Expansion and contraction of ribosomal DNA repeats in Saccharomyces cerevisiae: requirement of replication fork blocking (Fob1) protein and the role of RNA polymerase I - PubMed (original) (raw)
Expansion and contraction of ribosomal DNA repeats in Saccharomyces cerevisiae: requirement of replication fork blocking (Fob1) protein and the role of RNA polymerase I
T Kobayashi et al. Genes Dev. 1998.
Abstract
Saccharomyces cerevisiae carries approximately 150 copies of rDNA in tandem repeats. It was found that the absence of an essential subunit of RNA polymerase I (Pol I) in rpa135 deletion mutants triggers a gradual decrease in rDNA repeat number to about one-half the normal level. Reintroduction of the missing RPA135 gene induced a gradual increase in repeat number back to the normal level. Gene FOB1 was shown to be essential for both the decrease and increase of rDNA repeats. FOB1 was shown previously to be required for replication fork blocking (RFB) activity at RFB site in rDNA and for recombination hot-spot (HOT1) activity. Thus, DNA replication fork blockage appears to stimulate recombination and play an essential role in rDNA expansion/contraction and sequence homogenization, and possibly, in the instability of repeated sequences in general. RNA Pol I, on the other hand, appears to control repeat numbers, perhaps by stabilizing rDNA with the normal repeat numbers as a stable nucleolar structure.
Figures
Figure 1
Structure of rDNA repeats in S. cerevisiae. Locations of the 35S and 5S rRNA genes (the direction of transcription indicated by arrows), the two nontranscribed spacer regions (NTS1 and NTS2), ARS (replication origin), and the HOT1 I element are shown at top. _Bgl_II-A and B DNA fragments are also shown. NTS1 and its surrounding regions are expanded. Three solid bars represent the HOT1 E element, Pol I enhancer, and RFB site (also indicated by  ). Two open rectangles, indicated as 5S and NTS, are DNA regions used for the competitive PCR assay.
). Two open rectangles, indicated as 5S and NTS, are DNA regions used for the competitive PCR assay.
Figure 2
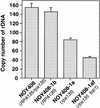
Southern hybridization analysis of rDNA copy numbers. The RPA135 gene was introduced by transformation into strains NOY408-1a (rpa135), NOY408-1af (rpa135 fob1), and NOY408-1af derivatives carrying FOB1 on a plasmid or control vector plasmid as indicated. DNA samples were prepared at 44 (lanes 2,6,9,12), 80 (lanes 3,7,10,13), and 116 generations (lanes 4,8,11,14) after introduction of RPA135. NOY408-1a, and NOY408-1af, which did not receive RPA135 (lanes 1,5, respectively), were also analyzed together with control strains NOY408 (lane 15) and NOY408-1b (lane 16). In addition to rDNA, a single copy gene, MCM2, was analyzed as a reference.
Figure 3
Absolute rDNA copy numbers determined by a competitive PCR assay. Values given are rDNA copy numbers per haploid genome determined as described in Materials and Methods and text.
Figure 4
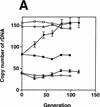
Pol I- and FOB1_-dependent increases of rDNA copy numbers. Experiments similar to the ones shown in Fig. 2 were carried out. (A) NOY408-1a (rpa135) and (B) NOY408-1af (rpa135 fob1), as well as their derivatives with (+YEp–_FOB1) and without (+YEplac195) the FOB1 gene were analyzed (YEp–FOB1 and YEplac195 are indicated as +pFOB1 and −pFOB1, respectively). Strains that contained RPA135 on plasmid pNOY117 are indicated by the strains’ designation, followed by +pRPA135.NOY408 (RPA135/rpa135) and NOY408-1b (RPA135) were also analyzed. Copy numbers of rDNA relative to the single copy gene MCM2 were analyzed at various generations after introduction of RPA135. Four independent transformants were analyzed. Values shown are the averages obtained for the four cultures (standard deviations are shown). Control cultures, which did not receive RPA135, were also analyzed in parallel (standard deviations are not shown).
Figure 4
Pol I- and FOB1_-dependent increases of rDNA copy numbers. Experiments similar to the ones shown in Fig. 2 were carried out. (A) NOY408-1a (rpa135) and (B) NOY408-1af (rpa135 fob1), as well as their derivatives with (+YEp–_FOB1) and without (+YEplac195) the FOB1 gene were analyzed (YEp–FOB1 and YEplac195 are indicated as +pFOB1 and −pFOB1, respectively). Strains that contained RPA135 on plasmid pNOY117 are indicated by the strains’ designation, followed by +pRPA135.NOY408 (RPA135/rpa135) and NOY408-1b (RPA135) were also analyzed. Copy numbers of rDNA relative to the single copy gene MCM2 were analyzed at various generations after introduction of RPA135. Four independent transformants were analyzed. Values shown are the averages obtained for the four cultures (standard deviations are shown). Control cultures, which did not receive RPA135, were also analyzed in parallel (standard deviations are not shown).
Figure 5
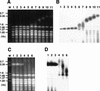
RFB activity analyzed by two-dimensional gel analysis. DNA was prepared from strains indicated, digested with _Bgl_II and _Sph_I and subjected to two-dimensional agarose gel electrophoresis followed by Southern hybridization with a rDNA probe (the _Hin_dIII–_Sph_I fragment; see Fig. 1). Spots indicated by arrows show accumulation of Y-form DNA molecules at RFB site.
Figure 6
Analysis of the size of chromosome XII by pulse-field gel electrophoresis. (A,B). Experiments similar to those in Figs. 2 and 4 were carried out. The RPA135 gene was introduced into NOY408-1a (lanes 8–10) and NOY408-1af (lanes 3–5). Control cultures without RPA135 introduction, NOY408-1a (lanes 6,7) and NOY408-1af (lanes 1,2) were also grown in parallel. Samples were taken at 44 (lanes 1,3,6,8), 80 (lanes 4,9), and 116 generations (lanes 2,5,7,10) after introduction of RPA135. Wild-type control culture, NOY408, was also analyzed (lane 11). (C,D). Haploid strains NOY408-2af (rpa135 fob1) (lanes 1–3) and NOY408-2a (rpa135) (lanes 4–6) were freshly constructed by tetrad dissection of diploid strains NOY408-f and NOY408, respectively. Samples were taken at 44 (lanes 1,4), 80 (lanes 2,5), and 116 generations (lanes 3,6) after germination of spores, and the size of chromosome XII was analyzed by pulse-field gel electrophoresis. (A,C) Chromosome patterns revealed by staining with ethidium bromide. (B,D) Autoradiographs obtained after hybridization with an rDNA probe. Size markers (lane M) are Hansenula wingei chromosomes (Bio-Rad, Hercules, CA). The background cross hybridization of the rDNA probe to the chromosomes without rDNA is not uniform in panels B and D. The reason is not clear, but its pattern was not reproducible. It should also be noted that heterogeneous chromosome XII (in B, lanes 6 and 7, and D lanes 4–6) wih specific hybridization overlapped chromosome IV with nonspecific hybridization, resulting in an artifactual strong band of ∼1.7 Mb in B, lane 7, and D, lanes 4–6.
Figure 7
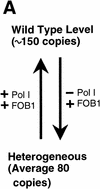
(A) Summary of the rDNA expansion/contraction system studied, and (B) fork block-dependent recombination model for rDNA expansion/contraction. The position of ARS and RFB are shown as filled oval and  , respectively. Individual lines represent chromatids with double-stranded DNA except for _c_′, where individual single-stranded DNAs are displayed to show detail of the structure formed after strand invasion. In this model, DNA replication starts from one of the ARS (ori-2) bidirectionally (a). In the yeast rDNA repeats, about one in three ARS sites is used as an active origin (Brewer and Fangman 1988; Linskens and Huberman 1988). A leftward replication fork is arrested at the RFB site and an exposed single-stranded region is cleaved by a nuclease (indicated by an open arrowhead in b). A recombination enzyme activates the resulting DSB end, and a strand invasion at a homologous duplex (a downstream sister chromatid near ori-3 in this example) takes place (c and _c_‘). A new replication fork is formed as a result of resolution of the Holliday junction (arrowheads). The new replication fork meets with the rightward replication fork from upstream, resulting in formation of two sister chromatids, one of which gains an extra copy of rDNA, indicated as boxed 35S-2 (d).
, respectively. Individual lines represent chromatids with double-stranded DNA except for _c_′, where individual single-stranded DNAs are displayed to show detail of the structure formed after strand invasion. In this model, DNA replication starts from one of the ARS (ori-2) bidirectionally (a). In the yeast rDNA repeats, about one in three ARS sites is used as an active origin (Brewer and Fangman 1988; Linskens and Huberman 1988). A leftward replication fork is arrested at the RFB site and an exposed single-stranded region is cleaved by a nuclease (indicated by an open arrowhead in b). A recombination enzyme activates the resulting DSB end, and a strand invasion at a homologous duplex (a downstream sister chromatid near ori-3 in this example) takes place (c and _c_‘). A new replication fork is formed as a result of resolution of the Holliday junction (arrowheads). The new replication fork meets with the rightward replication fork from upstream, resulting in formation of two sister chromatids, one of which gains an extra copy of rDNA, indicated as boxed 35S-2 (d).
Figure 7
(A) Summary of the rDNA expansion/contraction system studied, and (B) fork block-dependent recombination model for rDNA expansion/contraction. The position of ARS and RFB are shown as filled oval and  , respectively. Individual lines represent chromatids with double-stranded DNA except for _c_′, where individual single-stranded DNAs are displayed to show detail of the structure formed after strand invasion. In this model, DNA replication starts from one of the ARS (ori-2) bidirectionally (a). In the yeast rDNA repeats, about one in three ARS sites is used as an active origin (Brewer and Fangman 1988; Linskens and Huberman 1988). A leftward replication fork is arrested at the RFB site and an exposed single-stranded region is cleaved by a nuclease (indicated by an open arrowhead in b). A recombination enzyme activates the resulting DSB end, and a strand invasion at a homologous duplex (a downstream sister chromatid near ori-3 in this example) takes place (c and _c_‘). A new replication fork is formed as a result of resolution of the Holliday junction (arrowheads). The new replication fork meets with the rightward replication fork from upstream, resulting in formation of two sister chromatids, one of which gains an extra copy of rDNA, indicated as boxed 35S-2 (d).
, respectively. Individual lines represent chromatids with double-stranded DNA except for _c_′, where individual single-stranded DNAs are displayed to show detail of the structure formed after strand invasion. In this model, DNA replication starts from one of the ARS (ori-2) bidirectionally (a). In the yeast rDNA repeats, about one in three ARS sites is used as an active origin (Brewer and Fangman 1988; Linskens and Huberman 1988). A leftward replication fork is arrested at the RFB site and an exposed single-stranded region is cleaved by a nuclease (indicated by an open arrowhead in b). A recombination enzyme activates the resulting DSB end, and a strand invasion at a homologous duplex (a downstream sister chromatid near ori-3 in this example) takes place (c and _c_‘). A new replication fork is formed as a result of resolution of the Holliday junction (arrowheads). The new replication fork meets with the rightward replication fork from upstream, resulting in formation of two sister chromatids, one of which gains an extra copy of rDNA, indicated as boxed 35S-2 (d).
Similar articles
- Identification of DNA cis elements essential for expansion of ribosomal DNA repeats in Saccharomyces cerevisiae.
Kobayashi T, Nomura M, Horiuchi T. Kobayashi T, et al. Mol Cell Biol. 2001 Jan;21(1):136-47. doi: 10.1128/MCB.21.1.136-147.2001. Mol Cell Biol. 2001. PMID: 11113188 Free PMC article. - Transcription factor UAF, expansion and contraction of ribosomal DNA (rDNA) repeats, and RNA polymerase switch in transcription of yeast rDNA.
Oakes M, Siddiqi I, Vu L, Aris J, Nomura M. Oakes M, et al. Mol Cell Biol. 1999 Dec;19(12):8559-69. doi: 10.1128/MCB.19.12.8559. Mol Cell Biol. 1999. PMID: 10567580 Free PMC article. - The replication fork barrier site forms a unique structure with Fob1p and inhibits the replication fork.
Kobayashi T. Kobayashi T. Mol Cell Biol. 2003 Dec;23(24):9178-88. doi: 10.1128/MCB.23.24.9178-9188.2003. Mol Cell Biol. 2003. PMID: 14645529 Free PMC article. - [Gene amplification induced by the replication fork barrier site in yeast].
Kobayashi T, Takeuchi Y, Johzuka K, Horiuchi T. Kobayashi T, et al. Tanpakushitsu Kakusan Koso. 2001 Jun;46(8 Suppl):1004-12. Tanpakushitsu Kakusan Koso. 2001. PMID: 11436287 Review. Japanese. No abstract available. - [Recombination regulation by noncoding transcription in yeast rDNA repeats].
Kobayashi T. Kobayashi T. Tanpakushitsu Kakusan Koso. 2006 Nov;51(14 Suppl):2141-3. Tanpakushitsu Kakusan Koso. 2006. PMID: 17471925 Review. Japanese. No abstract available.
Cited by
- Regulation of ribosomal RNA production by RNA polymerase I: does elongation come first?
Albert B, Perez-Fernandez J, Léger-Silvestre I, Gadal O. Albert B, et al. Genet Res Int. 2012;2012:276948. doi: 10.1155/2012/276948. Epub 2012 Jan 12. Genet Res Int. 2012. PMID: 22567380 Free PMC article. - Expression of rRNA genes and nucleolus formation at ectopic chromosomal sites in the yeast Saccharomyces cerevisiae.
Oakes ML, Johzuka K, Vu L, Eliason K, Nomura M. Oakes ML, et al. Mol Cell Biol. 2006 Aug;26(16):6223-38. doi: 10.1128/MCB.02324-05. Mol Cell Biol. 2006. PMID: 16880531 Free PMC article. - The peculiar genetics of the ribosomal DNA blurs the boundaries of transgenerational epigenetic inheritance.
Bughio F, Maggert KA. Bughio F, et al. Chromosome Res. 2019 Mar;27(1-2):19-30. doi: 10.1007/s10577-018-9591-2. Epub 2018 Dec 4. Chromosome Res. 2019. PMID: 30511202 Free PMC article. Review. - Contrasting roles of checkpoint proteins as recombination modulators at Fob1-Ter complexes with or without fork arrest.
Mohanty BK, Bairwa NK, Bastia D. Mohanty BK, et al. Eukaryot Cell. 2009 Apr;8(4):487-95. doi: 10.1128/EC.00382-08. Epub 2009 Feb 20. Eukaryot Cell. 2009. PMID: 19234097 Free PMC article. - How do cells count multi-copy genes?: "Musical Chair" model for preserving the number of rDNA copies.
Iida T, Kobayashi T. Iida T, et al. Curr Genet. 2019 Aug;65(4):883-885. doi: 10.1007/s00294-019-00956-0. Epub 2019 Mar 23. Curr Genet. 2019. PMID: 30904990 Review.
References
- Bastia D, Mohanty BK. Mechanisms for completing DNA replication. In: DePamphilis ML, editor. DNA replication in eukaryotic cells. Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press; 1996. pp. 177–215.
- Brewer BJ, Fangman WL. The localization of replication origins on ARS plasmids in S. cerevisiae. Cell. 1987;51:463–471. - PubMed
- ————— A replication fork barrier at the 3′ end of yeast ribosomal RNA genes. Cell. 1988;55:637–643. - PubMed
- Brewer BJ, Lockshon D, Fangman WL. The arrest of replication forks in the rDNA of yeast occurs independently of transcription. Cell. 1992;71:267–276. - PubMed
- Brown DD, Dawid IB. Specific gene amplification in oocytes. Oocyte nuclei contain extrachromosomal replicas of the genes for ribosomal RNA. Science. 1968;160:272–280. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases