Characterization of the receptor-binding domain (RBD) of 2019 novel coronavirus: implication for development of RBD protein as a viral attachment inhibitor and vaccine - PubMed (original) (raw)
Characterization of the receptor-binding domain (RBD) of 2019 novel coronavirus: implication for development of RBD protein as a viral attachment inhibitor and vaccine
Wanbo Tai et al. Cell Mol Immunol. 2020 Jun.
Abstract
The outbreak of Coronavirus Disease 2019 (COVID-19) has posed a serious threat to global public health, calling for the development of safe and effective prophylactics and therapeutics against infection of its causative agent, severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), also known as 2019 novel coronavirus (2019-nCoV). The CoV spike (S) protein plays the most important roles in viral attachment, fusion and entry, and serves as a target for development of antibodies, entry inhibitors and vaccines. Here, we identified the receptor-binding domain (RBD) in SARS-CoV-2 S protein and found that the RBD protein bound strongly to human and bat angiotensin-converting enzyme 2 (ACE2) receptors. SARS-CoV-2 RBD exhibited significantly higher binding affinity to ACE2 receptor than SARS-CoV RBD and could block the binding and, hence, attachment of SARS-CoV-2 RBD and SARS-CoV RBD to ACE2-expressing cells, thus inhibiting their infection to host cells. SARS-CoV RBD-specific antibodies could cross-react with SARS-CoV-2 RBD protein, and SARS-CoV RBD-induced antisera could cross-neutralize SARS-CoV-2, suggesting the potential to develop SARS-CoV RBD-based vaccines for prevention of SARS-CoV-2 and SARS-CoV infection.
Keywords: 2019 novel coronavirus; SARS-CoV-2; cross-neutralization; receptor-binding domain; spike protein; viral inhibitor.
Conflict of interest statement
The authors declare no competing interests.
Figures
Fig. 1
Characterization of SARS-CoV-2 RBD. a Multiple sequence alignment of RBDs of SARS-CoV-2, SARS-CoV, and MERS-CoV spike (S) proteins. GenBank accession numbers are QHR63250.1 (SARS-CoV-2 S), AY278488.2 (SARS-CoV S), and AFS88936.1 (MERS-CoV S). Variable amino acid residues between SARS-CoV-2 and SARS-CoV are highlighted in cyan, and conserved residues among SARS-CoV-2, SARS-CoV, and MERS-CoV are highlighted in yellow. Asterisks represent fully conserved residues, colons represent highly conserved residues, and periods represent lowly conserved residues. The alignment was performed using Clustal Omega. SDS-PAGE (b) and Western blot (c, d) analysis of SARS-CoV-2 RBD. The protein molecular weight marker (kDa) is indicated on the left. SARS-CoV and MERS-CoV RBDs were included as controls. Antisera (1:3,000 dilution) from mice immunized with SARS-CoV RBD (c) and MERS-CoV RBD (d) were used for Western blot analysis
Fig. 2
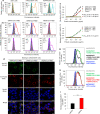
Detection of SARS-CoV-2 RBD binding to human ACE2 receptor. a Flow cytometry analysis of receptor expression in stable cell lines. (left panel) 293T cells alone expressed neither human ACE2 (hACE2) receptor (orange line) nor hDPP4 receptor (cyan line); (middle panel) hACE2-expressing 293T (hACE2/293T) cells expressed only hACE2 (orange line), but not hDPP4 (cyan line); (right panel) hDPP4-expressing 293T (hDPP4/293T) cells expressed only hDPP4 (cyan line), but not hACE2 (orange line). Mock-incubated cells (gray shading) were used as control. Representative images and median fluorescence intensity (MFI) ± standard error (s.e.m.) were shown (n = 4). b, c Flow cytometry analysis of SARS-CoV-2 RBD binding to cell-associated hACE2 receptor in hACE2/293T stable cell lines. SARS-CoV-2 RBD protein bound strongly to hACE2/293T cells (b (left panel, red line)), but not to hDPP4/293T cells (c (left panel, violet line)). SARS-CoV RBD protein bound to hACE2/293T cells (b (middle panel, red line)), but not to hDPP4/293T cells (c (middle panel, violet line)). MERS-CoV RBD protein did not bind to hACE2/293T cells (b (right panel, red line)), but rather bound to hDPP4/293T cells (c (right panel, violet line)). Human IgG Fc (hIgG-Fc, hereinafter hFc) protein-incubated cells (blue line) and mock-incubated cells (gray shading) were included as controls (b, c). Representative images and MFI ± s.e.m. were shown (n = 4). d Immunofluorescence detection of SARS-CoV-2 RBD binding to cell-associated hACE2 receptor in hACE2/293T cells. SARS-CoV-2 RBD protein (green) and SARS-CoV RBD protein (green), each of which was fused with a C-terminal hFc, were stained with FITC-labeled goat anti-human IgG antibody (1:500). hACE2 was stained with a goat-anti-hACE2 antibody (5 μg/ml) and Alexa-Fluor 647-labeled anti-goat antibody (red) (1:200). Fc-fused MERS-CoV RBD protein did not bind to hACE2, so only hACE2 (red), but not RBD (green), was detected in hACE2/293T cells. Nuclei were stained with 4’,6-diamidino-2-phenylindole (DAPI, blue). Scale bar: 10 μm. Representative images are shown. e Detection of dose-dependent binding of SARS-CoV-2 RBD protein to soluble hACE2 (sACE2) receptor by ELISA. The SARS-CoV-2 RBD binding to soluble hDPP4 (sDPP4) receptor (f), and the binding of both SARS-CoV RBD and MERS-CoV RBD proteins to sACE2 (e), or sDPP4 (f), were tested. Control: hFc protein. Data are presented as mean A450 ± s.e.m. (n = 4). 50% effective dose (EC50) was calculated for the binding between SARS-CoV-2 RBD (black) or SARS-CoV RBD (red) and hACE2 protein (e, sACE2), or the binding between MERS-CoV RBD and hDPP4 protein (sDPP4, green) (f). g–i Flow cytometry analysis of inhibition of SARS-CoV-2 RBD protein binding to hACE2/293T cells by sACE2. Binding of SARS-CoV-2 RBD to hACE2/293T cells (g, h, green line) was blocked by sACE2 (g, black line), but not by sDPP4 (h, red line). hFc protein-incubated cells (blue line) and mock-incubated cells (gray shading) were included as controls (g, h). Representative images are shown. i The blocking ability of sACE2 or sDPP4, as described above, was expressed as MFI ± s.e.m. (n = 4). Low MFI correlates with high blockage. Experiments were repeated twice and yielded similar results
Fig. 3
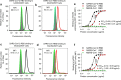
Comparison of SARS-CoV-2 RBD protein binding to human and bat ACE2 receptors. Flow cytometry analysis of SARS-CoV-2 RBD binding to hACE2 and bat ACE2 (bACE2) receptors in 293T cells transiently expressing hACE2 or bACE2. 293T cells were transiently transfected with hACE2 or bACE2 plasmid and incubated with SARS-CoV-2 RBD protein at various concentrations for analysis. SARS-CoV RBD and MERS-CoV RBD proteins were used as controls. Representative images of SARS-CoV-2 RBD protein (2.5 μg/ml) binding to bACE2/293T (a, black line), or hACE2/293T (c, black line), cells were shown. Binding of SARS-CoV RBD protein (2.5 μg/ml) to bACE2/293T (b, red line), or hACE2/293T (d, red line), cells were used as a comparison. MERS-CoV RBD protein (green line) and mock-incubated (gray shading) cells (a–d) were included as controls. e, f Dose-dependent binding of SARS-CoV-2 RBD protein to bACE2/293T (e), or hACE2/293T (f), cells by flow cytometry analysis. Significant differences between binding of SARS-CoV-2 RBD (black) and SARS-CoV RBD (red) to cell-associated bACE2 receptor (e), or hACE2 receptor (f) were identified based on the EC50 values. The data are presented as mean ± s.e.m. (n = 4). Experiments were repeated twice and yielded similar results
Fig. 4
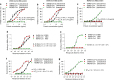
Ability of SARS-CoV-2 RBD to inhibit viral entry, as well as its cross-reactivity and cross-neutralizing activity with SARS-CoV. a Dose-dependent inhibition of SARS-CoV-2 RBD protein against pseudotyped SARS-CoV-2 entry into hACE2/293T cells. SARS-CoV and MERS-CoV RBDs, as well as hDPP4/293T cells, were included as controls. SARS-CoV-2 RBD protein inhibited entry of SARS-CoV-2 and SARS-CoV pseudoviruses into their respective target (hACE2/293T) cells (a), but not the entry of MERS-CoV pseudovirus into its target (hDPP4/293T) cells (a). SARS-CoV RBD protein inhibited both SARS-CoV-2 and SARS-CoV pseudovirus entry, but not MERS-CoV pseudovirus entry (b). MERS-CoV RBD inhibited neither SARS-CoV-2 nor SARS-CoV pseudovirus entry, but it did inhibit MERS-CoV pseudovirus entry (c). The data are presented as mean inhibition (%) ± s.e.m. (n = 4), and 50% inhibition concentration (IC50) was calculated for SARS-CoV-2 RBD (a, b, black), or SARS-CoV RBD (a, b, red), protein against SARS-CoV-2 pseudovirus and SARS-CoV pseudovirus and for MERS-CoV RBD protein (green) against MERS-CoV pseudovirus (c). d Cross-reactivity of SARS-CoV-2 RBD protein with SARS-CoV RBD-specific mouse sera by ELISA. Sera of mice immunized with mammalian cell-expressed SARS-CoV RBD protein were tested. Sera of mice immunized with mammalian cell-expressed MERS-CoV RBD protein were used as control. The data are presented as mean A450 ± s.e.m. (n = 4). The IgG antibody (Ab) titers were calculated as the endpoint dilution that remains positively detectable for SARS-CoV-2 RBD (black), or SARS-CoV RBD (red), binding to anti-SARS-CoV RBD sera (d) and for MERS-CoV RBD (green) binding to anti-MERS-CoV RBD sera (e). f Cross-neutralization of SARS-CoV RBD-immunized mouse sera against SARS-CoV-2 infection by pseudovirus neutralization assay. MERS-CoV RBD-immunized mouse sera were used as control. The data are presented as mean neutralization (%) ± s.e.m. (n = 4). 50% neutralizing antibody titers (NT50) were calculated against SARS-CoV-2 pseudovirus (black), or SARS-CoV pseudovirus (red), (f) infection in hACE2/293T target cells, as well as against MERS-CoV pseudovirus (green) (g) infection in hDPP4/293T cells. Experiments were repeated twice and yielded similar results
Similar articles
- Comparison of Severe Acute Respiratory Syndrome Coronavirus 2 Spike Protein Binding to ACE2 Receptors from Human, Pets, Farm Animals, and Putative Intermediate Hosts.
Zhai X, Sun J, Yan Z, Zhang J, Zhao J, Zhao Z, Gao Q, He WT, Veit M, Su S. Zhai X, et al. J Virol. 2020 Jul 16;94(15):e00831-20. doi: 10.1128/JVI.00831-20. Print 2020 Jul 16. J Virol. 2020. PMID: 32404529 Free PMC article. - Identification of SARS-CoV RBD-targeting monoclonal antibodies with cross-reactive or neutralizing activity against SARS-CoV-2.
Tai W, Zhang X, He Y, Jiang S, Du L. Tai W, et al. Antiviral Res. 2020 Jul;179:104820. doi: 10.1016/j.antiviral.2020.104820. Epub 2020 May 13. Antiviral Res. 2020. PMID: 32405117 Free PMC article. - Key residues of the receptor binding motif in the spike protein of SARS-CoV-2 that interact with ACE2 and neutralizing antibodies.
Yi C, Sun X, Ye J, Ding L, Liu M, Yang Z, Lu X, Zhang Y, Ma L, Gu W, Qu A, Xu J, Shi Z, Ling Z, Sun B. Yi C, et al. Cell Mol Immunol. 2020 Jun;17(6):621-630. doi: 10.1038/s41423-020-0458-z. Epub 2020 May 15. Cell Mol Immunol. 2020. PMID: 32415260 Free PMC article. - Targeting SARS-CoV2 Spike Protein Receptor Binding Domain by Therapeutic Antibodies.
Hussain A, Hasan A, Nejadi Babadaei MM, Bloukh SH, Chowdhury MEH, Sharifi M, Haghighat S, Falahati M. Hussain A, et al. Biomed Pharmacother. 2020 Oct;130:110559. doi: 10.1016/j.biopha.2020.110559. Epub 2020 Aug 1. Biomed Pharmacother. 2020. PMID: 32768882 Free PMC article. Review. - The SARS-CoV-2 Spike Glycoprotein as a Drug and Vaccine Target: Structural Insights into Its Complexes with ACE2 and Antibodies.
Papageorgiou AC, Mohsin I. Papageorgiou AC, et al. Cells. 2020 Oct 22;9(11):2343. doi: 10.3390/cells9112343. Cells. 2020. PMID: 33105869 Free PMC article. Review.
Cited by
- Novel human neutralizing mAbs specific for Spike-RBD of SARS-CoV-2.
Passariello M, Gentile C, Ferrucci V, Sasso E, Vetrei C, Fusco G, Viscardi M, Brandi S, Cerino P, Zambrano N, Zollo M, De Lorenzo C. Passariello M, et al. Sci Rep. 2021 May 26;11(1):11046. doi: 10.1038/s41598-021-90348-7. Sci Rep. 2021. PMID: 34040046 Free PMC article. - Screening of inhibitors against SARS-CoV-2 spike protein and their capability to block the viral entry mechanism: A viroinformatics study.
Farouk AE, Baig MH, Khan MI, Park T, Alotaibi SS, Dong JJ. Farouk AE, et al. Saudi J Biol Sci. 2021 Jun;28(6):3262-3269. doi: 10.1016/j.sjbs.2021.02.066. Epub 2021 Feb 26. Saudi J Biol Sci. 2021. PMID: 33654454 Free PMC article. - Evidence for SARS-CoV-2 Infection of Animal Hosts.
Abdel-Moneim AS, Abdelwhab EM. Abdel-Moneim AS, et al. Pathogens. 2020 Jun 30;9(7):529. doi: 10.3390/pathogens9070529. Pathogens. 2020. PMID: 32629960 Free PMC article. Review. - Potential inhibitors of the interaction between ACE2 and SARS-CoV-2 (RBD), to develop a drug.
Benítez-Cardoza CG, Vique-Sánchez JL. Benítez-Cardoza CG, et al. Life Sci. 2020 Sep 1;256:117970. doi: 10.1016/j.lfs.2020.117970. Epub 2020 Jun 15. Life Sci. 2020. PMID: 32553928 Free PMC article. - Genetic modification to design a stable yeast-expressed recombinant SARS-CoV-2 receptor binding domain as a COVID-19 vaccine candidate.
Chen WH, Wei J, Kundu RT, Adhikari R, Liu Z, Lee J, Versteeg L, Poveda C, Keegan B, Villar MJ, de Araujo Leao AC, Rivera JA, Gillespie PM, Pollet J, Strych U, Zhan B, Hotez PJ, Bottazzi ME. Chen WH, et al. Biochim Biophys Acta Gen Subj. 2021 Jun;1865(6):129893. doi: 10.1016/j.bbagen.2021.129893. Epub 2021 Mar 14. Biochim Biophys Acta Gen Subj. 2021. PMID: 33731300 Free PMC article.
References
- World Health Organization. Naming the coronavirus disease (COVID-19) and the virus that causes it. https://www.who.int/emergencies/diseases/novel-coronavirus-2019/technica... (2020).
Publication types
MeSH terms
Substances
Grants and funding
- R01 AI137472/AI/NIAID NIH HHS/United States
- R01 AI139092/AI/NIAID NIH HHS/United States
- R01AI139092/U.S. Department of Health & Human Services | National Institutes of Health (NIH)/International
- R01AI137472/U.S. Department of Health & Human Services | National Institutes of Health (NIH)/International
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous