The dawn of mRNA vaccines: The COVID-19 case - PubMed (original) (raw)
Review
The dawn of mRNA vaccines: The COVID-19 case
Rein Verbeke et al. J Control Release. 2021.
Abstract
In less than one year since the outbreak of the COVID-19 pandemic, two mRNA-based vaccines, BNT162b2 and mRNA-1273, were granted the first historic authorization for emergency use, while another mRNA vaccine, CVnCoV, progressed to phase 3 clinical testing. The COVID-19 mRNA vaccines represent a new class of vaccine products, which consist of synthetic mRNA strands encoding the SARS-CoV-2 Spike glycoprotein, packaged in lipid nanoparticles to deliver mRNA to cells. This review digs deeper into the scientific breakthroughs of the last decades that laid the foundations for the rapid rise of mRNA vaccines during the COVID-19 pandemic. As well as providing momentum for mRNA vaccines, SARS-CoV-2 represents an ideal case study allowing to compare design-activity differences between the different mRNA vaccine candidates. Therefore, a detailed overview of the composition and (pre)clinical performance of the three most advanced mRNA vaccines is provided and the influence of choices in their structural design on to their immunogenicity and reactogenicity profile is discussed in depth. In addition to the new fundamental insights in the mRNA vaccines' mode of action highlighted here, we also point out which unknowns remain that require further investigation and possibly, optimization in future mRNA vaccine development.
Copyright © 2021 Elsevier B.V. All rights reserved.
Figures
Graphical abstract
Fig. 1
Timeline of the development of three most advanced mRNA vaccines against COVID-19; BNT162b2 (Green dots), mRNA-1273 (Blue) and CVnCoV (Orange). (For interpretation of the references to colour in this figure legend, the reader is referred to the web version of this article.)
Fig. 2
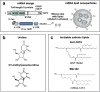
COVID-19 mRNA vaccine design. (a) The COVID-19 mRNA vaccines contain a mRNA sequence encoding the full length S protein with two proline substitutions (K986P and V987P). The S protein's genetic code is flanked by structural elements to produce a mature mRNA. Each of these elements can be optimized in order to modulate mRNA stability, translation capacity and innate immune activity. (b) While the CVnCoV vaccine candidate make use of unmodified uridines, BNT162b2 and mRNA-1273 are nucleoside-modified with a substitution of N1-methylpseudouridine (1mψ) for uridine (U). (c) Chemical structures of the ionizable cationic lipids ALC-0315 (((4-Hydroxybutyl)azanediyl)bis(hexane-6,1-diyl)bis(2-hexyldecanoate)) and SM-102 (Heptadecan-9-yl 8-((2-hydroxyethyl)(8-(nonyloxy)-8-oxooctyl)amino)octanoate) used in the LNP formulation of BNT162b2 and mRNA-1273, respectively. The ionizable cationic lipid used in CVnCoV has not (yet) been disclosed. Abbreviations: SS; Signal sequence, NTD; N-terminal domain, S1/S2; native furine cleavage site, TM; transmembrane domain.
Fig. 3
The mode of action of mRNA vaccines. (a –at the injection site) Upon endocytosis by muscle-resident cells, mRNA LNPs trigger a transient inflammatory response recruiting neutrophils, monocytes and DCs to the injection site. Local and recruited APC subsets transiently express the S protein mRNA and undergo maturation in response to innate immune sensing of the mRNA. The migration of targeted/activated APCs and direct lymphatic transport of mRNA LNPs and cell debris containing S proteins, brings the S antigen to B cells and T cells in draining lymph nodes. (b – at the cellular level) To avoid lysosomal degradation, mRNA must escape the endosomes and binds to ribosomes, known as a complex and rate-limiting process, which is facilitated by the ionizable LNP carrier. After translation and transport of S proteins through the endoplasmatic reticulum and Golgi apparatus, S proteins are exposed as prefusion-stabilized trimer constructs at the cell surface. This membrane-bound S antigen can efficiently be recognized and internalized by B cells, which leads to a series of events activating B cells responses towards neutralizing antibody generation against the S protein. Moreover, the expressed S antigens can gain access to the MHC class I antigen presentation pathway to prime CD8+ T cells that can eliminate infected cells, while recycling mechanisms allow the presentation of antigenic epitopes in MHC-II complexes to CD4+ helper T cells, especially needed to promote the antibody production by providing B cell help. Abbreviations: APC; Antigen presenting cell, RBD; Receptor binding domain, MHC; major histocompatibility complex.
Fig. 4
Proposed innate immune signaling in response to mRNA vaccination. The internalization of mRNA LNPs can be detected by innate immune sensors that are localized in the endosomes and cytosol. The detection of mRNA, by the endosomal TLR/8, recruits the MYD88 signal transduction adaptor and leads to the expression of type I IFNs (IFN-α and IFN-β) through IFN regulatory factor 7, and to the secretion of other proinflammatory cytokines through nuclear factor κB (NF-κB). In addition, dsRNA contaminants and/or secondary structures in the mRNA product can interact with TLR3 in the endosomes, recruiting TRIF, as well as upon their arrival in the cytosol be detected by RIG-I and MDA5, binding MAVS. The activation of TRIF and MAVS is followed by molecular cascades that results in the expression of type I IFNs in control of IRF3 and IRF7. In turn, the type I IFN cytokines bind autocrine or paracrine receptors, which eventually regulates the gene expression of hundreds of proteins involved in antiviral immunity. This includes the expression of MHC-I and co-stimulatory molecules, needed for T cell responses, as well as antiviral proteins involved with undesirable anti-RNA responses. Methods such as the introduction of modified nucleotides, the removal of dsRNA fragments, and sequence-engineering, can be utilized to minimize or control the type I IFN activity of mRNA. However, it remains unclear how to strike the perfect balance between obtaining sufficient mRNA-encoded antigen expression and adequate immunostimulation in order to support adaptive immunity. In addition, more research is needed to investigate whether and how the recognition of lipid components in the LNP vehicle might contribute to the innate immune response to mRNA vaccines. Abbreviations: IRF; interferon regulatory factor, ISG; interferon-stimulated gene, NF-κB; nuclear factor-κB, MAVS; mitochondrial antiviral signaling protein, MDA5; melanoma differentiation-associated protein 5, MYD88; myeloid differentiation primary response protein 88, TRIF, Toll-IL-1 receptor domain-containing adapter protein inducing IFNβ.
Similar articles
- Review of COVID-19 mRNA Vaccines: BNT162b2 and mRNA-1273.
Teo SP. Teo SP. J Pharm Pract. 2022 Dec;35(6):947-951. doi: 10.1177/08971900211009650. Epub 2021 Apr 12. J Pharm Pract. 2022. PMID: 33840294 Review. - Immunogenicity and reactogenicity of SARS-CoV-2 vaccines in people living with HIV in the Netherlands: A nationwide prospective cohort study.
Hensley KS, Jongkees MJ, Geers D, GeurtsvanKessel CH, Mueller YM, Dalm VASH, Papageorgiou G, Steggink H, Gorska A, Bogers S, den Hollander JG, Bierman WFW, Gelinck LBS, Schippers EF, Ammerlaan HSM, van der Valk M, van Vonderen MGA, Delsing CE, Gisolf EH, Bruns AHW, Lauw FN, Berrevoets MAH, Sigaloff KCE, Soetekouw R, Branger J, de Mast Q, Lammers AJJ, Lowe SH, de Vries RD, Katsikis PD, Rijnders BJA, Brinkman K, Roukens AHE, Rokx C. Hensley KS, et al. PLoS Med. 2022 Oct 27;19(10):e1003979. doi: 10.1371/journal.pmed.1003979. eCollection 2022 Oct. PLoS Med. 2022. PMID: 36301821 Free PMC article. - Real-world serological responses to extended-interval and heterologous COVID-19 mRNA vaccination in frail, older people (UNCoVER): an interim report from a prospective observational cohort study.
Vinh DC, Gouin JP, Cruz-Santiago D, Canac-Marquis M, Bernier S, Bobeuf F, Sengupta A, Brassard JP, Guerra A, Dziarmaga R, Perez A, Sun Y, Li Y, Roussel L, Langelier MJ, Ke D, Arnold C, Whelan M, Pelchat M, Langlois MA, Zhang X, Mazer BD; COVID-19 Immunity Task Force and UNCoVER Investigators. Vinh DC, et al. Lancet Healthy Longev. 2022 Mar;3(3):e166-e175. doi: 10.1016/S2666-7568(22)00012-5. Epub 2022 Feb 23. Lancet Healthy Longev. 2022. PMID: 35224524 Free PMC article. - Immunogenicity and safety of a booster dose of a self-amplifying RNA COVID-19 vaccine (ARCT-154) versus BNT162b2 mRNA COVID-19 vaccine: a double-blind, multicentre, randomised, controlled, phase 3, non-inferiority trial.
Oda Y, Kumagai Y, Kanai M, Iwama Y, Okura I, Minamida T, Yagi Y, Kurosawa T, Greener B, Zhang Y, Walson JL. Oda Y, et al. Lancet Infect Dis. 2024 Apr;24(4):351-360. doi: 10.1016/S1473-3099(23)00650-3. Epub 2023 Dec 20. Lancet Infect Dis. 2024. PMID: 38141632 Clinical Trial. - Lead SARS-CoV-2 Candidate Vaccines: Expectations from Phase III Trials and Recommendations Post-Vaccine Approval.
Tumban E. Tumban E. Viruses. 2020 Dec 31;13(1):54. doi: 10.3390/v13010054. Viruses. 2020. PMID: 33396343 Free PMC article. Review.
Cited by
- A Triazolium-Anchored Self-Immolative Linker Enables Self-Assembly-Driven siRNA Binding and Esterase-Induced Release.
Hollstein S, Ali LMA, Coste M, Vogel J, Bettache N, Ulrich S, von Delius M. Hollstein S, et al. Chemistry. 2023 Feb 7;29(8):e202203311. doi: 10.1002/chem.202203311. Epub 2022 Dec 16. Chemistry. 2023. PMID: 36346344 Free PMC article. - Humoral immune response to SARS-CoV-2 mRNA vaccines is associated with choice of vaccine and systemic adverse reactions.
Klingel H, Krüttgen A, Imöhl M, Kleines M. Klingel H, et al. Clin Exp Vaccine Res. 2023 Jan;12(1):60-69. doi: 10.7774/cevr.2023.12.1.60. Epub 2023 Jan 31. Clin Exp Vaccine Res. 2023. PMID: 36844685 Free PMC article. - Effects of Vaccination against COVID-19 in Chronic Spontaneous and Inducible Urticaria (CSU/CIU) Patients: A Monocentric Study.
Grieco T, Ambrosio L, Trovato F, Vitiello M, Demofonte I, Fanto M, Paolino G, Pellacani G. Grieco T, et al. J Clin Med. 2022 Mar 25;11(7):1822. doi: 10.3390/jcm11071822. J Clin Med. 2022. PMID: 35407429 Free PMC article. - Comparison of Physicochemical Properties of LipoParticles as mRNA Carrier Prepared by Automated Microfluidic System and Bulk Method.
Ayad C, Yavuz A, Salvi JP, Libeau P, Exposito JY, Ginet V, Monge C, Verrier B, Arruda DC. Ayad C, et al. Pharmaceutics. 2022 Jun 18;14(6):1297. doi: 10.3390/pharmaceutics14061297. Pharmaceutics. 2022. PMID: 35745869 Free PMC article. - Virus-Mimicking Cell Membrane-Coated Nanoparticles for Cytosolic Delivery of mRNA.
Park JH, Mohapatra A, Zhou J, Holay M, Krishnan N, Gao W, Fang RH, Zhang L. Park JH, et al. Angew Chem Int Ed Engl. 2022 Jan 10;61(2):e202113671. doi: 10.1002/anie.202113671. Epub 2021 Nov 29. Angew Chem Int Ed Engl. 2022. PMID: 34694684 Free PMC article.
References
- Verbeke R., Lentacker I., De Smedt S.C., Dewitte H. Three decades of messenger RNA vaccine development. Nano Today. 2019;28:100766.
- O’Neill L.A.J., Golenbock D., Bowie A.G. The history of toll-like receptors - redefining innate immunity. Nat. Rev. Immunol. 2013;13:453–460. - PubMed
- Kariko K., Ni H.P., Capodici J., Lamphier M., Weissman D. mRNA is an endogenous ligand for toll-like receptor 3. J. Biol. Chem. 2004;279:12542–12550. - PubMed
- Kariko K., Buckstein M., Ni H.P., Weissman D. Suppression of RNA recognition by toll-like receptors: the impact of nucleoside modification and the evolutionary origin of RNA. Immunity. 2005;23:165–175. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous