Activation of hypoxia-inducible factor 1alpha: posttranscriptional regulation and conformational change by recruitment of the Arnt transcription factor - PubMed (original) (raw)
Activation of hypoxia-inducible factor 1alpha: posttranscriptional regulation and conformational change by recruitment of the Arnt transcription factor
P J Kallio et al. Proc Natl Acad Sci U S A. 1997.
Abstract
In response to hypoxia the hypoxia-inducible factor-1 (HIF-1) mediates transcriptional activation of a network of genes encoding erythropoietin, vascular endothelial growth factor, and several glycolytic enzymes. HIF-1 consists of a heterodimer of two basic helix-loop-helix PAS (Per/Arnt/Sim) proteins, HIF-1alpha and Arnt. HIF-1alpha and Arnt mRNAs are constitutively expressed and were not altered upon exposure of HeLa or HepG2 cells to hypoxia, suggesting that the activity of the HIF-1alpha-Arnt complex may be regulated by some as yet unknown posttranscriptional mechanism. In support of this model, we demonstrate here that Arnt protein levels were not increased under conditions that induce an hypoxic response in HeLa and HepG2 cells. However, under identical conditions, HIF-1alpha protein levels were rapidly and dramatically up-regulated, as assessed by immunoblot analysis. In addition, HIF-1alpha acquired a new conformational state upon dimerization with Arnt, rendering HIF-1alpha more resistant to proteolytic digestion in vitro. Dimerization as such was not sufficient to elicit the conformational change in HIF-1alpha, since truncated forms of Arnt that are capable of dimerizing with HIF-1alpha did not induce this effect. Moreover, the high affinity DNA binding form of the HIF-1alpha-Arnt complex was only generated by forms of Arnt capable of eliciting the allosteric change in conformation. In conclusion, the combination of enhanced protein levels and allosteric change by dimerization defines a novel mechanism for modulation of transcription factor activity.
Figures
Figure 1
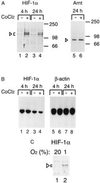
Hypoxic (CoCl2) induction of HIF-1α protein levels in HeLa cells. (A) HeLa cells were either treated with vehicle only (lanes 1, 3, and 5) or induced with 100 μM CoCl2 for 4 or 24 h, and whole cell extracts were prepared. Twenty-microgram aliquots of the extracts were separated on a 7.5% SDS/polyacrylamide gels and transferred onto a nitrocellulose filter. The transferred proteins were detected with rabbit anti-HIF-1α antiserum (1:400, lanes 1–4) or with rabbit anti-Arnt antibody (1:200 dilution, lanes 5 and 6) and visualized by chemiluminescence. Arrowheads indicate the specific HIF-1α and Arnt complexes. (B) Total cellular RNA was prepared from duplicate samples corresponding to lanes 1 to 4 in A and analyzed by RNA (Northern) blot for HIF-1α mRNA levels. As a control, the same blot was rehybridized with a β-actin probe (lanes 5–8). (C) Induction of HIF-1α protein levels by hypoxia. HepG2 cells were incubated under normoxic (20% oxygen, lane 1) or hypoxic (1% oxygen, lane 2) for 4 h, and whole cell extracts were prepared. Immunoblot analysis was performed as in A.
Figure 2
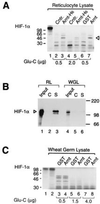
Arnt induces a conformational change in HIF-1α. (A) HIF-1α was in vitro translated in reticulocyte lysate and allowed to interact with vaccinia virus-expressed, histidine-tagged, and purified Arnt (Arnt-H6) or control (Cntr.) extracts (lanes 2–5) or with bacterially expressed, purified GST, or GST–Arnt fusion protein (lanes 6 and 7) at room temperature for 1 h. A total of 0.5 μg (lanes 2, 3, 6, and 7) or 2 μg (lanes 4 and 5) of S. aureus Glu-C protease was added, and the samples were incubated for 30 min at 30°C. Reaction products were run on a 12.5% SDS/polyacrylamide gel and visualized by fluorography. The Arnt-induced protease-resistant product is indicated by an arrowhead. Lane 1 represents full-length HIF-1α with no added protease. (B) Wheat germ lysate-expressed HIF-1α is not associated with hsp90. [35S]Methionine labeled full length HIF-1α, synthesized either in reticulocyte (RL) or in wheat germ lysate (WGL), was coimmunoprecipitated with either monoclonal anti-hsp90 (lanes S) IgM antibody or control (lanes C) IgM antibody as described. Immunoprecipitated proteins were analyzed by 7.5% SDS/PAGE and fluorography. Lanes 1 and 4 represent 10% of the inputs of in vitro synthesized HIF-1α. (C) Wheat germ lysate-expressed HIF-1α does not undergo an allosteric change in the presence of Arnt. Wheat germ lysate-translated HIF-1α (lane 2) was treated with either 0.5 μg (lanes 3 and 4), 1.5 μg (lanes 5 and 6), or 4.0 μg (lanes 7 and 8) Glu-C protease for 30 min at 30°C in the presence of GST (lanes 3, 5, and 7) or full-length GST–Arnt (lanes 4, 6 and 8). The reaction products were separated on a 12.5% SDS/polyacrylamide gel and analyzed by fluorography. Lane 1 represents HIF-1α expressed in reticulocyte lysate.
Figure 3
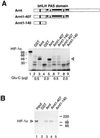
HIF-1α does not undergo an allosteric change in response to the GST–Arnt1-407 or GST–Arnt1-140 deletion mutants. (A) The Upper panel shows a schematic representation of full-length Arnt and the bacterially expressed Arnt bHLH and bHLH/PAS fragments Arnt1-140 and Arnt1-407, respectively. Full-length, [35S]methionine-labeled HIF-1α was expressed in reticulocyte lysate and incubated with either GST (lanes 2 and 4), full-length GST–Arnt (lanes 3 and 5), GST–Arnt1-407 (lanes 6 and 8), or GST–Arnt1-140 (lanes 7 and 9). Glu-C protease (0.5 μg: lanes 2, 3, 6, and 7; or 2.0 μg: lanes 4, 5, 8, and 9) was added and the incubation continued for an additional 30 min before analysis by SDS/PAGE and fluorography. (B) The bHLH and PAS regions of Arnt are sufficient for dimerization with HIF-1α. Ten microliters aliquots of in vitro synthetized and [35S]methionine-labeled HIF-1α expressed in reticulocyte lysate were incubated with either purified GST (lane 2) or GST–Arnt fusion proteins (lanes 3–5) for 1 h at room temperature. Twenty microliters of glutathione-Sepharose was added, and the samples were incubated on ice for 1 h. The precipitated pellet was analyzed on a 7.5% SDS/polyacrylamide gel. Lane 1 represents 10% of the HIF-1α input.
Figure 4
Full-length HIF-1α forms a DNA-binding complex with bacterially expressed Arnt. Six microliters of reticulocyte lysate-expressed HIF-1α was allowed to interact with either GST (lane 3), full-length Arnt (lanes 4–6, 9, and 10), Arnt1-407, or Arnt1-140 (lanes 7 and 8), respectively. In lanes 5 and 6, a 100-fold molar excess of unlabeled competitor DNA was added (specific and nonspecific, respectively), whereas in lanes 9 and 10 control (C) or specific (S) anti-HIF-1α antibodies were added to the reactions. The arrowhead indicates the HIF-1α–Arnt complex, whereas the asterisks denote nonspecific protein DNA complexes formed with unprogrammed reticulocyte lysate only (lane 2). Lane 1 represents free probe in the absence of any added protein. (B) The dioxin receptor forms a DNA binding complex with either full-length Arnt or Arnt1-407. Hepa 1c1c7 C4 cell cytosol was treated with 5 nM dioxin for 2 h to activate the dioxin receptor. Forty micrograms of dioxin-treated cytosol was incubated with GST (lane 3), full-length Arnt, Arnt1-407, or Arnt1-140 (lanes 4–6, respectively) at room temperature for 45 min. A 32P-labeled xenobiotic response element (XRE)-containing oligonucleotide was added as a probe and incubation continued for additional 15 min before electrophoresis. Lane 1, 32P-labeled XRE alone; lane 2, 32P-labeled XRE in the presence of the cytosolic extract without any added recombinant protein. Dioxin receptor–Arnt complexes are indicated by arrowheads.
Figure 5
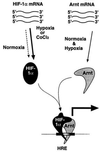
Model for activation of HIF-1 function. Both Arnt mRNA and protein are constitutively expressed. In contrast, hypoxia treatment results in dramatically elevated HIF-1α protein levels generated from a constitutively expressed pool of mRNA. HIF-1α–Arnt heterodimers then form, and dimerization induces a conformational change that allows the HIF-1α–Arnt complex to bind DNA. It is also possible that the structural change is propagated onto other functional domains of HIF-1α, most notably the transactivation domain. The present data indicate that the molecular chaperone hsp90 associates with de novo synthesized HIF-1α and gives rise to a conformation that is critical for dimerization with Arnt.
Similar articles
- Functional interference between hypoxia and dioxin signal transduction pathways: competition for recruitment of the Arnt transcription factor.
Gradin K, McGuire J, Wenger RH, Kvietikova I, fhitelaw ML, Toftgård R, Tora L, Gassmann M, Poellinger L. Gradin K, et al. Mol Cell Biol. 1996 Oct;16(10):5221-31. doi: 10.1128/MCB.16.10.5221. Mol Cell Biol. 1996. PMID: 8816435 Free PMC article. - The role of the aryl hydrocarbon receptor nuclear translocator (ARNT) in hypoxic induction of gene expression. Studies in ARNT-deficient cells.
Wood SM, Gleadle JM, Pugh CW, Hankinson O, Ratcliffe PJ. Wood SM, et al. J Biol Chem. 1996 Jun 21;271(25):15117-23. doi: 10.1074/jbc.271.25.15117. J Biol Chem. 1996. PMID: 8662957 - Insulin induces transcription of target genes through the hypoxia-inducible factor HIF-1alpha/ARNT.
Zelzer E, Levy Y, Kahana C, Shilo BZ, Rubinstein M, Cohen B. Zelzer E, et al. EMBO J. 1998 Sep 1;17(17):5085-94. doi: 10.1093/emboj/17.17.5085. EMBO J. 1998. PMID: 9724644 Free PMC article. - Structural and functional analysis of hypoxia-inducible factor 1.
Semenza GL, Agani F, Booth G, Forsythe J, Iyer N, Jiang BH, Leung S, Roe R, Wiener C, Yu A. Semenza GL, et al. Kidney Int. 1997 Feb;51(2):553-5. doi: 10.1038/ki.1997.77. Kidney Int. 1997. PMID: 9027737 Review. - Hypoxia and high altitude. The molecular response.
Höpfl G, Ogunshola O, Gassmann M. Höpfl G, et al. Adv Exp Med Biol. 2003;543:89-115. doi: 10.1007/978-1-4419-8997-0_7. Adv Exp Med Biol. 2003. PMID: 14713116 Review.
Cited by
- The Three A's in Asthma - Airway Smooth Muscle, Airway Remodeling & Angiogenesis.
Keglowich LF, Borger P. Keglowich LF, et al. Open Respir Med J. 2015 Jun 17;9:70-80. doi: 10.2174/1874306401509010070. eCollection 2015. Open Respir Med J. 2015. PMID: 26106455 Free PMC article. - Hypoxia and cytokines regulate carbonic anhydrase 9 expression in hepatocellular carcinoma cells in vitro.
Kockar F, Yildrim H, Sagkan RI, Hagemann C, Soysal Y, Anacker J, Hamza AA, Vordermark D, Flentje M, Said HM. Kockar F, et al. World J Clin Oncol. 2012 Jun 10;3(6):82-91. doi: 10.5306/wjco.v3.i6.82. World J Clin Oncol. 2012. PMID: 22724087 Free PMC article. - HIF-1α and RKIP: a computational approach for pancreatic cancer therapy.
Srivani G, Behera SK, Dariya B, Chalikonda G, Alam A, Nagaraju GP. Srivani G, et al. Mol Cell Biochem. 2020 Sep;472(1-2):95-103. doi: 10.1007/s11010-020-03788-6. Epub 2020 Jun 19. Mol Cell Biochem. 2020. PMID: 32562168 - Inhibition of N-Myc down regulated gene 1 in in vitro cultured human glioblastoma cells.
Said HM, Polat B, Stein S, Guckenberger M, Hagemann C, Staab A, Katzer A, Anacker J, Flentje M, Vordermark D. Said HM, et al. World J Clin Oncol. 2012 Jul 10;3(7):104-10. doi: 10.5306/wjco.v3.i7.104. World J Clin Oncol. 2012. PMID: 22787578 Free PMC article. - Up-regulation of hypoxia inducible factor-1α by cobalt chloride correlates with proliferation and apoptosis in PC-2 cells.
Dai ZJ, Gao J, Ma XB, Yan K, Liu XX, Kang HF, Ji ZZ, Guan HT, Wang XJ. Dai ZJ, et al. J Exp Clin Cancer Res. 2012 Mar 27;31(1):28. doi: 10.1186/1756-9966-31-28. J Exp Clin Cancer Res. 2012. PMID: 22453051 Free PMC article.
References
- Bunn H F, Poyton R O. Physiol Rev. 1996;76:839–885. - PubMed
- Goldberg M A, Dunning S P, Bunn H F. Science. 1988;242:1412–1415. - PubMed
- Wang G L, Semenza G L. Curr Opin Hematol. 1996;3:156–162. - PubMed
- Hankinson O. Annu Rev Pharmacol Toxicol. 1995;35:307–340. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous