Two separate signals act independently to localize a yeast late Golgi membrane protein through a combination of retrieval and retention - PubMed (original) (raw)
Two separate signals act independently to localize a yeast late Golgi membrane protein through a combination of retrieval and retention
N J Bryant et al. J Cell Biol. 1997.
Abstract
The localization of proteins to late-Golgi membranes (TGN) of Saccharomyces cerevisiae is conferred by targeting motifs containing aromatic residues in the cytosolic domains of these proteins. These signals could act by directing retrieval from a post-Golgi compartment or by preventing exit from the TGN. To investigate the mechanism of localization of yeast TGN proteins, we used the heterologous protein A-ALP (consisting of the cytosolic domain of dipeptidyl aminopeptidase A [DPAP A] fused to the transmembrane and luminal domains of the vacuolar protein alkaline phosphatase [ALP]), which localizes to the yeast TGN. Insertion of the aromatic residue-based TGN localization motif (FXFXD) of DPAP A into the cytosolic domain of ALP results in a protein that resides in the TGN. We demonstrate that the FXFXD motif confers Golgi localization through retrieval from a post-Golgi compartment by detecting a post-Golgi processed form of this protein in the TGN. We present an assay that uncouples retrieval-mediated Golgi localization from static retention-based localization, allowing measurement of the rate at which proteins exit the yeast TGN. We also demonstrate that the cytosolic domain of DPAP A contains additional information, separate from the retrieval motif, that slows exit from the TGN. We propose a model for DPAP A localization that involves two distinct mechanisms: one in which the FXFXD motif directs retrieval from a post-Golgi compartment, and a second that slows the rate at which DPAP A exits the TGN.
Figures
Figure 1
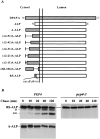
(A) Schematic illustration of modifications made within the cytosolic tail of DPAP A in the context of A-ALP. Deleted regions are indicated by a single line. (B) Immunoprecipitation of A-ALP from wild-type cells and RS-ALP from wild-type and pep4-3 cells. SNY36-9A (PEP4) cells harboring pSN55 (A-ALP) or pSN97 (RS-ALP) and SNY63 (pep4-3) cells harboring pSN97 (RS-ALP) were labeled with [35S]methionine for 10 min and chased by adding unlabeled methionine and cysteine each to a final concentration of 50 μg/ml. At the indicated times, the cells were spheroplasted and extracts were immunoprecipitated with a polyclonal antibody against ALP, followed by SDS-PAGE and fluorography. The product of the initial _PEP4_-dependent proteolysis is indicated by a single asterisk, with a double asterisk indicating a further breakdown product of this primary product. This breakdown pattern differs from that seen for A-ALP (which has only one _PEP4_-dependent breakdown product, e.g., Fig. 6), but clearly both of the lower bands shown here are produced in a _PEP4_-dependent manner.
Figure 6
Kinetics of processing of A-ALP, (F/A)A-ALP, RSALP, Vps10p, and Vps10p-10* (the recycling defective form of Vps10p) in wild-type and vps27Δ cells. SNY36-9A (wild-type) and NBY60 (vps27Δ) cells harboring pSN55 (A-ALP), pSN100 ((F/A)A-ALP) or pSN97 (RS-ALP), as well as NBY68 (vps1010*) and NBY69 (vps10-10* vps27Δ) cells were labeled with [35S]methionine for 10 min and chased by adding unlabeled methionine and cysteine, each to a final concentration of 50 μg/ml. At the indicated times, the cells were spheroplasted, and the extracts were immunoprecipitated with polyclonal antibodies against ALP and/or Vps10p, followed by SDS-PAGE and fluorography. The products of _PEP4_-dependent proteolysis are indicated using asterisks, as described in Fig. 1.
Figure 2
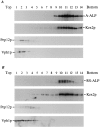
Sucrose gradient fractionation of A-ALP/RS-ALP containing membranes. High speed pellet membrane fractions (100,000 g) were prepared from 100 OD units of SNY36-9A cells harboring either pSN55 (A-ALP) or pSN97 (RS-ALP). These membranes were fractionated by a 20–50% sucrose density step gradient. After centrifugation, 14 × 1–ml fractions were collected from the top of the gradient, and the proteins were precipitated by the addition of TCA. Equal percentages of each fraction were subjected to immunoblot analysis after SDS-PAGE to detect the following proteins: A-ALP/RS-ALP using an anti-ALP mAb (1D3-A10); Kex2p was used as a marker protein for TGN membranes and detected using affinity-purified polyclonal antibodies; anti-Vph1p mAb 10D7-A7-B2 was used to detect Vph1p as a marker protein for vacuolar membranes; and Pep12p, a marker protein for membranes of the prevacuolar compartment, was detected using affinity-purified polyclonal antibodies.
Figure 3
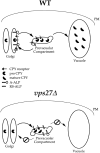
Model of Vps27p function in the retention of yeast TGN membrane proteins. TGN membrane proteins such as Vps10p, A-ALP, and RS-ALP continuously leave Golgi membranes and enter the PVC, from which they are retrieved via a mechanism that involves the recognition of an aromatic residue– containing motif. This retrieval requires Vps27p, and in vps27Δ cells, TGN membrane proteins accumulate in an exaggerated form of the PVC, the class E compartment (Raymond et al., 1992; Piper et al., 1995). The class E compartment is proteolytically active, and the rate at which proteolytically sensitive proteins are processed in vps27Δ cells can be used to measure the rate at which they enter this compartment. This is the basis for the assay to measure the leaving rates of A-ALP– and Vps10p–related proteins from the TGN. WT, wild type.
Figure 4
Induction of synthesis of Vps27p in vps27 cells. NBY67 (vps27) cells harboring either pSN55 (A-ALP), pSN100 ((F/A)AALP) or pSN97 (RS-ALP), and pHY5 (VPS27 under GAL1 control) (+) or empty vector (−) were grown in synthetic media containing 2% raffinose. At time t = 0, galactose was added to a final concentration of 2%, and cells were allowed to grow for an additional 90 min. At times t = 0 (− galactose) and t = 90 min (+ galactose), samples were removed and treated in the following ways (also see Fig. 5): (A) Whole-cell extracts were prepared from 10 OD units of NBY67 cells harboring pSN97 and pHY5 or empty vector to detect the presence of Vps27p using immunoblot analysis (similar results were obtained using the same cells harboring either pSN55 or pSN100 in conjunction with pHY5; data not shown); and (B) 0.5 OD units of cells were radiolabeled to determine whether the cells were secreting CPY.
Figure 5
Redistribution of A-ALP, (F/A)A-ALP, and RS-ALP upon induction of synthesis of Vps27p in vps27 cells. NBY67 (vps27) cells harboring either pSN55 (A-ALP), pSN100 ((F/A)A-ALP), or pSN97 (RS-ALP) and pHY5 (VPS27 under GAL1 control) (+) or empty vector (−) were grown in synthetic media containing 2% raffinose. At time t = 0, galactose was added to 2%, and cells were allowed to grow for an additional 90 min. At times (A) t = 0 (− galactose) and (B) t = 90 min (+ galactose) (also see Fig. 4), cells were fixed and prepared for immunofluorescence using anti-ALP antibodies in combination with biotin-linked secondary antibody and streptavidin-conjugated FITC, as well as anti-Vph1p antibodies in combination with rhodamine-conjugated goat anti–rabbit IgG (H + L).
Figure 7
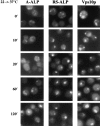
Redistribution of A-ALP, RS-ALP, and Vps10p in vps27-ts cells. RPY82 (vps27-ts) cells harboring either pSN55 (A-ALP) or pSN97 (RS-ALP) were grown in minimal media at 22°C, and then shifted to 37°C. Aliquots were removed and fixed at the times indicated. Fixed cells were spheroplasted and labeled with anti-ALP mAbs or affinity-purified anti-Vps10p polyclonal antibodies in combination with biotin-linked secondary antibody and streptavidin-conjugated FITC.
Figure 8
Kinetics of processing of (Δ2-11)A-ALP and (Δ68106)A-ALP in wild-type and vps27Δ cells. SNY36-9A (wild-type) and NBY60 (vps27Δ) cells harboring either pNB81 ((Δ2-11)AALP) or pSN27 ((Δ68-106)A-ALP) were labeled with [35S]methionine for 10 min and chased by adding unlabeled methionine and cysteine, each to a final concentration of 50 μg/ml. At the indicated times, the cells were spheroplasted, and the extracts were immunoprecipitated with a polyclonal antibody against ALP, followed by SDS-PAGE and fluorography. The products of _PEP4_-dependent proteolysis are indicated using asterisks, as described in Fig. 1.
Similar articles
- Retrieval of resident late-Golgi membrane proteins from the prevacuolar compartment of Saccharomyces cerevisiae is dependent on the function of Grd19p.
Voos W, Stevens TH. Voos W, et al. J Cell Biol. 1998 Feb 9;140(3):577-90. doi: 10.1083/jcb.140.3.577. J Cell Biol. 1998. PMID: 9456318 Free PMC article. - Membrane protein retention in the yeast Golgi apparatus: dipeptidyl aminopeptidase A is retained by a cytoplasmic signal containing aromatic residues.
Nothwehr SF, Roberts CJ, Stevens TH. Nothwehr SF, et al. J Cell Biol. 1993 Jun;121(6):1197-209. doi: 10.1083/jcb.121.6.1197. J Cell Biol. 1993. PMID: 8509444 Free PMC article. - The newly identified yeast GRD genes are required for retention of late-Golgi membrane proteins.
Nothwehr SF, Bryant NJ, Stevens TH. Nothwehr SF, et al. Mol Cell Biol. 1996 Jun;16(6):2700-7. doi: 10.1128/MCB.16.6.2700. Mol Cell Biol. 1996. PMID: 8649377 Free PMC article. - Multiple sorting pathways between the late Golgi and the vacuole in yeast.
Conibear E, Stevens TH. Conibear E, et al. Biochim Biophys Acta. 1998 Aug 14;1404(1-2):211-30. doi: 10.1016/s0167-4889(98)00058-5. Biochim Biophys Acta. 1998. PMID: 9714809 Review. - Protein transport from the late Golgi to the vacuole in the yeast Saccharomyces cerevisiae.
Bowers K, Stevens TH. Bowers K, et al. Biochim Biophys Acta. 2005 Jul 10;1744(3):438-54. doi: 10.1016/j.bbamcr.2005.04.004. Biochim Biophys Acta. 2005. PMID: 15913810 Review.
Cited by
- A novel mechanism for localizing membrane proteins to yeast trans-Golgi network requires function of synaptojanin-like protein.
Ha SA, Bunch JT, Hama H, DeWald DB, Nothwehr SF. Ha SA, et al. Mol Biol Cell. 2001 Oct;12(10):3175-90. doi: 10.1091/mbc.12.10.3175. Mol Biol Cell. 2001. PMID: 11598201 Free PMC article. - Vps52p, Vps53p, and Vps54p form a novel multisubunit complex required for protein sorting at the yeast late Golgi.
Conibear E, Stevens TH. Conibear E, et al. Mol Biol Cell. 2000 Jan;11(1):305-23. doi: 10.1091/mbc.11.1.305. Mol Biol Cell. 2000. PMID: 10637310 Free PMC article. - Golgi-to-late endosome trafficking of the yeast pheromone processing enzyme Ste13p is regulated by a phosphorylation site in its cytosolic domain.
Johnston HD, Foote C, Santeford A, Nothwehr SF. Johnston HD, et al. Mol Biol Cell. 2005 Mar;16(3):1456-68. doi: 10.1091/mbc.e04-07-0642. Epub 2005 Jan 12. Mol Biol Cell. 2005. PMID: 15647379 Free PMC article. - Global analysis of yeast endosomal transport identifies the vps55/68 sorting complex.
Schluter C, Lam KK, Brumm J, Wu BW, Saunders M, Stevens TH, Bryan J, Conibear E. Schluter C, et al. Mol Biol Cell. 2008 Apr;19(4):1282-94. doi: 10.1091/mbc.e07-07-0659. Epub 2008 Jan 23. Mol Biol Cell. 2008. PMID: 18216282 Free PMC article. - Cell-free transport to distinct Golgi cisternae is compartment specific and ARF independent.
Happe S, Weidman P. Happe S, et al. J Cell Biol. 1998 Feb 9;140(3):511-23. doi: 10.1083/jcb.140.3.511. J Cell Biol. 1998. PMID: 9456313 Free PMC article.
References
- Blum JS, Fiani ML, Stahl PD. Localization of cathepsin D in endosomes: characterization and biological importance. Adv Exp Med Biol. 1991;306:281–287. - PubMed
- Bretscher MS, Munro S. Cholesterol and the Golgi apparatus. Science (Wash DC) 1993;261:1280–1281. - PubMed
- Bryant NJ, Boyd A. Immunoisolation of Kex2p-containing organelles from yeast demonstrates colocalisation of three processing proteinases to a single Golgi compartment. J Cell Sci. 1993;106:815–822. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous