Epigenetic Mechanisms in Developmental Alcohol-Induced Neurobehavioral Deficits - PubMed (original) (raw)
Review
Epigenetic Mechanisms in Developmental Alcohol-Induced Neurobehavioral Deficits
Balapal S Basavarajappa et al. Brain Sci. 2016.
Abstract
Alcohol consumption during pregnancy and its damaging consequences on the developing infant brain are significant public health, social, and economic issues. The major distinctive features of prenatal alcohol exposure in humans are cognitive and behavioral dysfunction due to damage to the central nervous system (CNS), which results in a continuum of disarray that is collectively called fetal alcohol spectrum disorder (FASD). Many rodent models have been developed to understand the mechanisms of and to reproduce the human FASD phenotypes. These animal FASD studies have provided several molecular pathways that are likely responsible for the neurobehavioral abnormalities that are associated with prenatal alcohol exposure of the developing CNS. Recently, many laboratories have identified several immediate, as well as long-lasting, epigenetic modifications of DNA methylation, DNA-associated histone proteins and microRNA (miRNA) biogenesis by using a variety of epigenetic approaches in rodent FASD models. Because DNA methylation patterns, DNA-associated histone protein modifications and miRNA-regulated gene expression are crucial for synaptic plasticity and learning and memory, they can therefore offer an answer to many of the neurobehavioral abnormalities that are found in FASD. In this review, we briefly discuss the current literature of DNA methylation, DNA-associated histone proteins modification and miRNA and review recent developments concerning epigenetic changes in FASD.
Keywords: DNA and histone modification; FAS; FASD; Learning and memory; Synaptic plasticity.
Figures
Figure 1
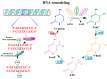
Graphic representation of DNA methylation and its regulation by enzymatic mechanisms. Methylation of DNA begins with the covalent addition of a methyl group from s-adenyl methionine (SAM) [41] to the fifth carbon of the cytosine pyrimidine ring to form 5-methylcytosine (5-mC), a process that is catalyzed by a family of DNA methyltransferases (DNMTs). The majority of DNA methylation usually occurs at genes on cytosines that precede a guanine nucleotide or CpG islands. De novo methyltransferases (e.g., DNMT3a/b) transfer methyl groups to naked DNA CpG pairs (e.g., CpG/GpC to mCpG/GpC) [42,43]. DNMT1 is the maintenance methyltransferase that transfers methyl groups to hemimethylated DNA strands (e.g., mCpG/GpC to mCpG/GpCm) and maintains the parental DNA methylation pattern during replication [44]. 5-mC undergoes sequential oxidation to 5-caC by TET1 activities. 5-caC, through base-excision-repair (BER) mechanisms, results in the regeneration of cytosine [39,45,46]. 5-methylcytosine (5-mC); 5-hydroxymethylcytosine (5-hmC); 5-formylcytosine (5-fC); 5-carboxylcytosine (5-caC).
Figure 2
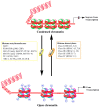
Schematic representation of DNA-associated histone protein acetylation and deacetylation by histone acetyltransferase (HAT) and histone deacetylase (HDAC) enzyme families. The net acetylation at lysine residues of histone proteins of nucleosomes is decided by the interplay between acetylation and deacetylation processes through HAT and HDAC enzyme activities, respectively. The box provides different families and classes of HAT and HDAC enzymes. CBP, cyclic adenomonophosphate response element-binding (CREB) binding protein; GNAT, Gcn5-related _N_-acetyltransferases; hGCN5, human general control of amino acid synthesis protein 5-like 2; PCAF, p300/CBP-associated factor; ELP3, elongation protein 3; TIP60, TAT interacting proteins 60; TFIIIC90, transcription factor IIIC 90kDa; TAF1, TATA Box Binding Protein-Associated Factor; SRC1, steroid receptor coactivator 1; ACTR, activator of thyroid receptor; p160, receptor coactivators proteins 160; CLOCK, Clock Circadian Regulator.
Figure 3
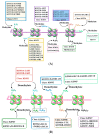
DNA-associated histone protein lysine methylation (A) and demethylation (B) by histone methyltransferase (KMTs) and histone demethylase (KDMs) enzyme families. Histone H3 and H4 tails with known lysine methyltransferases (KMT1-8) and demethylases (KDM1-7) are shown under each specific lysine residue.
Figure 4
The schematic diagram of miRNA biogenesis and function. In the nucleus, RNA polymerase II (Pol II)-dependent transcription of a miRNA-encoding gene, which may include both intron- and exon-coding regions, results in the formation of a long primary miRNA transcript (pri-miRNA) that is 50-capped and 30-polyadenylated. This pri-miRNA transcript is subject to nuclear processing by the microprocessor complex, which includes DCGR8 and Drosha, into the precursor miRNA (pre-miRNA) transcript. Pre-miRNAs are transported out of the nucleus to the cytoplasm by Exp5 and Ran-GTP. Pre-miRNA can then be further cleaved by Dicer/TARBP to generate a mature miRNA. The mature miRNA is incorporated into the RNA-induced silencing complex (RISC), which will select mRNA transcripts for the down-regulation of protein expression. In the synapses, miRNA can regulate the local down-regulation of protein expression. Certain conditions, such as neuronal activity, could affect the signaling events as well as miRNA formation. Additionally, genes regulated by miRNA can act on synaptic activity-dependent signaling pathways that promote the activation of epigenetic factors (e.g., CREB and MeCP2), which in turn can control miRNA transcription in the nucleus.
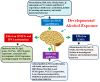
Figure 5
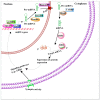
Graphical summary of developmental alcohol-induced epigenetic defects. Developmental alcohol exposure has been shown to affect DNA methylation, HATs/HDACs, KMTs and miRNAs, followed by several changes in genes and protein expression that are important for cognitive and other brain function.
Similar articles
- The Impact of Oxidative Stress on the Epigenetics of Fetal Alcohol Spectrum Disorders.
Terracina S, Tarani L, Ceccanti M, Vitali M, Francati S, Lucarelli M, Venditti S, Verdone L, Ferraguti G, Fiore M. Terracina S, et al. Antioxidants (Basel). 2024 Mar 28;13(4):410. doi: 10.3390/antiox13040410. Antioxidants (Basel). 2024. PMID: 38671857 Free PMC article. Review. - Synaptic Plasticity Abnormalities in Fetal Alcohol Spectrum Disorders.
Basavarajappa BS, Subbanna S. Basavarajappa BS, et al. Cells. 2023 Jan 29;12(3):442. doi: 10.3390/cells12030442. Cells. 2023. PMID: 36766783 Free PMC article. Review. - Fetal Alcohol Spectrum Disorder: Potential Role of Endocannabinoids Signaling.
Basavarajappa BS. Basavarajappa BS. Brain Sci. 2015 Oct 29;5(4):456-93. doi: 10.3390/brainsci5040456. Brain Sci. 2015. PMID: 26529026 Free PMC article. Review. - Overview of the Genetic Basis and Epigenetic Mechanisms that Contribute to FASD Pathobiology.
Liyanage VR, Curtis K, Zachariah RM, Chudley AE, Rastegar M. Liyanage VR, et al. Curr Top Med Chem. 2017;17(7):808-828. doi: 10.2174/1568026616666160414124816. Curr Top Med Chem. 2017. PMID: 27086780 Review. - Changes to histone modifications following prenatal alcohol exposure: An emerging picture.
Chater-Diehl EJ, Laufer BI, Singh SM. Chater-Diehl EJ, et al. Alcohol. 2017 May;60:41-52. doi: 10.1016/j.alcohol.2017.01.005. Epub 2017 Feb 4. Alcohol. 2017. PMID: 28431792 Review.
Cited by
- The Impact of Oxidative Stress on the Epigenetics of Fetal Alcohol Spectrum Disorders.
Terracina S, Tarani L, Ceccanti M, Vitali M, Francati S, Lucarelli M, Venditti S, Verdone L, Ferraguti G, Fiore M. Terracina S, et al. Antioxidants (Basel). 2024 Mar 28;13(4):410. doi: 10.3390/antiox13040410. Antioxidants (Basel). 2024. PMID: 38671857 Free PMC article. Review. - Alcohol Feeding in Mice Promotes Colonic Hyperpermeability and Changes in Colonic Organoid Stem Cell Fate.
Forsyth CB, Shaikh M, Bishehsari F, Swanson G, Voigt RM, Dodiya H, Wilkinson P, Samelco B, Song S, Keshavarzian A. Forsyth CB, et al. Alcohol Clin Exp Res. 2017 Dec;41(12):2100-2113. doi: 10.1111/acer.13519. Epub 2017 Nov 7. Alcohol Clin Exp Res. 2017. PMID: 28992396 Free PMC article. - CB1R regulates CDK5 signaling and epigenetically controls Rac1 expression contributing to neurobehavioral abnormalities in mice postnatally exposed to ethanol.
Joshi V, Subbanna S, Shivakumar M, Basavarajappa BS. Joshi V, et al. Neuropsychopharmacology. 2019 Feb;44(3):514-525. doi: 10.1038/s41386-018-0181-y. Epub 2018 Aug 22. Neuropsychopharmacology. 2019. PMID: 30143782 Free PMC article. - Neonatal Ethanol Disturbs the Normal Maturation of Parvalbumin Interneurons Surrounded by Subsets of Perineuronal Nets in the Cerebral Cortex: Partial Reversal by Lithium.
Saito M, Smiley JF, Hui M, Masiello K, Betz J, Ilina M, Saito M, Wilson DA. Saito M, et al. Cereb Cortex. 2019 Apr 1;29(4):1383-1397. doi: 10.1093/cercor/bhy034. Cereb Cortex. 2019. PMID: 29462278 Free PMC article. - DNA modifications in models of alcohol use disorders.
Tulisiak CT, Harris RA, Ponomarev I. Tulisiak CT, et al. Alcohol. 2017 May;60:19-30. doi: 10.1016/j.alcohol.2016.11.004. Epub 2016 Nov 9. Alcohol. 2017. PMID: 27865607 Free PMC article. Review.
References
- Kumar A., Choi K.H., Renthal W., Tsankova N.M., Theobald D.E., Truong H.T., Russo S.J., Laplant Q., Sasaki T.S., Whistler K.N., et al. Chromatin remodeling is a key mechanism underlying cocaine-induced plasticity in striatum. Neuron. 2005;48:303–314. doi: 10.1016/j.neuron.2005.09.023. - DOI - PubMed
- Madrigano J., Baccarelli A., Mittleman M.A., Wright R.O., Sparrow D., Vokonas P.S., Tarantini L., Schwartz J. Prolonged exposure to particulate pollution, genes associated with glutathione pathways, and DNA methylation in a cohort of older men. Environ. Health Perspect. 2011;119:977–982. doi: 10.1289/ehp.1002773. - DOI - PMC - PubMed
- Subbanna S., Nagaraja N.N., Umapathy N.S., Pace B.S., Basavarajappa B.S. Ethanol exposure induces neonatal neurodegeneration by enhancing CB1R Exon1 histone H4K8 acetylation and up-regulating CB1R function causing neurobehavioral abnormalities in adult mice. Int. J. Neuropsychopharmacol. 2015 doi: 10.1093/ijnp/pyu028. - DOI - PMC - PubMed
- Subbanna S., Nagre N.N., Shivakumar M., Umapathy N.S., Psychoyos D., Basavarajappa B.S. Ethanol induced acetylation of histone at G9a Exon1 and G9a-mediated histone H3 dimethylation leads to neurodegeneration in neonatal mice. Neuroscience. 2014;258:422–432. doi: 10.1016/j.neuroscience.2013.11.043. - DOI - PMC - PubMed
Publication types
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous