The human homologue of Saccharomyces cerevisiae Gle1p is required for poly(A)+ RNA export - PubMed (original) (raw)
The human homologue of Saccharomyces cerevisiae Gle1p is required for poly(A)+ RNA export
J L Watkins et al. Proc Natl Acad Sci U S A. 1998.
Abstract
The mechanism of mRNA export is a complex issue central to cellular physiology. We characterized previously yeast Gle1p, a protein with a leucine-rich (LR) nuclear export sequence (NES) that is essential for poly(A)+ RNA export in Saccharomyces cerevisiae. To characterize elements of the vertebrate mRNA export pathway, we identified a human homologue of yeast Gle1p and analyzed its function in mammalian cells. hGLE1 encodes a predicted 75-kDa polypeptide with high sequence homology to yeast Gle1p, but hGle1p does not contain a sequence motif matching any of the previously characterized NESs. hGLE1 can complement a yeast gle1 temperature-sensitive export mutant only if a LR-NES is inserted into it. To determine whether hGle1p played a role in nuclear export, anti-hGle1p antibodies were microinjected into HeLa cells. In situ hybridization of injected cells showed that poly(A)+ RNA export was inhibited. In contrast, there was no effect on the nuclear import of a glucocorticoid receptor reporter. We conclude that hGle1p functions in poly(A)+ RNA export, and that human cells facilitate such export with a factor similar to yeast but without a recognizable LR-NES. With hGle1p localized at the nuclear pore complexes, hGle1p is positioned to act at a terminal step in the export of mature RNA messages to the cytoplasm.
Figures
Figure 1
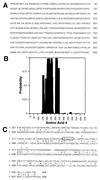
Amino acid sequence of hGle1p. (A) hGle1p amino acid sequence. (B) Coiled-coil probability for hGle1p. A middle region of hGle1p shows a high probability of forming a coiled-coil structure by analysis with the
coils
program (48). (C) Comparison of the C-terminal regions of yeast and human Gle1p.
align
analysis (43) between the yeast (upper line) and human proteins (lower line) reveals significant homology. The center line designates identical (capital letter) and conserved (: or .) residues. The LR-NES in yeast Gle1p is boxed. The slash (/) designates the breakpoint for Regions 3 and 4 (Fig. 3_A_). The plus signs (+) mark residues changed in yeast gle1 alleles (P380G or G384R).
Figure 2
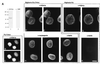
hGle1p localizes to the NPC and is cytoplasmically accessible. (A) Immunoblot analysis with whole-cell HeLa extracts and affinity-purified rabbit antibodies: anti-hGle1p (lane 1) or anti-MBP (lane 2). Molecular mass markers are in kDa. (B_–_K) Intracellular distribution of hGle1p. HeLa cells were processed for labeling with primary antibodies as designated: Fix-Triton (B and C), Digitonin-Fix-Triton (D, E, and H_–_J), or Digitonin-Fix (F, G, and K). hGle1p localization was observed with the affinity-purified rabbit anti-hGle1p antibody (B_–_G), nucleoporin staining (H and I) with mAb414 (40), and lamin staining with a rabbit anti-lamin B serum (J and K) (41). Pairs of cells (B_–_C, D_–_E, H_–_I, and F_–_G) were photographed in two different focal planes for the punctate (C, D, H, and F) and nuclear rim (B, E, G, and I) staining. (Bars = 10 μm.)
Figure 3
Complementation analysis of yeast gle1–4 cells with hGLE1 chimeras. (A) Schematic representation of the structural regions in yeast (red) and human (blue) Gle1p proteins, with the LR-NES designated (yellow). (B) Yeast gle1–4 strains expressing the designated yeast deletion (Δ) or yeast–human chimeric Gle1p were grown at 23°C (Right) or 30°C (Left). Serial dilutions (from left to right) of equivalent cell numbers were plated on synthetic minimal medium lacking leucine. Expression of full-length yeast Gle1p supports growth at 30°C (top row), whereas vector alone does not (second row). Immunoblot analysis confirmed expression of all polypeptides (data not shown).
Figure 4
Injection of anti-hGle1p antibodies inhibits poly(A)+ RNA export in HeLa cells. The distribution of poly(A)+ RNA was examined in HeLa cells co-microinjected with Texas red–dextran and either affinity-purified rabbit anti-hGle1p antibody alone (2.7 mg/ml, Left) or anti-hGle1p premixed with purified GST-hGle1p (2.7 mg/ml IgG and 2.25 mg/ml recombinant, Right). The anti-hGle1p antibody is monospecific for hGle1p (see Fig. 2_A_). Only a subset of the cells in a given field were injected, and all the cells were cytoplasmically injected except for the uppermost injected cell in the Right column (by Texas red localization, Middle). Cells were incubated for 12 hr at 37°C before in situ hybridization with a digoxigenin-labeled oligo(dT30) probe and fluorescein isothiocyanate antidigoxigenin Fab. Arrowhead highlights a typical injected cell, and arrow indicates an uninjected cell (Top). Nuclear DNA was stained with 4′,6-diamidino-2-phenylindole (Bottom). (Bar = 20 μm.)
Figure 5
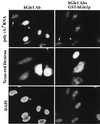
Nuclear import is not affected by microinjection of anti-hGle1p antibody. HeLa cells transiently expressing GR-GFP (A, D, G, and J) were co-microinjected with Texas red–dextran and either affinity-purified rabbit antibody: anti-hGle1p (2 mg/ml, A_–_F) or anti-MBP (2 mg/ml, G_–_L). The anti-MBP antibody served as a control for nonspecific effects of antibody injection. Texas red–dextran (Center, B, E, H, and K) differentiates between injected and uninjected cells. In a given field, only a subset of the cells were cytoplasmically injected. Cells were incubated for 12 hr at 37°C, and those in D_–_F and J_–_L were treated with medium containing 10 μg/ml dexamethasone for 30 min before fixation and processing for fluorescence microscopy. Cells in A_–_C and G_–_I were untreated. Nuclear DNA was stained with 4′,6-diamidino-2-phenylindole (C, F, I, and L). (Bar = 20 μm.)
Similar articles
- The RNA export factor Gle1p is located on the cytoplasmic fibrils of the NPC and physically interacts with the FG-nucleoporin Rip1p, the DEAD-box protein Rat8p/Dbp5p and a new protein Ymr 255p.
Strahm Y, Fahrenkrog B, Zenklusen D, Rychner E, Kantor J, Rosbach M, Stutz F. Strahm Y, et al. EMBO J. 1999 Oct 15;18(20):5761-77. doi: 10.1093/emboj/18.20.5761. EMBO J. 1999. PMID: 10610322 Free PMC article. - An RNA-export mediator with an essential nuclear export signal.
Murphy R, Wente SR. Murphy R, et al. Nature. 1996 Sep 26;383(6598):357-60. doi: 10.1038/383357a0. Nature. 1996. PMID: 8848052 - The NES-Crm1p export pathway is not a major mRNA export route in Saccharomyces cerevisiae.
Neville M, Rosbash M. Neville M, et al. EMBO J. 1999 Jul 1;18(13):3746-56. doi: 10.1093/emboj/18.13.3746. EMBO J. 1999. PMID: 10393189 Free PMC article. - Rat8p/Dbp5p is a shuttling transport factor that interacts with Rat7p/Nup159p and Gle1p and suppresses the mRNA export defect of xpo1-1 cells.
Hodge CA, Colot HV, Stafford P, Cole CN. Hodge CA, et al. EMBO J. 1999 Oct 15;18(20):5778-88. doi: 10.1093/emboj/18.20.5778. EMBO J. 1999. PMID: 10523319 Free PMC article. - Movement of macromolecules between the cytoplasm and the nucleus in yeast.
Bossie MA, Silver PA. Bossie MA, et al. Curr Opin Genet Dev. 1992 Oct;2(5):768-74. doi: 10.1016/s0959-437x(05)80137-6. Curr Opin Genet Dev. 1992. PMID: 1458025 Review.
Cited by
- The importance of managing the patient and not the gene: expanded phenotype of _GLE1_-associated arthrogryposis.
Tan QK, McConkie-Rosell A, Juusola J, Gustafson KE, Pizoli CE, Buckley AF, Jiang YH. Tan QK, et al. Cold Spring Harb Mol Case Stud. 2017 Nov 21;3(6):a002063. doi: 10.1101/mcs.a002063. Print 2017 Nov. Cold Spring Harb Mol Case Stud. 2017. PMID: 28729373 Free PMC article. - mRNA nuclear export and human disease.
Hurt JA, Silver PA. Hurt JA, et al. Dis Model Mech. 2008 Sep-Oct;1(2-3):103-8. doi: 10.1242/dmm.000745. Dis Model Mech. 2008. PMID: 19048072 Free PMC article. - Proteomic Profiles and Protein Network Analysis of Primary Human Leukocytes Revealed Possible Clearance Biomarkers for Staphylococcus aureus Infection.
Sirichoat A, Kaewseekhao B, Nithichanon A, Roytrakul S, Faksri K. Sirichoat A, et al. Curr Microbiol. 2023 Sep 4;80(10):335. doi: 10.1007/s00284-023-03450-6. Curr Microbiol. 2023. PMID: 37665379 - Nup42 and IP6 coordinate Gle1 stimulation of Dbp5/DDX19B for mRNA export in yeast and human cells.
Adams RL, Mason AC, Glass L, Aditi, Wente SR. Adams RL, et al. Traffic. 2017 Dec;18(12):776-790. doi: 10.1111/tra.12526. Epub 2017 Oct 16. Traffic. 2017. PMID: 28869701 Free PMC article. - Survival beyond the perinatal period expands the phenotypes caused by mutations in GLE1.
Said E, Chong JX, Hempel M, Denecke J, Soler P, Strom T, Nickerson DA, Kubisch C; University of Washington Center for Mendelian Genomics; Bamshad MJ, Lessel D. Said E, et al. Am J Med Genet A. 2017 Nov;173(11):3098-3103. doi: 10.1002/ajmg.a.38406. Epub 2017 Sep 8. Am J Med Genet A. 2017. PMID: 28884921 Free PMC article.
References
- Lee M S, Silver P A. Curr Opin Genet Dev. 1997;7:212–219. - PubMed
- Nakielny S, Dreyfuss G. Curr Opin Cell Biol. 1997;9:420–429. - PubMed
- Ullman K S, Powers M A, Forbes D J. Cell. 1997;90:967–970. - PubMed
- Legrain P, Rosbash M. Cell. 1989;57:573–583. - PubMed
- Chang D D, Sharp P A. Cell. 1989;59:789–795. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases