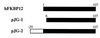A three-hybrid system for detecting small ligand-protein receptor interactions - PubMed (original) (raw)
A three-hybrid system for detecting small ligand-protein receptor interactions
E J Licitra et al. Proc Natl Acad Sci U S A. 1996.
Abstract
Small ligand-receptor interactions underlie many fundamental processes in biology and form the basis for pharmacological intervention of human diseases in medicine. We report herein a genetic system, named the yeast three-hybrid system, for detecting ligand-receptor interactions in vivo. This system is adapted from the yeast two-hybrid system with which a third synthetic hybrid ligand is combined. The feasibility of this system was demonstrated using as the hybrid ligand a heterodimer of covalently linked dexamethasone and FK506. Yeast expressing fusion proteins of the hormone binding domain of the rat glucocorticoid receptor fused to the LexA DNA-binding domain and FKBP12 fused to a transcriptional activation domain activated reporter genes when plated on medium containing the dexamethasone-FK506 heterodimer. The reporter gene activation is completely abrogated in a competitive manner by the presence of excess FK506. Using this system, we screened a Jurkat cDNA library fused to the transcriptional activation domain in yeast expressing the hormone binding domain of rat glucocorticoid receptor-LexA DNA binding domain fusion protein in the presence of dexamethasone-FK506 heterodimer. We isolated overlapping clones of human FKBP12. These results demonstrate that the three-hybrid system can be used to discover receptors for small ligands and to screen for new ligands to known receptors.
Figures
Figure 1
Synthesis and structure of the dexamethasone–FK506 hybrid ligand (4).
Figure 2
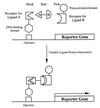
Diagrammatic representation of the components of the yeast three-hybrid system. The synthetic “bait” hybrid ligand consists of ligand A (triangle) and ligand B (semicircle) connected by a linker. The “hook” fusion protein consists of the receptor for ligand A fused to the DNA-binding domain of a transcription factor. The “fish” fusion protein consists of the receptor for ligand B fused to a transactivation domain of a transcription factor. The relative sizes of the three hybrid molecules are not drawn to scale. The “hook,” the “bait,” and the “fish” form a trimeric complex, and the resulting proximity of the transactivation domain and the DNA-binding domain leads to activation of the reporter gene.
Figure 3
(A) Control fusion protein (pSH17-4) between LexA DNA-binding domain (solid bars) and Gal4 transactivation domain (shaded bars) and variants of “hook” fusion proteins between LexA DNA-binding domain and hormone-binding domain of rGR (open bars). The first and the last amino acid of the LexA DNA binding protein, Gal4 transactivation domain, and the hormone binding domain of rGR, as well as the relative position of the two mutations in the hormone-binding domain of rGR are indicated above each construct. (B) Activation of the lacZ reporter gene by dexamethasone–FK506 hybrid ligand. EGY48 transformed with different “hook” fusion protein constructs (numbered as in_A_), the pJG-FKBP12 “fish” fusion protein construct and the reporter pSH18-34 from a master plate were replica plated onto SC medium (His−, Ura−, Trp−) containing galactose, 5-bromo-4-chloro-3-indolyl β-
d
-galactoside, and 1 μM dexamethasone–FK506 and incubated at 30°C for 3 days. (C) Inhibition of the_lacZ_ reporter gene activation by FK506 competition. All conditions and strains of yeast are identical to those in_B_ except that 10 μM of FK506 was present in the medium.
Figure 3
(A) Control fusion protein (pSH17-4) between LexA DNA-binding domain (solid bars) and Gal4 transactivation domain (shaded bars) and variants of “hook” fusion proteins between LexA DNA-binding domain and hormone-binding domain of rGR (open bars). The first and the last amino acid of the LexA DNA binding protein, Gal4 transactivation domain, and the hormone binding domain of rGR, as well as the relative position of the two mutations in the hormone-binding domain of rGR are indicated above each construct. (B) Activation of the lacZ reporter gene by dexamethasone–FK506 hybrid ligand. EGY48 transformed with different “hook” fusion protein constructs (numbered as in_A_), the pJG-FKBP12 “fish” fusion protein construct and the reporter pSH18-34 from a master plate were replica plated onto SC medium (His−, Ura−, Trp−) containing galactose, 5-bromo-4-chloro-3-indolyl β-
d
-galactoside, and 1 μM dexamethasone–FK506 and incubated at 30°C for 3 days. (C) Inhibition of the_lacZ_ reporter gene activation by FK506 competition. All conditions and strains of yeast are identical to those in_B_ except that 10 μM of FK506 was present in the medium.
Figure 4
FKBP12 cDNA variants isolated through the yeast three-hybrid screen of a Jurkat cDNA library. pJG-1 encodes FKBP12 minus the first 5 amino acids; pJG-2, isolated in duplicate, encodes FKBP12 with an extended 29-amino acid fragment at its amino terminus.
Similar articles
- A novel FK506 binding protein can mediate the immunosuppressive effects of FK506 and is associated with the cardiac ryanodine receptor.
Lam E, Martin MM, Timerman AP, Sabers C, Fleischer S, Lukas T, Abraham RT, O'Keefe SJ, O'Neill EA, Wiederrecht GJ. Lam E, et al. J Biol Chem. 1995 Nov 3;270(44):26511-22. doi: 10.1074/jbc.270.44.26511. J Biol Chem. 1995. PMID: 7592869 - Identification of an 11-kDa FKBP12-rapamycin-binding domain within the 289-kDa FKBP12-rapamycin-associated protein and characterization of a critical serine residue.
Chen J, Zheng XF, Brown EJ, Schreiber SL. Chen J, et al. Proc Natl Acad Sci U S A. 1995 May 23;92(11):4947-51. doi: 10.1073/pnas.92.11.4947. Proc Natl Acad Sci U S A. 1995. PMID: 7539137 Free PMC article. - A GAL4-based yeast three-hybrid system for the identification of small molecule-target protein interactions.
Henthorn DC, Jaxa-Chamiec AA, Meldrum E. Henthorn DC, et al. Biochem Pharmacol. 2002 May 1;63(9):1619-28. doi: 10.1016/s0006-2952(02)00884-5. Biochem Pharmacol. 2002. PMID: 12007565 - FAP48, a new protein that forms specific complexes with both immunophilins FKBP59 and FKBP12. Prevention by the immunosuppressant drugs FK506 and rapamycin.
Chambraud B, Radanyi C, Camonis JH, Shazand K, Rajkowski K, Baulieu EE. Chambraud B, et al. J Biol Chem. 1996 Dec 20;271(51):32923-9. doi: 10.1074/jbc.271.51.32923. J Biol Chem. 1996. PMID: 8955134 - FK506 and the role of immunophilins in nerve regeneration.
Gold BG. Gold BG. Mol Neurobiol. 1997 Dec;15(3):285-306. doi: 10.1007/BF02740664. Mol Neurobiol. 1997. PMID: 9457703 Review.
Cited by
- RNA-protein interactions in the yeast three-hybrid system: affinity, sensitivity, and enhanced library screening.
Hook B, Bernstein D, Zhang B, Wickens M. Hook B, et al. RNA. 2005 Feb;11(2):227-33. doi: 10.1261/rna.7202705. Epub 2004 Dec 21. RNA. 2005. PMID: 15613539 Free PMC article. - Design and synthesis of a cell-permeable synthetic transcription factor mimic.
Xiao X, Yu P, Lim HS, Sikder D, Kodadek T. Xiao X, et al. J Comb Chem. 2007 Jul-Aug;9(4):592-600. doi: 10.1021/cc070023a. Epub 2007 May 27. J Comb Chem. 2007. PMID: 17530904 Free PMC article. - Interaction trap/two-hybrid system to identify interacting proteins.
Golemis EA, Serebriiskii I, Finley RL Jr, Kolonin MG, Gyuris J, Brent R. Golemis EA, et al. Curr Protoc Cell Biol. 2001 May;Chapter 17:Unit 17.3. doi: 10.1002/0471143030.cb1703s08. Curr Protoc Cell Biol. 2001. PMID: 18228339 Free PMC article. - Transcriptional regulation improves the throughput of three-hybrid counter selections in Saccharomyces cerevisiae.
Harton MD, Wingler LM, Cornish VW. Harton MD, et al. Biotechnol J. 2013 Dec;8(12):1485-91. doi: 10.1002/biot.201300186. Epub 2013 Dec 4. Biotechnol J. 2013. PMID: 24318638 Free PMC article. - Analysis of a peptide hormone-receptor interaction in the yeast two-hybrid system.
Zhu J, Kahn CR. Zhu J, et al. Proc Natl Acad Sci U S A. 1997 Nov 25;94(24):13063-8. doi: 10.1073/pnas.94.24.13063. Proc Natl Acad Sci U S A. 1997. PMID: 9371800 Free PMC article.
References
- Yamamoto K R. Annu Rev Genet. 1985;19:209–252. - PubMed
- Schreiber S L. Science. 1991;251:283–287. - PubMed
- Jakoby W B, Wilchek M. Methods Enzymol. 1974;46:1–196. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources