Miele MT, Rapaccini V, Grelli S, Curatolo P, Sinibaldi-Vallebona P, Pasini A, Balestrieri E. Cipriani C, et al. Int J Mol Sci. 2018 Oct 23;19(11):3286. doi: 10.3390/ijms19113286. Int J Mol Sci. 2018. PMID: 30360480 Free PMC article. - Recent policies that support clinical application of induced pluripotent stem cell-based regenerative therapies.
Azuma K, Yamanaka S. Azuma K, et al. Regen Ther. 2016 Mar 1;4:36-47. doi: 10.1016/j.reth.2016.01.009. eCollection 2016 Jun. Regen Ther. 2016. PMID: 31245486 Free PMC article. Review.
References
- Thomson JA, et al. Embryonic stem cell lines derived from human blastocysts. Science. 1998;282(5391):1145–1147. - PubMed
- Takahashi K, et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell. 2007;131(5):861–872. - PubMed
- Yu J, et al. Induced pluripotent stem cell lines derived from human somatic cells. Science. 2007;318(5858):1917–1920. - PubMed
- Newman AM, Cooper JB. Lab-specific gene expression signatures in pluripotent stem cells. Cell Stem Cell. 2010;7(2):258–262. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials