Receptor recognition mechanisms of coronaviruses: a decade of structural studies - PubMed (original) (raw)
Review
Receptor recognition mechanisms of coronaviruses: a decade of structural studies
Fang Li. J Virol. 2015 Feb.
Abstract
Receptor recognition by viruses is the first and essential step of viral infections of host cells. It is an important determinant of viral host range and cross-species infection and a primary target for antiviral intervention. Coronaviruses recognize a variety of host receptors, infect many hosts, and are health threats to humans and animals. The receptor-binding S1 subunit of coronavirus spike proteins contains two distinctive domains, the N-terminal domain (S1-NTD) and the C-terminal domain (S1-CTD), both of which can function as receptor-binding domains (RBDs). S1-NTDs and S1-CTDs from three major coronavirus genera recognize at least four protein receptors and three sugar receptors and demonstrate a complex receptor recognition pattern. For example, highly similar coronavirus S1-CTDs within the same genus can recognize different receptors, whereas very different coronavirus S1-CTDs from different genera can recognize the same receptor. Moreover, coronavirus S1-NTDs can recognize either protein or sugar receptors. Structural studies in the past decade have elucidated many of the puzzles associated with coronavirus-receptor interactions. This article reviews the latest knowledge on the receptor recognition mechanisms of coronaviruses and discusses how coronaviruses have evolved their complex receptor recognition pattern. It also summarizes important principles that govern receptor recognition by viruses in general.
Copyright © 2015, American Society for Microbiology. All Rights Reserved.
Figures
FIG 1
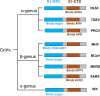
Receptor recognition pattern of coronaviruses.
FIG 2
Crystal structures of coronavirus S1-CTDs complexed with their respective receptor. (A to D) These structures include β-genus SARS-CoV S1-CTD complexed with ACE2 (Protein Data Bank identifier [PDB ID]: 2AJF) (76) (A), β-genus MERS-CoV S1-CTD complexed with DPP4 (PDB ID: 4KR0) (77) (B), α-genus HCoV-NL63 S1-CTD complexed with ACE2 (PDB ID: 3KBH) (79) (C), and α-genus PRCV S1-CTD complexed with APN (PDB ID: 4F5C) (80) (D). In the structures of the complexes, the receptors are in green, and the cores and RBMs of S1-CTDs are in cyan and red, respectively. (E and F) The structural topologies of the four coronavirus S1-CTDs are shown as schematic illustrations, where β-strands are depicted as arrows and α-helices as cylinders. In the tertiary structures and structural topologies of the S1-CTDs, the secondary structures of all of the S1-CTDs are colored and numbered in the same way as for HCoV-NL63 S1-CTD.
FIG 3
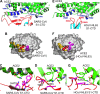
Virus-binding hot spots on ACE2 that are critical for the binding of SARS-CoV and HCoV-NL63. (A) Enlarged view of the SARS-CoV–ACE2 interface. VBMs on ACE2 and RBM on SARS-CoV S1-CTD are in blue and red, respectively. (B) Footprint of SARS-CoV on the surface of ACE2. The view is derived from the one in panel A by rotating ACE2 by 90° along a horizontal axis in such a way that the edge facing the viewer moves up. VBM1 residues are in orange, VBM2 residues in magenta, VBM3 residues in red, and VBM1b residues in green. (C) A virus-binding hot spot on ACE2 centering on Lys31 is critical for the binding of SARS-CoV S1-CTD. Mutation of residue 479 on SARS-CoV S1-CTD is critical for the transmission of SARS-CoV from palm civets to humans. (D) A second virus-binding hot spot on ACE2 centering on Lys353 is also critical for the binding of SARS-CoV S1-CTD. Mutation of residue 487 on SARS-CoV S1-CTD is critical for the transmission of SARS-CoV from human to human. (E) Enlarged view of the HCoV-NL63–ACE2 interface. (F) Footprint of HCoV-NL63 on the surface of ACE2. (G) The same virus-binding hot spot on ACE2 centering on Lys353 is also critical for the binding of HCoV-NL63 S1-CTD.
FIG 4
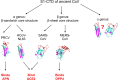
Proposed origin and evolution of coronavirus S1-CTDs. The question mark indicates possible tertiary structures of γ-coronavirus S1-CTD.
FIG 5
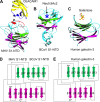
Crystal structures of coronavirus S1-NTDs. (A and B) These structures include β-genus MHV S1-NTD complexed with its receptor CEACAM1 (PDB ID: 3R4D) (81) (A) and β-genus BCoV S1-NTD by itself (PDB ID: 4H14) (53) (B). (C) The structure of human galectin-3 (PDB ID: 1A3K) is shown as a comparison. In the structure of the MHV S1-NTD-CEACAM1 complex, the VBM on CEACAM1 and the RBM on MHV S1-NTD are in blue and red, respectively. In the structure of BCoV S1-NTD, the sugar-binding pocket as identified by mutagenesis studies is indicated by a five-pointed star. (D and E) The structural topologies of the two coronavirus S1-NTDs (D) and human galectin-3 (E) are shown as schematic illustrations, where β-strands are depicted as arrows and α-helices as cylinders. In the structural topologies of these proteins, the secondary structures are colored and numbered in the same way as for the MHV S1-NTD.
FIG 6
Proposed origin and evolution of coronavirus S1-NTDs. Orange arrows indicate the locations of CEACAM1 or sugar that binds coronavirus NTDs. Question marks indicate the postulated structures of hypothetical evolutionary intermediates.
FIG 7
Summary of the receptor recognition mechanisms of coronaviruses in a three-dimensional view. The overall structure of trimeric SARS-CoV spike complexed with ACE2 is shown; it includes both the schematic topology of the spike and the negative-stain electron micropic images of the spike ectodomain (upper right). TM, transmembrane anchor. IC, intracellular tail. The structures and functions of coronavirus S1 domains are listed. The question marks indicate possible tertiary structures of coronavirus S1 domains.
Similar articles
- Cryo-Electron Microscopy Structure of Porcine Deltacoronavirus Spike Protein in the Prefusion State.
Shang J, Zheng Y, Yang Y, Liu C, Geng Q, Tai W, Du L, Zhou Y, Zhang W, Li F. Shang J, et al. J Virol. 2018 Feb 15;92(4):e01556-17. doi: 10.1128/JVI.01556-17. J Virol. 2018. PMID: 29070693 Free PMC article. - Structure of mouse coronavirus spike protein complexed with receptor reveals mechanism for viral entry.
Shang J, Wan Y, Liu C, Yount B, Gully K, Yang Y, Auerbach A, Peng G, Baric R, Li F. Shang J, et al. PLoS Pathog. 2020 Mar 9;16(3):e1008392. doi: 10.1371/journal.ppat.1008392. eCollection 2020 Mar. PLoS Pathog. 2020. PMID: 32150576 Free PMC article. - Structure, Function, and Evolution of Coronavirus Spike Proteins.
Li F. Li F. Annu Rev Virol. 2016 Sep 29;3(1):237-261. doi: 10.1146/annurev-virology-110615-042301. Epub 2016 Aug 25. Annu Rev Virol. 2016. PMID: 27578435 Free PMC article. Review. - Middle East respiratory syndrome coronavirus and bat coronavirus HKU9 both can utilize GRP78 for attachment onto host cells.
Chu H, Chan CM, Zhang X, Wang Y, Yuan S, Zhou J, Au-Yeung RK, Sze KH, Yang D, Shuai H, Hou Y, Li C, Zhao X, Poon VK, Leung SP, Yeung ML, Yan J, Lu G, Jin DY, Gao GF, Chan JF, Yuen KY. Chu H, et al. J Biol Chem. 2018 Jul 27;293(30):11709-11726. doi: 10.1074/jbc.RA118.001897. Epub 2018 Jun 10. J Biol Chem. 2018. PMID: 29887526 Free PMC article. - A structural view of coronavirus-receptor interactions.
Reguera J, Mudgal G, Santiago C, Casasnovas JM. Reguera J, et al. Virus Res. 2014 Dec 19;194:3-15. doi: 10.1016/j.virusres.2014.10.005. Epub 2014 Oct 14. Virus Res. 2014. PMID: 25451063 Free PMC article. Review.
Cited by
- [Comparative analysis of structural characteristics and epitopes in S proteins between SARS-CoV-2 and SARS-CoV].
Lun Y, Liu B, Dong W, Sun J, Pan L. Lun Y, et al. Zhejiang Da Xue Xue Bao Yi Xue Ban. 2020 May 25;49(3):315-323. doi: 10.3785/j.issn.1008-9292.2020.05.01. Zhejiang Da Xue Xue Bao Yi Xue Ban. 2020. PMID: 32762171 Free PMC article. Chinese. - Molecular Basis for Pathogenicity of Human Coronaviruses.
Pourrajab F, Zare-Khormizi MR, Sheikhha MH. Pourrajab F, et al. Infect Drug Resist. 2020 Jul 17;13:2385-2405. doi: 10.2147/IDR.S255156. eCollection 2020. Infect Drug Resist. 2020. PMID: 32765013 Free PMC article. Review. - Mouse Adapted SARS-CoV-2 protects animals from lethal SARS-CoV challenge.
Muruato A, Vu MN, Johnson BA, Davis-Gardner ME, Vanderheiden A, Lokugmage K, Schindewolf C, Crocquet-Valdes PA, Langsjoen RM, Plante JA, Plante KS, Weaver SC, Debbink K, Routh AL, Walker D, Suthar MS, Xie X, Shi PY, Xie X, Menachery VD. Muruato A, et al. bioRxiv [Preprint]. 2021 May 4:2021.05.03.442357. doi: 10.1101/2021.05.03.442357. bioRxiv. 2021. PMID: 33972939 Free PMC article. Updated. Preprint. - Molecular and cellular mechanisms involved in tissue-specific metabolic modulation by SARS-CoV-2.
Dos Santos AAC, Rodrigues LE, Alecrim-Zeza AL, de Araújo Ferreira L, Trettel CDS, Gimenes GM, da Silva AF, Sousa-Filho CPB, Serdan TDA, Levada-Pires AC, Hatanaka E, Borges FT, de Barros MP, Cury-Boaventura MF, Bertolini GL, Cassolla P, Marzuca-Nassr GN, Vitzel KF, Pithon-Curi TC, Masi LN, Curi R, Gorjao R, Hirabara SM. Dos Santos AAC, et al. Front Microbiol. 2022 Nov 10;13:1037467. doi: 10.3389/fmicb.2022.1037467. eCollection 2022. Front Microbiol. 2022. PMID: 36439786 Free PMC article. Review. - Investigation of Transmission and Evolution of PEDV Variants and Co-Infections in Northeast China from 2011 to 2022.
Zhao F, Ma X, Yang J, Wei Z, Li J, Jiang Y, Cui W, Shan Z, Tang L. Zhao F, et al. Animals (Basel). 2024 Jul 25;14(15):2168. doi: 10.3390/ani14152168. Animals (Basel). 2024. PMID: 39123693 Free PMC article.
References
- Ksiazek TG, Erdman D, Goldsmith CS, Zaki SR, Peret T, Emery S, Tong SX, Urbani C, Comer JA, Lim W, Rollin PE, Dowell SF, Ling AE, Humphrey CD, Shieh WJ, Guarner J, Paddock CD, Rota P, Fields B, DeRisi J, Yang JY, Cox N, Hughes JM, LeDuc JW, Bellini WJ, Anderson LJ. 2003. A novel coronavirus associated with severe acute respiratory syndrome. N Engl J Med 348:1953–1966. doi:10.1056/NEJMoa030781. - DOI - PubMed
- Peiris JSM, Lai ST, Poon LLM, Guan Y, Yam LYC, Lim W, Nicholls J, Yee WKS, Yan WW, Cheung MT, Cheng VCC, Chan KH, Tsang DNC, Yung RWH, Ng TK, Yuen KY. 2003. Coronavirus as a possible cause of severe acute respiratory syndrome. Lancet 361:1319–1325. doi:10.1016/S0140-6736(03)13077-2. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous