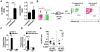Humoral Immunity in the Gut Selectively Targets Phenotypically Virulent Attaching-and-Effacing Bacteria for Intraluminal Elimination - PubMed (original) (raw)
Humoral Immunity in the Gut Selectively Targets Phenotypically Virulent Attaching-and-Effacing Bacteria for Intraluminal Elimination
Nobuhiko Kamada et al. Cell Host Microbe. 2015.
Abstract
Virulence factors expressed by enteric bacteria are pivotal for pathogen colonization and induction of intestinal disease, but the mechanisms by which host immunity regulates pathogen virulence are largely unknown. Here we show that specific antibody responses are required for downregulation of virulence gene expression in Citrobacter rodentium, an enteric pathogen that models human infections with attaching-and-effacing bacteria. In the absence of antibodies against the pathogen, phenotypically virulent C. rodentium, accumulated and infected the epithelium and subsequently invaded the lamina propia, causing host lethality. IgG induced after infection recognized virulence factors and bound virulent bacteria within the intestinal lumen, leading to their engulfment by neutrophils, while phenotypically avirulent pathogens remained in the intestinal lumen and were eventually outcompeted by the microbiota. Thus, the interplay of the innate and adaptive immune system selectively targets virulent C. rodentium in the intestinal lumen to promote pathogen eradication and host survival.
Copyright © 2015 Elsevier Inc. All rights reserved.
Figures
Figure. 1. Host adaptive immunity is required for down-regulation of LEE virulence during C. rodentium infection
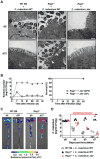
A, B, SPF (A) and GF (B) WT B6 and _Rag1_-/- mice (n= 7) were infected orally with 1×109 cfu of C. rodentium and pathogen load in feces (left) and mouse survival (right) were determined over the indicated time. Data points are given as mean ± SD. Results are representative of at least 3 independent experiments. † denotes bacterial loads could not be determined beyond this time due to mouse lethality. **; p<0.01 by Log-rank test. C, Bioluminescent imaging of ler expression in the intestines of GF WT and _Rag1_-/- mice infected with the ler-lux C. rodentium reporter strain. Imaging was performed on day 5 and 14 post-infection and the signal was quantified based on the color scale shown. Results are representative of 3 individual mice. D, Expression of ler (left) and bacterial burden (right) in fecal pellets of GF mice infected with the reporter ler-lux C. rodentium strain at the indicated day post-infection (dpi). Results show luminescence of ler-lux (relative light units) and C. rodentium cfu in the same samples. Data expressed as mean ± SD of individual mice (n=3). Results are representative of at least 2 experiments. *, p<0.05; N.S., not significant by Student's t test. See also Figure S1.
Figure. 2. Inappropriate retention of mucosa-associated LEE virulence and pathogen invasion in mice lacking adaptive immunity
A, GF WT and Rag1_-/- deficient mice were infected with WT and Δ_ler mutant C. rodentium. On day 5 (top) or day 21 (bottom) post-infection, mucosa-associated bacteria in ceca were assessed by Transmission Electron Microscopy. Original magnification: 13,500×. Scale bar: 1 μm. Results are representative of 2 experiments. B, SPF and GF _Rag1_-/- mice (n= 5) were infected orally with 1×109 cfu of C. rodentium and pathogen load in feces (left) and mouse survival (right) were determined over the indicated time. Data points are given as mean ± SD. Results are representative of at least 2 independent experiments. C, GF WT and _Rag1_-/- mice were infected with the ler-lux C. rodentium strain for 5 or 21 days. Cecum and colonic tissues were collected at the indicated day and then washed with PBS to remove non-adherent bacteria. Bioluminescent imaging of ler expression of C. rodentium attached to the cecum (top) and colon (bottom). Results are representative of 2 experiments using 3-4 different mice. D, GF WT and _Rag1_-/- mice were infected with WT and Δler mutant C. rodentium. On day 10 post infection, mice were co-housed with SPF mice to transfer microbiota (1:1). Pathogen load was determined in feces on indicated days after co-housing. Dots represent individual mice. See also Figure S2.
Figure. 3. Pathogen-specific IgG responses are required for downregulation of LEE virulence expression
A, Production of total IgG and IgA against C. rodentium in the serum and luminal content of QM mice before (d0) and after (day 14) oral infection with C. rodentium. Dots represent individual mice. *, p<0.05; ***, p<0.001; N.S., not significant by Dunn's test. B, WT and QM mice were infected with C. rodentium and pathogen load in feces were determined over the indicated time. Dots represent individual mice and pooled 3 independent experiments. C, WT and QM mice were infected with the ler-lux C. rodentium strain for 5 or 14 days. Cecum and colonic tissues were collected at the indicated day and then washed with PBS to remove non-adherent bacteria. Bioluminescent imaging of ler expression of C. rodentium attached to the cecum (top) and colon (bottom). Results are representative of 3 independent experiments. See also Figure S3.
Figure. 4. Virulence factor-specific IgG is induced in the late phase of C. rodentium infection
A, GF WT mice were infected with GFP-expressing C. rodentium. Cecal bacteria were harvested at indicated days post-infection and binding of IgG and IgA was analyzed by flow cytometry. Results are representative of 3 experiments. B, Bacterial lysates of WT and Δler mutant C. rodentium were loaded with SDS-PAGE. For confirmation of amount of loaded protein, gels were stained with silver staining reagent (left panel). Serum or luminal content were obtained from naïve (d0) and _C. rodentium_-infected (d21) GF mice, and used as primary antibodies. _C. rodentium_-specific IgG was detected by anti-mouse IgG secondary Ab. C, WT (GFP+) and Δler mutant (GFP-) C. rodentium were cultured in DMEM for 6 hrs. Cultured bacteria were then washed with PBS and mixed at 1:1 ratio. Mixed bacteria were then incubated with 5% of serum from naïve and C. rodentium pre-infected (day 21 post-infection) GF mice for 30 min on ice. After incubation, bacteria were washed and IgG-binding to the bacteria was detected by biotin-conjugated anti-mouse IgG antibody and streptavidin-APC. Rat IgG was used as a control staining. IgG binding to each bacterial strain was analyzed by flow cytometry. Results are representative of 2 independent experiments. See also Figure S4.
Figure 5. Targeting of LEE virulence by IgG promotes selective elimination of phenotypically virulent pathogens
A, GF WT mice were infected with C. rodentium. At day 7 post-infection, mucosal-associated and luminal bacteria were harvested from the cecum (n=10) and ler and rpoB mRNA levels were determined by qPCR. Expression was normalized to that of the 16S rRNA gene rrsA. Results are given as mean ± SD. ***; p<0.001; N.S., not significant by Mann-Whitney U test. B, GF WT mice were infected with 1:1 ratio of WT (GFP+) and Δ_ler_ mutant (GFP-) C. rodentium. After 21 days, cecal bacteria were harvested and binding of IgG was analyzed by flow cytometry. Results are representative of 3 experiments. C, WT (ChlR) and Δler (KanR) C. rodentium were mixed at a 1:1 ratio (1×107 cfu in 200 ml PBS), and injected into the peritoneal cavity of naïve and C. rodentium infected (d21 post infection) mice that had been pre-injected with thioglycollate to induce neutrophil recruitment into the peritoneal cavity. Neutrophils were harvested from the peritoneal cavity 30 min after injection of bacteria, and numbers of WT and Δler bacteria engulfed by neutrophils were assessed. Data are given as mean ± SD (n=3). Results are representative of 2 experiments. D, WT (ChlR) and Δler (KanR) C. rodentium were mixed at a 1:1 ratio, and then incubated with serum from naïve or C. rodentium infected (d21 post infection) GF WT mice for 30 min on ice. After washing, serum-treated bacteria were injected into the peritoneal cavity of naïve WT mice pre-injected with thioglycollate. The numbers of engulfed bacteria by neutrophils were assessed as described in (C). Data are given as mean ± SD (n=3). Results are representative of 2 experiments. ***; p<0.001 by Student's t test. E, GF WT and Rag1_-/- mice were infected with 1:1 ratio of WT (ChlR) and Δler mutant (KanR) C. rodentium. After 3 and 21 days, intestinal burden of WT and mutant bacteria was measured. Dots represent colonization index (CI; ratio of WT/Δ_ler) of individual mice and pooled 3 independent experiments with 3 mice. *; p<0.05, ***; p<0.001 by Mann-Whitney U test.
Figure. 6. Neutrophils elicit selective elimination of opsonized virulent bacteria
A, SPF WT mice were infected with GFP-expressing C. rodentium. At day 11 post infection, cecal content was harvested. Luminal debris and bacteria were removed by filtering and centrifugation, and then stained with antibodies for CD45, CD11b, Ly6G, and 7-AAD and analyzed by flow cytometry. Data are representative of 3 independent experiments. B, LysMCreMcl1wt/wt (control) and LysMCreMcl1fl/fl (neutrophil deficient) chimeric mice (n= 16; Mcl1wt/wt, n=9; Mcl1fl/fl) were infected with C. rodentium and mouse mortality was determined over the indicated time. ***; p<0.001 by Log-rank test. C, LysMCreMcl1wt/wt and LysMCreMcl1fl/fl chimeric mice were infected with C. rodentium and luminal emigration of neutrophils was analyzed on day 5 post-infection. 7-AAD-CD45+CD11b+ cells in luminal content were gated and further analyzed for Ly6G expression. Results are representative of 2 individual mice. D, LysMCreMcl1wt/wt and LysMCreMcl1fl/fl chimeric mice were infected ler-lux C. rodentium strain for 5 or 8 days. Cecum and colonic tissues were collected at the indicated day and then washed with PBS to remove non-adherent bacteria. Bioluminescent imaging of ler expression in C. rodentium attached to the cecum (top) and colon (bottom). Results are representative of 3 independent experiments. See also Figure S5.
Comment in
- IgG "detoxes" the intestinal mucosa.
Eckmann L, Stappenbeck TS. Eckmann L, et al. Cell Host Microbe. 2015 May 13;17(5):538-9. doi: 10.1016/j.chom.2015.05.001. Cell Host Microbe. 2015. PMID: 25974292 - Host response: Sifting out virulent bacteria.
Attar N. Attar N. Nat Rev Microbiol. 2015 Jun;13(6):330. doi: 10.1038/nrmicro3499. Nat Rev Microbiol. 2015. PMID: 25978704 No abstract available.
Similar articles
- IgG "detoxes" the intestinal mucosa.
Eckmann L, Stappenbeck TS. Eckmann L, et al. Cell Host Microbe. 2015 May 13;17(5):538-9. doi: 10.1016/j.chom.2015.05.001. Cell Host Microbe. 2015. PMID: 25974292 - Host response: Sifting out virulent bacteria.
Attar N. Attar N. Nat Rev Microbiol. 2015 Jun;13(6):330. doi: 10.1038/nrmicro3499. Nat Rev Microbiol. 2015. PMID: 25978704 No abstract available. - Regulated virulence controls the ability of a pathogen to compete with the gut microbiota.
Kamada N, Kim YG, Sham HP, Vallance BA, Puente JL, Martens EC, Núñez G. Kamada N, et al. Science. 2012 Jun 8;336(6086):1325-9. doi: 10.1126/science.1222195. Epub 2012 May 10. Science. 2012. PMID: 22582016 Free PMC article. - Citrobacter rodentium-host-microbiota interactions: immunity, bioenergetics and metabolism.
Mullineaux-Sanders C, Sanchez-Garrido J, Hopkins EGD, Shenoy AR, Barry R, Frankel G. Mullineaux-Sanders C, et al. Nat Rev Microbiol. 2019 Nov;17(11):701-715. doi: 10.1038/s41579-019-0252-z. Epub 2019 Sep 20. Nat Rev Microbiol. 2019. PMID: 31541196 Review. - Citrobacter rodentium: a model enteropathogen for understanding the interplay of innate and adaptive components of type 3 immunity.
Silberger DJ, Zindl CL, Weaver CT. Silberger DJ, et al. Mucosal Immunol. 2017 Sep;10(5):1108-1117. doi: 10.1038/mi.2017.47. Epub 2017 Jun 14. Mucosal Immunol. 2017. PMID: 28612839 Free PMC article. Review.
Cited by
- Virulence factors enhance Citrobacter rodentium expansion through aerobic respiration.
Lopez CA, Miller BM, Rivera-Chávez F, Velazquez EM, Byndloss MX, Chávez-Arroyo A, Lokken KL, Tsolis RM, Winter SE, Bäumler AJ. Lopez CA, et al. Science. 2016 Sep 16;353(6305):1249-53. doi: 10.1126/science.aag3042. Epub 2016 Sep 15. Science. 2016. PMID: 27634526 Free PMC article. - Glycan-based shaping of the microbiota during primate evolution.
Singh S, Bastos-Amador P, Thompson JA, Truglio M, Yilmaz B, Cardoso S, Sobral D, Soares MP. Singh S, et al. Elife. 2021 May 19;10:e67450. doi: 10.7554/eLife.67450. Elife. 2021. PMID: 34009123 Free PMC article. - A subpopulation of high IL-21-producing CD4(+) T cells in Peyer's Patches is induced by the microbiota and regulates germinal centers.
Jones L, Ho WQ, Ying S, Ramakrishna L, Srinivasan KG, Yurieva M, Ng WP, Subramaniam S, Hamadee NH, Joseph S, Dolpady J, Atarashi K, Honda K, Zolezzi F, Poidinger M, Lafaille JJ, Curotto de Lafaille MA. Jones L, et al. Sci Rep. 2016 Aug 8;6:30784. doi: 10.1038/srep30784. Sci Rep. 2016. PMID: 27499025 Free PMC article. - Twin-Arginine Translocation System Is Involved in Citrobacter rodentium Fitness in the Intestinal Tract.
Otake T, Fujimoto M, Hoshino Y, Ishihara T, Haneda T, Okada N, Miki T. Otake T, et al. Infect Immun. 2020 Feb 20;88(3):e00892-19. doi: 10.1128/IAI.00892-19. Print 2020 Feb 20. Infect Immun. 2020. PMID: 31818958 Free PMC article. - Cooperation between physiological defenses and immune resistance produces asymptomatic carriage of a lethal bacterial pathogen.
Chen GY, Thorup NR, Miller AJ, Li YC, Ayres JS. Chen GY, et al. bioRxiv [Preprint]. 2023 Feb 26:2023.01.22.525099. doi: 10.1101/2023.01.22.525099. bioRxiv. 2023. PMID: 36711884 Free PMC article. Updated. Preprint.
References
- Bry L, Brenner MB. Critical role of T cell-dependent serum antibody, but not the gut-associated lymphoid tissue, for surviving acute mucosal infection with Citrobacter rodentium, an attaching and effacing pathogen. J Immunol. 2004;172:433–441. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- P30 CA046592/CA/NCI NIH HHS/United States
- T32DK094775/DK/NIDDK NIH HHS/United States
- T32 HL007517/HL/NHLBI NIH HHS/United States
- 5P30DK034933/DK/NIDDK NIH HHS/United States
- P30 DK034933/DK/NIDDK NIH HHS/United States
- R01 DK095782/DK/NIDDK NIH HHS/United States
- T32 DK094775/DK/NIDDK NIH HHS/United States
- NIH DK095782/DK/NIDDK NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources