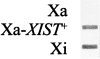Stabilization and localization of Xist RNA are controlled by separate mechanisms and are not sufficient for X inactivation - PubMed (original) (raw)
Stabilization and localization of Xist RNA are controlled by separate mechanisms and are not sufficient for X inactivation
C M Clemson et al. J Cell Biol. 1998.
Abstract
These studies address whether XIST RNA is properly localized to the X chromosome in somatic cells where human XIST expression is reactivated, but fails to result in X inactivation (Tinker, A.V., and C.J. Brown. 1998. Nucl. Acids Res. 26:2935-2940). Despite a nuclear RNA accumulation of normal abundance and stability, XIST RNA does not localize in reactivants or in naturally inactive human X chromosomes in mouse/ human hybrid cells. The XIST transcripts are fully stabilized despite their inability to localize, and hence XIST RNA localization can be uncoupled from stabilization, indicating that these are separate steps controlled by distinct mechanisms. Mouse Xist RNA tightly localized to an active X chromosome, demonstrating for the first time that the active X chromosome in somatic cells is competent to associate with Xist RNA. These results imply that species-specific factors, present even in mature, somatic cells that do not normally express Xist, are necessary for localization. When Xist RNA is properly localized to an active mouse X chromosome, X inactivation does not result. Therefore, there is not a strict correlation between Xist localization and chromatin inactivation. Moreover, expression, stabilization, and localization of Xist RNA are not sufficient for X inactivation. We hypothesize that chromosomal association of XIST RNA may initiate subsequent developmental events required to enact transcriptional silencing.
Figures
Figure 1
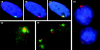
Fluorescence in situ hybridization detection of XIST RNA in normal human and mouse/human hybrid cells. Digoxigenin and biotinylated probes were hybridized in situ and detected with fluorochrome-conjugated avidin or antidigoxigenin antibody. Fluorochromes used were FITC (green), rhodamine (red), and DAPI (blue). (A) In normal human diploid fibroblasts (46, XX WI-38 cells), the XIST RNA (green) occupies a discrete location in the nucleus. (B) The XIST RNA has a similar shape to the X chromosome territory defined by the whole X chromosome library signal (red; the weak red signal in the middle of this cell is the active X chromosome which is out of the plane of focus). (C) The XIST RNA essentially paints the X chromosome in these normal nuclei (overlap is yellow). (D) In mouse/human hybrids containing a single active human X chromosome (AHA-A5) that expresses XIST through treatment with the demethylating agent 5azadC, the XIST RNA (red) is disperse, and spreads throughout the nucleus. (E) The XIST RNA (red) does not associate or strictly localize to the human whole X chromosome library signal (green) in the hybrid cells. (F) In mouse/human hybrid cells (t86-B1maz1b-3a) that contain a single human inactive X that expresses XIST endogenously (no 5azadC treatment), the human XIST RNA (red) also shows aberrant localization to the whole X chromosome library signal (green).
Figure 2
Reactivated and endogenous XIST expression is similar in hybrid cells. Total cellular RNA from the following cells were transferred to nitrocellulose and hybridized with an XIST cDNA probe: AHA-11aB1, the active X-containing hybrid cells (Xa); AHA-2C-5C-9, the demethylated derivative that ectopically expresses XIST (Xa-XIST +); and t11-4Aaz5 cells containing a human inactive X chromosome that normally expresses XIST (Xi). The blots were scanned, and the density of signal from the Xi and Xa-XIST+ clones were essentially identical, demonstrating that the RNA is produced in similar quantities from endogenously expressing and demethylated XIST genes.
Figure 3
Stability of the XIST RNA after treatment of cells with actinomycin. Cells were harvested after 0–8 h of actinomycin treatment (A) or control DMSO treatment (D) as the actinomycin is dissolved in DMSO. RT-PCR was performed on the RNA isolated from these cells with primers for XIST (A and B) and actin (not shown). The RT-PCR products for a female lymphoblast (GM07350; A) and an inactive X-containing human/ mouse somatic cell hybrid (t75-2maz34-1a; B) are shown, and the ratio of XIST product to actin product (normalized to the ratio of XIST to actin for the control DMSO treatments) is plotted in C. The black line shows the ratio for the female cells, while the grey line shows the ratio for the hybrid cell line.
Figure 4
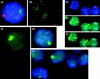
In situ RNA and murine X whole chromosome library detection in AS-2B mouse/human hybrid cells that contain both an active human and mouse X chromosome. Both human and murine Xist RNA is expressed from these chromosomes as a result of 5azadC treatment. (A and B) The difference between the reactivated murine Xist RNA and human XIST RNA is illustrated in these cells, with the murine RNA appearing highly localized (red) and the human XIST RNA (green) showing a focus of expression (most likely representing the site of transcription) with particles that drift away into the nucleoplasm. Reactivated murine Xist RNA localizes to the active X chromosome (C–E). (C) The murine Xist RNA (red) is discretely contained in the hybrid cell nucleus (D) in a pattern similar to the murine X chromosome signal (green; the mouse X chromosome library cross-reacts slightly with the human X chromosome, giving a second green signal apparent in the middle cell in D); (E) the murine Xist RNA colocalizes with the active murine X chromosome (overlap is yellow).Figure 5. The mouse X chromosome painted by the Xist RNA does not inactivate. Codetection of murine Xist RNA (green) and Zfx RNA or Pgk1 RNA (red) in undenatured 1B-5C-3B cells. (A and B) Zfx RNA (red) is expressed from cells that also express mouse Xist RNA (green); (C) Pgk1 RNA expression (red) is also expressed from chromosomes that express Xist RNA (green), suggesting that the active X chromosome is not inactivated as a result of Xist localization. Expression of mouse genes appears to be unaffected in cells that express mouse Xist. (A) There is a similar pattern and intensity of expression of Zfx RNA in cells that express mouse Xist (top), and those that do not (bottom). (C). Similarly, there is comparable Pgk1 expression in cells that express mouse Xist (left) and cells that do not (far right).
Similar articles
- Reactivation of XIST in normal fibroblasts and a somatic cell hybrid: abnormal localization of XIST RNA in hybrid cells.
Hansen RS, Canfield TK, Stanek AM, Keitges EA, Gartler SM. Hansen RS, et al. Proc Natl Acad Sci U S A. 1998 Apr 28;95(9):5133-8. doi: 10.1073/pnas.95.9.5133. Proc Natl Acad Sci U S A. 1998. PMID: 9560241 Free PMC article. - The human XIST gene: analysis of a 17 kb inactive X-specific RNA that contains conserved repeats and is highly localized within the nucleus.
Brown CJ, Hendrich BD, Rupert JL, Lafrenière RG, Xing Y, Lawrence J, Willard HF. Brown CJ, et al. Cell. 1992 Oct 30;71(3):527-42. doi: 10.1016/0092-8674(92)90520-m. Cell. 1992. PMID: 1423611 - Xist has properties of the X-chromosome inactivation centre.
Herzing LB, Romer JT, Horn JM, Ashworth A. Herzing LB, et al. Nature. 1997 Mar 20;386(6622):272-5. doi: 10.1038/386272a0. Nature. 1997. PMID: 9069284 - X chromosome inactivation and the Xist gene.
Brockdorff N, Duthie SM. Brockdorff N, et al. Cell Mol Life Sci. 1998 Jan;54(1):104-12. doi: 10.1007/s000180050129. Cell Mol Life Sci. 1998. PMID: 9487391 Free PMC article. Review. - The cell biology of a novel chromosomal RNA: chromosome painting by XIST/Xist RNA initiates a remodeling cascade.
Hall LL, Lawrence JB. Hall LL, et al. Semin Cell Dev Biol. 2003 Dec;14(6):369-78. doi: 10.1016/j.semcdb.2003.09.011. Semin Cell Dev Biol. 2003. PMID: 15015744 Review.
Cited by
- XIST RNA exhibits nuclear retention and exhibits reduced association with the export factor TAP/NXF1.
Cohen HR, Panning B. Cohen HR, et al. Chromosoma. 2007 Aug;116(4):373-83. doi: 10.1007/s00412-007-0100-1. Epub 2007 Mar 1. Chromosoma. 2007. PMID: 17333237 - The X chromosome is organized into a gene-rich outer rim and an internal core containing silenced nongenic sequences.
Clemson CM, Hall LL, Byron M, McNeil J, Lawrence JB. Clemson CM, et al. Proc Natl Acad Sci U S A. 2006 May 16;103(20):7688-93. doi: 10.1073/pnas.0601069103. Epub 2006 May 8. Proc Natl Acad Sci U S A. 2006. PMID: 16682630 Free PMC article. - XIST RNA and architecture of the inactive X chromosome: implications for the repeat genome.
Hall LL, Lawrence JB. Hall LL, et al. Cold Spring Harb Symp Quant Biol. 2010;75:345-56. doi: 10.1101/sqb.2010.75.030. Epub 2011 Mar 29. Cold Spring Harb Symp Quant Biol. 2010. PMID: 21447818 Free PMC article. Review. - Derivation of consensus inactivation status for X-linked genes from genome-wide studies.
Balaton BP, Cotton AM, Brown CJ. Balaton BP, et al. Biol Sex Differ. 2015 Dec 30;6:35. doi: 10.1186/s13293-015-0053-7. eCollection 2015. Biol Sex Differ. 2015. PMID: 26719789 Free PMC article. - Ordered chromatin changes and human X chromosome reactivation by cell fusion-mediated pluripotent reprogramming.
Cantone I, Bagci H, Dormann D, Dharmalingam G, Nesterova T, Brockdorff N, Rougeulle C, Vallot C, Heard E, Chaligne R, Merkenschlager M, Fisher AG. Cantone I, et al. Nat Commun. 2016 Aug 10;7:12354. doi: 10.1038/ncomms12354. Nat Commun. 2016. PMID: 27507283 Free PMC article.
References
- Brockdorff N, Ashworth A, Kay GF, McCabe VM, Norris DP, Cooper PJ, Swift S, Rastan S. The product of the mouse Xist gene is a 15 kb inactive X-specific transcript containing no conserved ORF and located in the nucleus. Cell. 1992;71:515–526. - PubMed
- Brown CJ, Ballabio A, Rupert JL, Lafreniere RG, Grompe M, Tonlorenzi R, Willard HF. A gene from the region of the human X inactivation centre is expressed exclusively from the inactive X chromosome. Nature. 1991;349:38–44. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases