Mode of Ezrin-Membrane Interaction as a Function of PIP2 Binding and Pseudophosphorylation - PubMed (original) (raw)
Mode of Ezrin-Membrane Interaction as a Function of PIP2 Binding and Pseudophosphorylation
Victoria Shabardina et al. Biophys J. 2016.
Abstract
Ezrin, a protein of the ezrin, radixin, moesin (ERM) family, provides a regulated linkage between the plasma membrane and the cytoskeleton. The hallmark of this linkage is the activation of ezrin by phosphatidylinositol-4,5-bisphosphate (PIP2) binding and a threonine phosphorylation at position 567. To analyze the influence of these activating factors on the organization of ezrin on lipid membranes and the proposed concomitant oligomer-monomer transition, we made use of supported lipid bilayers in conjunction with atomic force microscopy and fluorescence microscopy. Bilayers doped with either PIP2 as the natural receptor lipid of ezrin or a Ni-nitrilotriacetic acid-equipped lipid to bind the proteins via their His6-tags to the lipid membrane were used to bind two different ezrin variants: ezrin wild-type and ezrin T567D mimicking the phosphorylated state. Using a combination of reflectometric interference spectroscopy, atomic force microscopy, and Förster resonance energy transfer experiments, we show that only the ezrin T567D mutant, upon binding to PIP2-containing bilayers, undergoes a remarkable conformational change, which we attribute to an opening of the conformation resulting in monomeric protein on the lipid bilayer.
Copyright © 2016 Biophysical Society. Published by Elsevier Inc. All rights reserved.
Figures
Figure 1
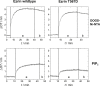
Binding of ezrin wt and ezrin T567D to SLBs composed of DOPC/DOGS-Ni-NTA or POPC/PIP2 (96:4), analyzed by RIfS. (a) Addition of 0.8 _μ_M protein solution leads to an increase in OT. (b) After saturation, the surface is rinsed with buffer.
Figure 2
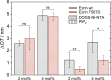
Results of the RIfS experiments. The change in OT is plotted for ezrin wt (gray) and ezrin T567D (red) bound to either 2 or 4 mol % DOGS-Ni-NTA (crosswise striated) or PIP2 (longitudinally striated). Three to eight independent experiments were performed in each case. The error bars are the standard deviation of the mean (∗p < 0.04, ∗∗p < 0.01; ns, not significant; two-sample (nonpaired) Student’s _t_-test). To see this figure in color, go online.
Figure 3
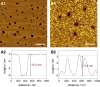
Atomic force micrographs of an SLB before and after protein incubation. (A) Atomic force micrograph of a lipid bilayer (DPPC/PIP2, 97:3) on a silicon/silicon dioxide substrate (A1). The corresponding line profile (A2) along the blue line shown in the topography image (A1) demonstrates the relative height of a single bilayer (interface set to 0 nm) with respect to a membrane defect. (B) Atomic force micrograph of a lipid bilayer (DPPC/PIP2, 97:3) that was incubated with N-ERMAD (0.5 _μ_M, 15 h) and subsequently rinsed with buffer (B1). The height profile of the protein clusters (B2) along the blue line shown in the topography image (B1) demonstrates the height difference between the lipid bilayer (interface set to 0 nm) and the adsorbed protein clusters. Scale bars, 1 _μ_m. To see this figure in color, go online.
Figure 4
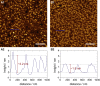
Atomic force micrographs of SLBs (1) composed of DPPC/PIP2 (97:3) on silicon/silicon dioxide substrates, which were incubated with (A) ezrin wt (0.5 μM) or (B) ezrin T567D (0.5 μM). The SLBs were rinsed with buffer before imaging. The corresponding line profiles of protein clusters (2) marked in the topography image (blue line) show a significantly lower protein height for ezrin T567D. Scale bars: 1 μm. To see this figure in color, go online.
Figure 5
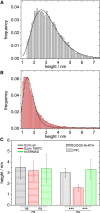
Summary of results obtained after height analysis of atomic force micrographs. (A and B) Histograms obtained for (A) ezrin wt bound to DPPC/PIP2 (97:3) and (B) ezrin T567D bound to DPPC/PIP2 (97:3). (C) Summary of protein heights determined by AFM images according to the procedure described in the Materials and Methods section for ezrin wt (gray), ezrin T567D (red), and N-ERMAD (green) bound to PIP2 (longitudinally striated) or DOGS-Ni-NTA (crosswise striated). The error bars are the standard deviation of the mean. The significance of the data was analyzed with Student’s t_-tests (∗∗∗_p < 0.001; ns, not significant). To see this figure in color, go online.
Figure 6
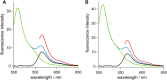
(A and B) Fluorescence spectra of (A) ezrin wt and (B) ezrinT567D. Ezrin-eGFP was excited at 488 nm (green line), and Cy3-ezrin-eGFP was excited at 488 nm (blue line) and 550 nm (red line, fluorescence intensity divided by 5). The black line is the difference spectrum of the spectra shown in blue and green. To see this figure in color, go online.
Figure 7
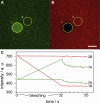
Acceptor bleaching of Cy3-ezrin wt-eGFP bound to a SLB composed of DOPC/DOGS-Ni-NTA (96:4). (A) eGFP, fluorescence, (B) Cy3 fluorescence. The yellow circle 1 marks the ROI after bleaching of Cy3, circle 2 marks the reference ROI. Scale bar: 5 μm. (C) Time trace of the bleaching process showing the increase in eGFP fluorescence (green line, 1A) and the decrease in Cy3 fluorescence (red line, 1B), numbers 1 and 2 correspond to bleached ROI and reference ROI, respectively. To see this figure in color, go online.
Figure 8
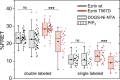
Summary of the FRET analysis of SLB-bound ezrin. The double-labeled ezrin constructs (left: wt, Cy3-ezrin wt-eGFP; T567D, Cy3-ezrin T567D-eGFP) and 1:1 mixtures of single-labeled protein constructs (right: wt, Cy3-ezrin wt/ezrin wt-eGFP; T567D, Cy3-ezrin T567D/ezrin T567D-eGFP) were bound to DOGS-Ni-NTA (crosswise striated) or PIP2 (longitudinally striated) containing lipid bilayers and the FRET efficiency was measured by acceptor bleaching. The solid line in each box is the median and the dot marks the mean value. Each data point is plotted next to the respective box (black/red dots). The FRET efficiency (%FRET, Eq. 1) was calculated using FRETcalc in ImageJ. The statistical significance of the data was analyzed with a nonparametric Mann-Whitney U-test, which showed differences for ezrin T567D with ∗∗∗p < 0.001, and nonsignificant ones (ns) for ezrin wt. To see this figure in color, go online.
Figure 9
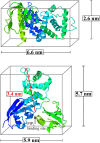
Crystallographic dimensions of N-ERMAD composed of three subdomains (F1, F2, and F3). The top image shows the minimum and maximum dimensions of N-ERMAD. The bottom image shows the possible orientation of N-ERMAD bound to PIP2, resulting in the observed height of 3.4 nm. To see this figure in color, go online.
Figure 10
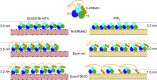
Model of different binding scenarios for ezrin composed of N-ERMAD (F1, F2, and F3) and C-ERMAD on an SLB, demonstrating a more loosely packed ezrin monolayer in the case of the fully activated state (ezrin T567D on PIP2). To see this figure in color, go online.
Similar articles
- Quantitative analysis of the binding of ezrin to large unilamellar vesicles containing phosphatidylinositol 4,5 bisphosphate.
Blin G, Margeat E, Carvalho K, Royer CA, Roy C, Picart C. Blin G, et al. Biophys J. 2008 Feb 1;94(3):1021-33. doi: 10.1529/biophysj.107.110213. Epub 2007 Sep 7. Biophys J. 2008. PMID: 17827228 Free PMC article. - Phosphatidylinositol 4,5-bisphosphate alters the number of attachment sites between ezrin and actin filaments: a colloidal probe study.
Braunger JA, Brückner BR, Nehls S, Pietuch A, Gerke V, Mey I, Janshoff A, Steinem C. Braunger JA, et al. J Biol Chem. 2014 Apr 4;289(14):9833-43. doi: 10.1074/jbc.M113.530659. Epub 2014 Feb 5. J Biol Chem. 2014. PMID: 24500715 Free PMC article. - Actin binding of ezrin is activated by specific recognition of PIP2-functionalized lipid bilayers.
Janke M, Herrig A, Austermann J, Gerke V, Steinem C, Janshoff A. Janke M, et al. Biochemistry. 2008 Mar 25;47(12):3762-9. doi: 10.1021/bi702542s. Epub 2008 Feb 27. Biochemistry. 2008. PMID: 18302339 - Cooperative adsorption of ezrin on PIP2-containing membranes.
Herrig A, Janke M, Austermann J, Gerke V, Janshoff A, Steinem C. Herrig A, et al. Biochemistry. 2006 Oct 31;45(43):13025-34. doi: 10.1021/bi061064a. Biochemistry. 2006. PMID: 17059219 - Activation of F-actin binding capacity of ezrin: synergism of PIP₂ interaction and phosphorylation.
Bosk S, Braunger JA, Gerke V, Steinem C. Bosk S, et al. Biophys J. 2011 Apr 6;100(7):1708-17. doi: 10.1016/j.bpj.2011.02.039. Biophys J. 2011. PMID: 21463584 Free PMC article.
Cited by
- Ezrin Phosphorylation at T567 Modulates Cell Migration, Mechanical Properties, and Cytoskeletal Organization.
Zhang X, Flores LR, Keeling MC, Sliogeryte K, Gavara N. Zhang X, et al. Int J Mol Sci. 2020 Jan 9;21(2):435. doi: 10.3390/ijms21020435. Int J Mol Sci. 2020. PMID: 31936668 Free PMC article. - Role of Phosphorylation in Moesin Interactions with PIP2-Containing Biomimetic Membranes.
Lubart Q, Vitet H, Dalonneau F, Le Roy A, Kowalski M, Lourdin M, Ebel C, Weidenhaupt M, Picart C. Lubart Q, et al. Biophys J. 2018 Jan 9;114(1):98-112. doi: 10.1016/j.bpj.2017.10.041. Biophys J. 2018. PMID: 29320700 Free PMC article. - Monoallelic loss of the F-actin-binding protein radixin facilitates startle reactivity and pre-pulse inhibition in mice.
Hausrat TJ, Vogl C, Neef J, Schweizer M, Yee BK, Strenzke N, Kneussel M. Hausrat TJ, et al. Front Cell Dev Biol. 2022 Nov 28;10:987691. doi: 10.3389/fcell.2022.987691. eCollection 2022. Front Cell Dev Biol. 2022. PMID: 36518539 Free PMC article. - A biophysical perspective of the regulatory mechanisms of ezrin/radixin/moesin proteins.
Senju Y, Tsai FC. Senju Y, et al. Biophys Rev. 2022 Jan 28;14(1):199-208. doi: 10.1007/s12551-021-00928-0. eCollection 2022 Feb. Biophys Rev. 2022. PMID: 35340609 Free PMC article. Review. - FERM domains recruit ample PI(4,5)P2s to form extensive protein-membrane attachments.
Ehret T, Heißenberg T, de Buhr S, Aponte-Santamaría C, Steinem C, Gräter F. Ehret T, et al. Biophys J. 2023 Apr 4;122(7):1325-1333. doi: 10.1016/j.bpj.2023.02.027. Epub 2023 Feb 22. Biophys J. 2023. PMID: 36814382 Free PMC article.
References
- Bretscher A., Edwards K., Fehon R.G. ERM proteins and merlin: integrators at the cell cortex. Nat. Rev. Mol. Cell Biol. 2002;3:586–599. - PubMed
- Mangeat P., Roy C., Martin M. ERM proteins in cell adhesion and membrane dynamics. Trends Cell Biol. 1999;9:187–192. - PubMed
- Tsukita S., Yonemura S. Cortical actin organization: lessons from ERM (ezrin/radixin/moesin) proteins. J. Biol. Chem. 1999;274:34507–34510. - PubMed
- Sauvanet C., Wayt J., Bretscher A. Structure, regulation, and functional diversity of microvilli on the apical domain of epithelial cells. Annu. Rev. Cell Dev. Biol. 2015;31:593–621. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials