Ruminococcin C, a promising antibiotic produced by a human gut symbiont - PubMed (original) (raw)
. 2019 Sep 25;5(9):eaaw9969.
doi: 10.1126/sciadv.aaw9969. eCollection 2019 Sep.
Clarisse Roblin 2 3, Sylvie Kieffer-Jaquinod 4, Sybille Tachon 2, Chloé Leprètre 1, Christian Basset 1, Dwi Aditiyarini 1, Hamza Olleik 2, Cendrine Nicoletti 2, Olivier Bornet 5, Olga Iranzo 2, Marc Maresca 2, Renaud Hardré 2, Michel Fons 6, Thierry Giardina 2, Estelle Devillard 3, Françoise Guerlesquin 5, Yohann Couté 4, Mohamed Atta 1, Josette Perrier 2, Mickael Lafond 2, Victor Duarte 1
Affiliations
- PMID: 31579822
- PMCID: PMC6760926
- DOI: 10.1126/sciadv.aaw9969
Ruminococcin C, a promising antibiotic produced by a human gut symbiont
Steve Chiumento et al. Sci Adv. 2019.
Abstract
A major public health challenge today is the resurgence of microbial infections caused by multidrug-resistant strains. Consequently, novel antimicrobial molecules are actively sought for development. In this context, the human gut microbiome is an under-explored potential trove of valuable natural molecules, such as the ribosomally-synthesized and post-translationally modified peptides (RiPPs). The biological activity of the sactipeptide subclass of RiPPs remains under-characterized. Here, we characterize an antimicrobial sactipeptide, Ruminococcin C1, purified from the caecal contents of rats mono-associated with Ruminococcus gnavus E1, a human symbiont. Its heterologous expression and post-translational maturation involving a specific sactisynthase establish a thioether network, which creates a double-hairpin folding. This original structure confers activity against pathogenic Clostridia and multidrug-resistant strains but no toxicity towards eukaryotic cells. Therefore, the Ruminococcin C1 should be considered as a valuable candidate for drug development and its producer strain R. gnavus E1 as a relevant probiotic for gut health enhancement.
Copyright © 2019 The Authors, some rights reserved; exclusive licensee American Association for the Advancement of Science. No claim to original U.S. Government Works. Distributed under a Creative Commons Attribution NonCommercial License 4.0 (CC BY-NC).
Figures
Fig. 1. Biosynthesis of sactipeptides.
(A) Thioether network in previously described sactipeptides. Thioether bridges, disulfide bridges, and head-to-tail cyclization are indicated by purple, yellow, and black lines, respectively. (B) Gene regulon encoding ruminococcins C, subtilosin A, thuricin CD, and thurincin H. Purple, radical SAM enzymes; light blue, precursor peptides; dark blue, transporter systems; green, signal peptidases; yellow, response regulators; pink, immunity systems; and gray, genes of unknown function. (C) Alignment of the five RumC peptide isoforms. (D) Thioether bond formation in sactipeptides catalyzed by radical SAM enzymes.
Fig. 2. Purification and characterization of the five RumC isoforms produced in vivo.
(A) Protocol for extraction from cecal contents to obtain a purified mixture of RumCs. Fractions were selected on the basis of their anti-Cp activity throughout the purification steps. PBS, phosphate-buffered saline. (B and C) LC-MS analyses of the two fractions containing the different RumC (* and † indicate succinimide and deamidated forms of C5 and C3, respectively). RumC5 was present in two consecutive C18 reversed-phase fractions. (D) Deconvoluted mass spectrum of RumC1 eluted at 21.6 min in nano–LC-MS analyses. (E) Deconvoluted mass spectrum of RumC1 after dithiothreitol (DTT)/iodoacetamide treatment. (F) Deconvoluted mass spectrum of synthetic unmodified RumC1.
Fig. 3. Tandem mass spectra of mature RumC1 peptide from in vivo and in vitro preparations and thioether network (see table S1 for theoretical and observed masses of interest).
(A) Deconvoluted MS/MS spectrum of in vivo–matured RumC1 (1 to 44, bold sequence) showing prominent y/b and c/z fragments induced by breaking of the amide bonds preceding the residues bound to cysteines in thioether bridges. The very structured peptide produced high-intensity and unusual internal fragments (blue italics), particularly ANSH (A12-H15) and RNANANVA (R34-A41), corresponding to fragments located between two linked residues. (B) Deconvoluted MS/MS spectrum of the heterologously matured mRumC1 [containing leader peptide (italics) and four additional GAMD amino acids for cloning purposes (gray italics)], revealing the same characteristic fragmentation pattern. Peaks below 500 Da (identical for the y series of in vivo RumC1) are not shown to improve overall visibility. All masses considered are monoisotopic masses. M (last peak in each spectrum) corresponds to the nonfragmented peptide. (C) Deconvoluted MS spectrum of mRumC1 after DTT/iodoalkylation showing no mass increment. Mass of 6694.09 (ammonia loss) corresponds to a succinimide (*) form produced as a by-product of high-temperature reduction of RumC1hm before iodoalkylation. (D) Identification of bridging partners. (E) Double-hairpin–like structure of mRumC1. Cysteine residues bridged via thioether bonds are shown in purple, and their amino acid partners are indicated in orange.
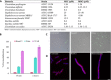
Fig. 4. RumC1 antimicrobial activity.
(A) Activity spectrum of RumC1 against selected Gram-positive strains. MIC and MBC were >100 μM for the following Gram-negative strains tested: Salmonella enterica (CIP 80.39), E. coli (ATCC 8739), E. coli MR4 (DSMZ 22314), Pseudomonas aeruginosa (ATCC 9027), P. aeruginosa fluoroquinolone resistant (CIP 107398), Acinetobacter baumanii (CIP 103572), A. baumanii multiresistant (CIP 110431), and Klebsiella pneumoniae MR4 (DSMZ 26371). (B and C) Membrane permeabilization assay on Cp cells treated with RumC1 or nisin based on measurement of PI incorporation (B) or SYTOX Green staining (C). (B) Cells incubated with cetyltrimethylammonium bromide (CTAB) were used as a positive lysis control, and untreated cells were used as a negative control. (C) Cells were treated for 15 min before staining. Scale bar, 10 μm. (D) Confocal imaging of control Cp cells or Cp cells treated with RumC1 or metronidazole. Membranes were stained with FM4-64FX, and DNA was stained with DAPI (4′,6-diamidino-2-phenylindole). RumC1 treatment leads to three morphotypes identical to the ones induced by metronidazole (fig. S9). This figure shows one of these three morphotypes, i.e., one regular cell associated with a cell three to four times longer and with uncondensed DNA throughout the cells with a few spots of highly condensed DNA. Scale bar, 2 μm.
Fig. 5. Proposed mechanism of maturation and activation of RumC1 in the human gut.
After induction of the two-component system, conventional transcription, and translation of the gene regulon Ruminococcins C, the intracellular RumC1 maturation process involves (i) an in situ posttranslational modification of the core peptide by RumMc1, leading to the inactive mRumC1; (ii) a partial cleavage of the leader peptide by RumPc, leading to the still inactive mRumC1c; (iii) an export in the intestinal lumen by RumTc; and (iv) an ex situ cleavage of the five remaining N-terminal amino acids of the leader peptide by pancreatic trypsin, leading to an active mRumC1cc (i.e., RumC1).
Similar articles
- The unusual structure of Ruminococcin C1 antimicrobial peptide confers clinical properties.
Roblin C, Chiumento S, Bornet O, Nouailler M, Müller CS, Jeannot K, Basset C, Kieffer-Jaquinod S, Couté Y, Torelli S, Le Pape L, Schünemann V, Olleik H, De La Villeon B, Sockeel P, Di Pasquale E, Nicoletti C, Vidal N, Poljak L, Iranzo O, Giardina T, Fons M, Devillard E, Polard P, Maresca M, Perrier J, Atta M, Guerlesquin F, Lafond M, Duarte V. Roblin C, et al. Proc Natl Acad Sci U S A. 2020 Aug 11;117(32):19168-19177. doi: 10.1073/pnas.2004045117. Epub 2020 Jul 27. Proc Natl Acad Sci U S A. 2020. PMID: 32719135 Free PMC article. - Ruminococcin C, an anti-clostridial sactipeptide produced by a prominent member of the human microbiota Ruminococcus gnavus.
Balty C, Guillot A, Fradale L, Brewee C, Boulay M, Kubiak X, Benjdia A, Berteau O. Balty C, et al. J Biol Chem. 2019 Oct 4;294(40):14512-14525. doi: 10.1074/jbc.RA119.009416. Epub 2019 Jul 23. J Biol Chem. 2019. PMID: 31337708 Free PMC article. - The Multifunctional Sactipeptide Ruminococcin C1 Displays Potent Antibacterial Activity In Vivo as Well as Other Beneficial Properties for Human Health.
Roblin C, Chiumento S, Jacqueline C, Pinloche E, Nicoletti C, Olleik H, Courvoisier-Dezord E, Amouric A, Basset C, Dru L, Ollivier M, Bogey-Lambert A, Vidal N, Atta M, Maresca M, Devillard E, Duarte V, Perrier J, Lafond M. Roblin C, et al. Int J Mol Sci. 2021 Mar 23;22(6):3253. doi: 10.3390/ijms22063253. Int J Mol Sci. 2021. PMID: 33806791 Free PMC article. - Ruminococcus gnavus: friend or foe for human health.
Crost EH, Coletto E, Bell A, Juge N. Crost EH, et al. FEMS Microbiol Rev. 2023 Mar 10;47(2):fuad014. doi: 10.1093/femsre/fuad014. FEMS Microbiol Rev. 2023. PMID: 37015876 Free PMC article. Review. - The Ruminococci: key symbionts of the gut ecosystem.
La Reau AJ, Suen G. La Reau AJ, et al. J Microbiol. 2018 Mar;56(3):199-208. doi: 10.1007/s12275-018-8024-4. Epub 2018 Feb 28. J Microbiol. 2018. PMID: 29492877 Review.
Cited by
- Bacteriocins: potentials and prospects in health and agrifood systems.
Reuben RC, Torres C. Reuben RC, et al. Arch Microbiol. 2024 Apr 25;206(5):233. doi: 10.1007/s00203-024-03948-y. Arch Microbiol. 2024. PMID: 38662051 Free PMC article. Review. - Genome Mining and Characterization of Biosynthetic Gene Clusters in Two Cave Strains of Paenibacillus sp.
Lebedeva J, Jukneviciute G, Čepaitė R, Vickackaite V, Pranckutė R, Kuisiene N. Lebedeva J, et al. Front Microbiol. 2021 Jan 11;11:612483. doi: 10.3389/fmicb.2020.612483. eCollection 2020. Front Microbiol. 2021. PMID: 33505378 Free PMC article. - Comparative Structure-Activity Analysis of the Antimicrobial Activity, Cytotoxicity, and Mechanism of Action of the Fungal Cyclohexadepsipeptides Enniatins and Beauvericin.
Olleik H, Nicoletti C, Lafond M, Courvoisier-Dezord E, Xue P, Hijazi A, Baydoun E, Perrier J, Maresca M. Olleik H, et al. Toxins (Basel). 2019 Sep 3;11(9):514. doi: 10.3390/toxins11090514. Toxins (Basel). 2019. PMID: 31484420 Free PMC article. - Recent Advances in Discovery, Bioengineering, and Bioactivity-Evaluation of Ribosomally Synthesized and Post-translationally Modified Peptides.
Zhong G, Wang ZJ, Yan F, Zhang Y, Huo L. Zhong G, et al. ACS Bio Med Chem Au. 2022 Dec 21;3(1):1-31. doi: 10.1021/acsbiomedchemau.2c00062. eCollection 2023 Feb 15. ACS Bio Med Chem Au. 2022. PMID: 37101606 Free PMC article. Review. - Mechanistic and functional aspects of the Ruminococcin C sactipeptide isoforms.
Shamseddine L, Roblin C, Veyrier I, Basset C, De Macedo L, Boyeldieu A, Maresca M, Nicoletti C, Brasseur G, Kieffer-Jaquinod S, Courvoisier-Dezord É, Amouric A, Carpentier P, Campo N, Bergé M, Polard P, Perrier J, Duarte V, Lafond M. Shamseddine L, et al. iScience. 2023 Aug 6;26(9):107563. doi: 10.1016/j.isci.2023.107563. eCollection 2023 Sep 15. iScience. 2023. PMID: 37664601 Free PMC article.
References
- O’Neill A. J., New antibacterial agents for treating infections caused by multi-drug resistant Gram-negative bacteria. Expert Opin. Investig. Drugs 17, 297–302 (2008). - PubMed
- J. O’Neill, Tackling Drug-Resistant Infections Globally: Final Report and Recommendations (Review on Antimicrobial Resistance, 2016).
- Arnison P. G., Bibb M. J., Bierbaum G., Bowers A. A., Bugni T. S., Bulaj G., Camarero J. A., Campopiano D. J., Challis G. L., Clardy J., Cotter P. D., Craik D. J., Dawson M., Dittmann E., Donadio S., Dorrestein P. C., Entian K.-D., Fischbach M. A., Garavelli J. S., Göransson U., Gruber C. W., Haft D. H., Hemscheidt T. K., Hertweck C., Hill C., Horswill A. R., Jaspars M., Kelly W. L., Klinman J. P., Kuipers O. P., Link A. J., Liu W., Marahiel M. A., Mitchell D. A., Moll G. N., Moore B. S., Müller R., Nair S. K., Nes I. F., Norris G. E., Olivera B. M., Onaka H., Patchett M. L., Piel J., Reaney M. J. T., Rebuffat S., Ross R. P., Sahl H.-G., Schmidt E. W., Selsted M. E., Severinov K., Shen B., Sivonen K., Smith L., Stein T., Süssmuth R. D., Tagg J. R., Tang G.-L., Truman A. W., Vederas J. C., Walsh C. T., Walton J. D., Wenzel S. C., Willey J. M., van der Donk W. A., Ribosomally synthesized and post-translationally modified peptide natural products: Overview and recommendations for a universal nomenclature. Nat. Prod. Rep. 30, 108–160 (2013). - PMC - PubMed
- M. C. Rea, R. P. Ross, P. D. Cotter, C. Hill, Classification of bacterocins from Gram-positive bacteria, in Prokaryotic Antimicrobial Peptides: From genes to Applications, D. Drider, S. Rebuffat, Eds. (Springer, 2011), pp. 29–53.
- Mathur H., Rea M. C., Cotter P. D., Hill C., Ross R. P., The sactibiotic subclass of bacteriocins: An update. Curr. Protein Pept. Sci. 16, 549–558 (2015). - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources