Increased Frequency of De Novo Copy Number Variations in Congenital Heart Disease by Integrative Analysis of SNP Array and Exome Sequence Data (original) (raw)
. Author manuscript; available in PMC: 2015 Oct 24.
Abstract
Rationale
Congenital heart disease (CHD) is among the most common birth defects. Most cases are of unknown etiology.
Objective
To determine the contribution of de novo copy number variants (CNVs) in the etiology of sporadic CHD.
Methods and Results
We studied 538 CHD trios using genome-wide dense single nucleotide polymorphism (SNP) arrays and/or whole exome sequencing (WES). Results were experimentally validated using digital droplet PCR. We compared validated CNVs in CHD cases to CNVs in 1,301 healthy control trios. The two complementary high-resolution technologies identified 63 validated de novo CNVs in 51 CHD cases. A significant increase in CNV burden was observed when comparing CHD trios with healthy trios, using either SNP array (_p_=7x10−5, Odds Ratio (OR)=4.6) or WES data (_p_=6x10−4, OR=3.5) and remained after removing 16% of de novo CNV loci previously reported as pathogenic (_p_=0.02, OR=2.7). We observed recurrent de novo CNVs on 15q11.2 encompassing CYFIP1, NIPA1, and NIPA2 and single de novo CNVs encompassing DUSP1, JUN, JUP, MED15, MED9, PTPRE SREBF1, TOP2A, and ZEB2, genes that interact with established CHD proteins NKX2-5 and GATA4. Integrating de novo variants in WES and CNV data suggests that ETS1 is the pathogenic gene altered by 11q24.2-q25 deletions in Jacobsen syndrome and that CTBP2 is the pathogenic gene in 10q sub-telomeric deletions.
Conclusions
We demonstrate a significantly increased frequency of rare de novo CNVs in CHD patients compared with healthy controls and suggest several novel genetic loci for CHD.
Keywords: De novo copy number variation, congenital heart disease, SNP-array, whole exome sequencing, CNV burden, congenital cardiac defect, microarray, genomics
INTRODUCTION
Congenital heart disease (CHD) is the most frequent birth defect, affecting approximately 7 in 1000 live births,1 and is a significant cause of childhood morbidity and mortality.2 Rare Mendelian disorders, specific chromosomal abnormalities, and copy number variants (CNVs) are known to explain a subset of CHD cases,2-4 but the cause of over 80% of CHD remains unexplained.5-12
The application of evolving technologies that detect structural variation throughout the genome has demonstrated a considerable contribution of CNVs to CHD. Early cytogenetic studies recognized an increased prevalence of de novo chromosomal abnormalities in syndromic CHD patients, observations that were replicated and extended to non-syndromic CHD with successive generations of CNV detection technologies including array CGH and low density SNP arrays. Using these techniques, researchers have demonstrated significant burden of large de novo CNV in some specific CHD lesions. Such CNVs are reported to occur in 13.9% of infants with single ventricles compared to 4.4% in controls,13 in 10% of non-syndromic tetralogy of Fallot (TOF) compared to 4% of controls,5 and in 12.7% children with hypoplastic left heart syndrome compared to 2% of controls.20 Among different CHD lesions, the frequency of large de novo CNVs is similar.20 While many large CNVs are unique to a single CHD patient, several are recurrent in CHD cohorts. A 3-Mb 22q11.2 deletion is the most common recurrent de novo CNV associated with syndromic conotruncal defects (CTDs) and is found overall in at least 10% of TOF, 35% of truncus and 50% of interrupted aortic arch (IAA) type B cases.23 Recurrent de novo CNVs in CHD patients reported in multiple studies also occur at chromosomes 1q21.1, 3p25.1, 7q11.13, 8p23.1, 11q24-25, and 16p13.11.
The identification of CHD loci that are altered by CNVs provides opportunities to elucidate disease pathogenesis. However, discerning the causal gene(s) and inferring critical networks and pathways that cause or contribute to CHD has been difficult because low-resolution technologies used in many studies (array CGH and low-density SNP arrays) typically define large CNVs (>100kb) involving many genes. To address these issues, we capitalized on two independent strategies, high-density SNP genotyping arrays (Illumina Omni-1.0 and 2.5M) and whole exome sequencing (WES), to detect smaller de novo CNVs in a family-based trio study of sporadic CHD cases with conotruncal, heterotaxy, and left ventricular outflow tract defects.24 We compared CNVs found in CHD trios to those identified in healthy control trios. Through these analyses we sought to compare the robustness of genome-wide CNV detection using array-based and sequence-based technologies to determine if there was an increased burden of smaller de novo CNVs in CHD patients as was demonstrated with larger CNVs, and to determine if fewer genes altered by these CNVs enabled more precise detection of gene networks and pathways contributing to the pathogenesis of CHD.
METHODS
Ethics statement
The protocol was approved by the Institutional Review Boards of Boston Children's Hospital, Brigham and Women's Hospital, Great Ormond St. Hospital, Children's Hospital of Los Angeles, Children's Hospital of Philadelphia, Columbia University Medical Center, Icahn School of Medicine and Mt. Sinai, Rochester School of Medicine and Dentistry, Steven and Alexandra Cohen Children's Medical Center of New York, and Yale School of Medicine. Written informed consent was obtained from each participating subject or their parent/guardian.
Patient cohorts
CHD probands and parents were recruited into the CHD Genes Study of the Pediatric Cardiac Genomics Consortium (CHD genes: ClinicalTrials.gov identifier NCT01196182) as previously described,24 using protocols approved by Institutional Review Boards of each institution. Trios selected for this study had no history of CHD in first-degree relatives. CHD diagnoses were obtained from echocardiograms, catheterization and operative reports; extra-cardiac findings were extracted from medical records and included dysmorphic features, major anomalies, non-cardiac medical problems, and deficiencies in growth or developmental delay. The etiologies for CHD were unknown; patients with previously identified cytogenetic anomalies or pathogenic CNVs identified through routine clinical evaluation were excluded. Whole blood samples were collected and genomic DNA extracted.
CHD trios were studied by SNP arrays (n=414) or by WES (n=356), including a subset (n=233) that were analyzed by both methods. The distribution by CHD lesions in patients genotyped by arrays was: 403 (61%) left ventricular obstruction (LVO); 197 (30%) conotruncal defects (CTD); 49 (7%) heterotaxy (HTX); and 12 (2%) other cardiac diagnoses (Supplementary Table I). The distribution by CHD lesions in patients studied by WES was 284 (46.1%) left ventricular obstruction (LVO); 235 (38.1%) conotruncal defects (CTD); 78 (12.7%) heterotaxy (HTX); and 19 (3.1%) with other cardiac diagnoses (Supplemental Table II).
Control trios were the unaffected sibling and parents of a child with autism who were consented and recruited through the Simons Simplex Collection (SSC). CNVs were identified in the same way in the control trios as in the cases using SNP arrays (n=814) or WES (n=872), including a subset (n=385) analyzed by both methods.
Additional data on the distribution and prevalence of previously reported CNVs in the general population was derived from the Database of Genomic Variants (http://dgv.tcag.ca) and from 649 de-identified control subjects who had participated in an unrelated psychiatric case-control study, genotyped on the same high density SNP array platforms at the same genotyping center as the CHD probands (438 on the Illumina Omni-1M and 211 on the Illumina Omni-2.5M). These controls were used only to prioritize the de novo CNVs identified by SNP array methods that were selected for confirmation analyses.
Array genotyping and CNV identification
A total of 360 CHD parental samples genotyped on the Omni1M and 654 on Omni2.5M arrays were applied for cluster definition using Illumina Genome Studio clustering algorithm. Raw data is publicly available through the database of genotypes and phenotypes (dbGaP) National Heart, Lung, and Blood Institute (NHLBI) Bench to Bassinet Program: The Pediatric Cardiac Genetics Consortium (PCGC) under dbGaP Study Accession: phs000571.v1.p1.We removed clusters with outlier values of SNP call rate, Hardy-Weinberg equilibrium, AA/AB/BB cluster means, and minor allele frequency to improve the intensity noise (Log R ratio standard deviation) from a mean of 0.2 (using the default cluster file from Illumina) to 0.1 for CHD samples. Briefly, individual samples were filtered through a standard quality control pipeline.25 B-allele frequency (BAF) and Log R ratio (LRR) values were exported from Illumina Genome Studio. Only samples with SNP call rate > 98%, standard deviation (SD) of normalized intensity (LRR) < 0.3, absolute value of GC-corrected LRR <0.005, as well as CNV call count <800 for Omni1-Quadv1 or <300 for Omni2.5-8v1 were included.63 Samples with high inbreeding coefficients, that were duplicated, or had gender mismatches, and trios with Mendelian errors > 1% were removed from analyses. We started with 1,536 genotyped samples (512 trios), including 561 on the Illumina Omni-1M and 969 on the Illumina Omni-2.5M. Four hundred and sixty-one trios had the same array version for all family members. Upon completion of these QC procedures 1,245 samples, including 447 genotyped on the Illumina Omni-1M and 798 on the Illumina Omni-2.5M high-density SNP array platforms, were taken forward for analysis, constituting 415 complete trios (Supplemental Table III).
Three groups (CHOP, Harvard, Yale) independently analyzed genotyping data using slightly different algorithms to detect putative de novo CNVs. For each of the three independent analyses, CNVs were called for each subject using PennCNV 64 with the hidden Markov model algorithm and custom-made population frequency of B-allele (PFB) and GC model files. CNVs were called when 10 or more consecutive probes demonstrated consistent copy number change. The PennCNV detect_cnv --trio option was used to boost transmission probability of CNV calling for initially de novo scored CNVs. Fragmented CNV calls were merged using clean_cnv. All candidate CNVs were visually inspected to ensure the appropriate pattern of LRR and B-allele frequency was consistent with the CNV call. Additionally, Gnosis,25 QuantiSNP,65 and Nexus (biodiscovery.com) were used to increase specificity. De novo CNVs were prioritized for quality by genomic length, number of probes, confidence score based on signal strength, 50% overlap of two or more algorithms, low parental origin p-value using infer_snp_allele, and visual BAF/LRR review. CNVs with a minor allele frequency > 1% were removed, leaving rare CNVs. All putative de novo CNVs were experimentally evaluated by digital droplet PCR (ddPCR, Supplemental Figure I), and only validated CNVs are reported.
De novo CNV loci that were previously reported as pathogenic were defined by reported recurrence in at least two publications using independent data. Although some of the CNVs reported here overlap with previously reported CNVs in CHD patients based on review of the literature,66, they do not meet our frequency constraint for previously reported pathogenic de novo CNV loci.
CNV identification and variant calling from WES data
WES data from 356 CHD trios were analyzed for de novo CNVs (Supplemental Table II). WES samples were captured with the Nimblegen SeqCap Exome V2 chemistry and sequenced on the Illumina HiSeq 2000 platform as previously described.26 Sequence reads were aligned to the human reference genome hg19 using Novoalign (http://novocraft.com), BWA,67 and ELAND.68 Duplicates were marked with Picard (http://picard.sourceforge.net). Indel realignment and Base Quality Score Recalibration was done with GATK. XHMM is an algorithm to detect exon-level copy number variation and assign CNV quality metrics38 and was used at four of the PCGC analysis sites (CHOP, Harvard, Columbia and Mount Sinai) to identify de novo CNVs (Supplemental Figure II). Candidate de novo CNVs were inspected visually. Putative de novo CNVs were prioritized for confirmation based on genomic length, low sequence depth variability and low prevalence in the XHMM call set data (AF<1%). All putative de novo CNVs were independently confirmed by ddPCR.
SNVs and short insertions/deletions (indels) were called from the Novoalign alignment of WES trios using a pipeline derived from GATK version 2.7 best practices.69 Briefly, aligned reads were first compressed using the GATK ReducedReads module and variants were called on all CHD WES trios using the UnifiedGenotyper joint variant calling module. Identified variants were filtered using GATK variant quality score recalibration. Variants were annotated using SnpEff.70 De novo SNVs and indels were independently confirmed using Sanger sequencing.
CNV confirmation with digital droplet PCR
Putative CNVs were experimentally confirmed with ddPCR as previously reported71 using an 18-27 base pair FAM probe designed within each candidate CNV region, avoiding homopolymer runs or probes that began with G. A VIC probe targeting the RPP30 gene was used as reference. Reaction mixtures of 20 μL volume comprising ddPCR Master Mix (Bio-Rad), relevant forward and reverse primers and probe(s) and 100 ng of digested DNA were prepared, ensuring that approximately 25-75% of the 10,000 droplets ultimately produced were positive for FAM or VIC signal. For de novo CNV confirmations, DNA from the CHD patient and parents was used. After thermal cycling, plates were transferred to a droplet reader (Bio-Rad) that flows droplets single-file past a two-color fluorescence detector. Differentiation between droplets that contain target and those that did not was achieved by applying a global fluorescence amplitude threshold in QuantaSoft (Bio-Rad). The threshold was set manually based on visual inspection at approximately the midpoint between the average fluorescence amplitude of positives and negative droplet clusters on each of the FAM and VIC channels. Confirmed CNV duplications had approximately 50% increase in the ratio of positive to negative droplets as did the reference channel. Conversely confirmed CNV deletions had approximately half the ratio of positive to negative droplets as did the reference channel.
Network analysis
Three bioinformatic algorithms were utilized: DAVID,35 DAPPLE,36 and WebGestalt.37 Four different gene lists derived from the de novo CNV loci were used (Supplemental Table IV). The lists were constructed as follows: (1) All genes contained within de novo CNV intervals; (2) Published “causative” genes from previously reported CHD CNVs intervals in addition to all genes in novel CHD CNV intervals. “Causal” genes in previously reported CNV intervals included ELN (Williams syndrome), RAI1 (Smith-Magenis syndrome),TBX1 (22q11 deletion), GATA4 (8p23.1 deletion), GJA5 (1q21.1 duplication), and NKX2.5 (5q35.1 deletion); (3) Genes contained solely within novel CHD CNV intervals (e.g., exclude genes from previously published CNVs); (4) Genes contained within de novo CNV intervals that are highly expressed in the developing mouse heart (top quartile of all genes expressed E14.5 mouse heart).26 We anticipated that genes in list 2 and list 4 would have increased specificity for CHD in comparison to genes in list 1 and that genes in list 3 would be biased towards new disease networks.
We expanded network analysis input gene lists by including both de novo CNV genes and de novo single nucleotide variants (SNV) that were previously identified in CHD probands by WES.26 Only de novo SNVs predicted to be deleterious (e.g., loss of function (LOF): nonsense, frame-shift, and splice site mutations and missense variants that alter highly conserved amino acid residues or predicted to be deleterious by SIFT or PolyPhen2) were included in the expanded gene list. The additional gene lists included: (5) All genes within a de novo CNV interval (e.g., list 1) and protein-altering SNVs and (6) Published “causative” genes from previously reported CHD CNVs intervals in addition to all genes in novel CHD CNV intervals (e.g., list 2) and protein altering SNVs.
Statistical analysis
Burden calculations were done with a Fisher exact test computed in the R statistical computing environment. For analyses using DAVID, networks with an enrichment of genes impacted by CNVs were assigned a _p_-value with Benjamini and Hochberg correction for multiple testing with a false discovery rate of 0.05. In DAPPLE, type I error was controlled through permutation. _p_-values of less than 0.05 were considered significant.
RESULTS
Identification of De Novo CNVs
We studied 415 CHD trios genotyped by SNP arrays and 356 trios by WES analysis, including 233 trios studied by both methods. No trios had an affected first-degree relative and the genetic cause of CHD in all studied children was unknown (Supplementary Tables 1 and 2).
Sixty-five de novo CNVs identified in CHD cases were independently confirmed by ddPCR. De novo CNVs were identified in 51 unique probands (9.8%). These CNVs ranged in size from 0.1 kb to 12.8 Mb. 48 of these (76%) were <500 kb and half were smaller than 110 kb. The number of genes in the CNV intervals ranged from 1 to 175 with 42 (67%) having ≤ 5 genes. Four de novo intervals contained no genes. Six probands had two de novo CNVs, two had three CNVs and one had four CNVs.
The parental origin of deletion CNVs was determined when the haplotype of the remaining copy could be uniquely assigned to one parent. Seven de novo CNVs arose on maternal chromosomes and 10 on paternal chromosomes. The remainder could not be assigned due to uninformative or insufficient numbers of informative parent-of-origin SNPs.
Comparison of SNP Array and WES CNV calling
To consider the accuracy of identifying de novo CNVs from SNP array data, we first considered a set of 40 high-confidence PennCNV de novo CNV calls that contained ≥10 adjacent SNPs, were >10 kb in length, and passed visual inspection. Among these 40 high-confidence putative CNVs, 40 were experimentally validated by ddPCR in the proband and 32 (80%) were experimentally confirmed to be de novo, representing a false positive rate of 20%. For smaller de novo CNVs identified using the high-density array data, we considered a set of 97 high-confidence PennCNV putative de novo CNV calls based on 7-9 SNPs. While 88% were experimentally validated by ddPCR in the proband, only four of the 97 (5%) were confirmed to be de novo.
From the WES data, we selected an initial set of 29 putative CNVs with a size range spanning six orders of magnitude from 530 bases in length (two exons) to more than 8 Mb in length covering hundreds of exons. Twenty-six of the 29 CNVs (90%) confirmed experimentally to be de novo, representing a false positive rate of 10%. The three false positive CNVs included one 530-bp region that contained only two exon targets and two different inherited CNVs that were miscalled as de novo because both parents harbored CNVs at the locus. Based on these considerations, we restricted subsequent WES de novo CNV calls to those containing ≥3 exons and for which each parental dataset contained no CNVs within the locus.
To evaluate false negative rates of the two platforms and analyses, we tested our ability to detect four CNVs (two 22q11 deletions, one 17p11 duplication, and one 10q terminal deletion; Supplemental Table V in clinical cases previously diagnosed with these CNVs. These four CNVs served as positive controls and were distinct from the PCGC cohort. Both the SNP array and WES platforms detected each of these four large, clinically significant CNVs.
We also compared the results of de novo CNVs analysis from the 233 trios studied by both SNP array and WES. Among 42 confirmed de novo CNVs in these trios, 24% (10/42) were identified by both platforms while 40% (17/42) were identified only with the SNP arrays and 35% (15/42) only by WES (Figure 1). The recognized technical limitations of each platform prevented detection of some CNVs. For example, CNVs that occur exclusively in noncoding sequences are not captured by WES whilst CNVs in coding or non-coding genomic regions where the SNP density is sparse can escape detection by SNP arrays.
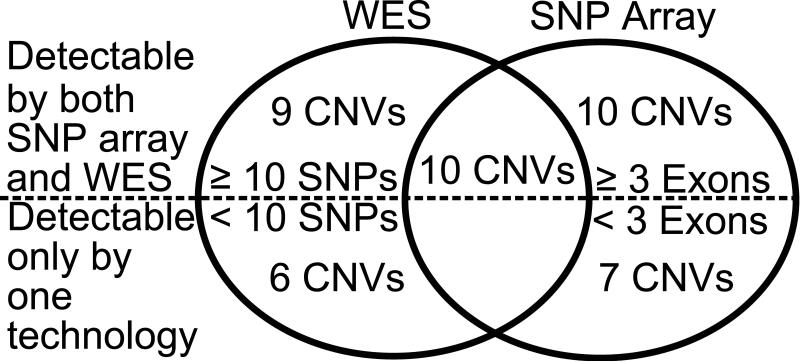
Figure 1.
Comparison of SNP array and WES platforms in detection of the 42 validated de novo CNVs in the subset of 233 probands studied by both technologies. 10 of the 42 were detected by both methods, 32 were called by one method. The figures below the dotted line show the number of CNVs that were below the detection limits of the second platform (CNVs that span <10 SNPs on SNP arrays or <3 exons on WES) and hence could not have been called. The figures above the dotted line show the number of CNVs with sufficient SNPs and/or exons to enable high confirmation rates, but that were not called.
From our studies we deduced that de novo CNVs were accurately detected by arrays when ≥10 adjacent SNPs were impacted or by WES when greater than three adjacent exons were impacted. In our dataset, 29 of 42 CNVs fulfilled both of these criteria and should have been identified by both technologies (Figure 1). However, only 34% (10/29) of these ddPCR confirmed CNVs were identified by both platforms. SNP arrays uniquely identified 34% (10/29) and WES analyses uniquely identified 31% (9/29). Taken together, the false negative rate of each methodology is approximately 30-35%. Overall, the genome-wide analyses of de novo CNVs identified by SNP arrays was reasonably concordant with WES data, but each also identified complementary CNVs. The minimum CNV size that we reliably detected by SNP arrays was 10 kb and by WES was 1 kb, although some smaller CNVs identified by these techniques were validated.
CNV burden analysis
The burden of de novo CNVs in CHD cases and control trios was initially compared using analyses from SNP arrays. De novo CNVs were assessed in 841 control trios, studied using the Illumina Omni1M array to match the case trio array resolution and called using the PennCNV algorithm using computational parameters described previously25. Nine de novo ddPCR-validated CNVs were identified among 841 control trios. Twenty-two de novo ddPCR-validated CNVs were identified among 462 CHD trios with SNP arrays. These data define a significant burden of CNVs in CHD cases compared to controls (OR: 4.6, Fisher p-value: 7 x 10−5; Table 2). After excluding nine previously identified CHD-associated CNVs, the calculated burden of novel CNVs identified in CHD cases remained modestly significant (OR: 2.7, Fisher _p_=0.02).
Table 2.
Case Control de novo CNV Burden
| N Probands | N (%) CNVs | OR | P-value | ||
|---|---|---|---|---|---|
| SNP Array | SSC1 | 841 | 9 (1%) | - | - |
| PCGC: all CNVs | 462 | 22 (4.7%) | 4.6 | 7 × 10−5 | |
| PCGC: novel loci | 13 (2.8%) | 2.7 | 0.02 | ||
| WES | SSC2 | 872 | 14 (1.6%) | - | - |
| PCGC: all CNVs | 356 | 19 (5.6%) | 3.5 | 6 × 10−4 | |
| PCGC: novel loci | 13 (3.9%) | 2.3 | 0.03 |
To provide further support for this finding, we analyzed the burden of de novo CNVs that were identified by WES. WES in CHD cases and control trios were technically comparable, including the same Nimblegen V2 exome capture chemistry and similar sequence read depths obtained on identical Illumina platforms. Sixty percent of control trios were sequenced at the same site (Yale Center for Genome Analysis) that sequenced the cases. Raw sequence reads were processed through the identical short read aligner (Novoalign) for CNV burden analysis. SNP genotyping of CHD and control datasets and principal component analysis did not identify any systematic biases (Supplemental Figure V). Cases and controls were matched for gender as best as possible with slight excess of male cases. Using an identical XHMM pipeline (CNVs involving ≥3 exons and no parental CNVs within 1 MB), we identified 19 de novo ddPCR-validated CNVs in 356 CHD trios, and 14 de novo ddPCR-validated CNVs in 872 control trios (OR: 3.5, Fisher _p_=6 x 10−4; Table 2). Excluding the six de novo CNVs previously identified as CHD-associated, we identified a similar OR and _p_-value as in the SNP array data (OR: 2.3, Fisher _p_=0.03).
Our data identify an increased burden of CNVs, detected by SNP arrays or WES, in CHD patients compared to controls. We observed a larger mean size of de novo CNVs with increased burden in CHD patients (3.6 Mb) than controls (495 kb; t-test _p_=0.035) with the distribution of CHD CNVs skewed towards the largest CNVs identified in CHD cases. The median size of de novo CNVs from CHD cases (522 kb) was also significantly larger than controls (118 kb; Mann-Whitney _p_=0.028). Of the CNVs identified by SNP array which were capable of detecting CNVs outside of coding regions, there was a trend towards an increased number of de novo CNVs in controls that contained no coding exon (4/9) compared to PCGC cases (3/22; Fisher _p_= 0.15).
Putative CHD loci at 15q11.2 and 2p13.3
Overlapping de novo CNVs found in multiple cases and not in controls likely contain disease genes. Sixteen of 65 (25%) de novo CNVs in CHD probands have been previously implicated in CHD5, including four 22q11.2 deletions, three 8p23 deletions (involving GATA4), two 1q21.1 duplications (involving PRKAB2, PDIA3P, FMO5, CHD1L, BCL9, ACP6 and GJA5), one 22q11.2 distal duplication, one 2q22.3 deletion (that causes Mowat-Wilson syndrome), one 11q24.2-q25 deletion (that causes Jacobsen syndrome) and four with CNVs in 15q11.2.
CNVs in four CHD probands (two deletions, two duplications) at the BP1-BP2 15q11.2 locus that spans approximately 225 kb (chr15:22,836,000-23,062,000) were observed as recurrent de novo events (Supplemental Figures III and IV). Both patients with duplications (1-00192, 1-00315) and one with a deletion (1-00243) had LVO due to aortic coarctation. The remaining proband (1-01396) had TOF with pulmonary atresia. As there was no de novo CNV identified in this region among 814 and 872 control trios studied respectively by SNP arrays or WES, this locus has a significant burden of de novo CNVs in CHD cases (4/538 CHD vs. 0/1301 controls; Fisher _p_=0.007). CNVs at the 15q11.2 locus were observed at low frequency (AF<1%) in the Database for Genomic Variants (DGV). Among the three genes altered by this CNV (CYFIP1, NIPA1, and NIPA2), only CYFIP1 is highly expressed in the developing mouse heart.26 CYFIP1 encodes the cytoplasmic FMR1-interacting protein 1, which has dual roles in inhibiting local protein synthesis and in promoting actin remodeling.27 An earlier study observed an increased burden of inherited deletions in CHD cases at 15q11.21 and a recent paper identified a single proband with a 6-Mb de novo duplication at 15q11.2-q13.120 and two additional cases with inherited 300-400-kb duplications at 15q11.2. Our data provide additional evidence that de novo CNVs at 15q11.2 may contribute to disease risk in CHD.
In addition, a recurrent CNV was observed to alter a novel locus at chromosome 2p13.3. A de novo 190-kb deletion was identified in a TOF proband (1-01536) and was maternally inherited in a proband with truncus arteriosus (1-01805). No 2p13.3 CNV was found in control samples or in DGV. Among three genes included in the CNV interval (ASPRV1, PCBP1 and PCBP1-AS1), only PCBP1 is highly expressed in the developing mouse heart.26 PCBP1 encodes a major cellular poly(rC)-binding protein, which controls translation from mRNAs containing the DICE (differentiation control element).28 In DECIPHER, patient 257771 with an atrioventricular canal defect had a 7-Mb overlapping deletion of 2p13.3, suggesting this locus may also contribute to disease risk in CHD.
Integration of CNV and sequence data to identify CHD genes
To improve the identification of specific genes altered by CNVs that might cause or contribute to CHD, we searched the WES data for de novo, rare loss-of-function (LOF) variants in genes encoded in CNV intervals. We identified a terminal deletion of chromosome 11q24.2-q25, which causes Jacobsen syndrome in one CHD patient (1-01486) with clinical manifestations typical of this dominant disorder (hypoplastic left heart, coarctation of the aorta, mitral and aortic valve atresia, strabismus, and short stature). ETS1 has been proposed as the critical CHD gene in the Jacobsen syndrome locus based on impaired ventricular development in an _Ets1_-null mouse.29 WES analyses identified a de novo ETS1 frameshift mutation (chr11:128350159GTCCT>G, c.1046_1049delAGGA, [p.K349fs]) in another CHD patient without the chromosome 11q24.2-q25 deletion with cardiac abnormalities observed in Jacobsen syndrome (hypoplastic left heart and mitral valve atresia). Our data provide the first human genetic evidence to suggest that ETS1 mutations contribute to the cause of cardiac malformations in Jacobsen syndrome.
We also assessed whether de novo CNVs in combination with a rare or novel deleterious variant on the other allele might produce recessive forms of CHD. One CHD patient (1-01179) with a de novo 10q25-26 deletion also had a novel CTBP2 variant (p.R134W) on the remaining allele. The hemizygous variant was absent from public genome databases, is predicted to be damaging (Polyphen2 score of 0.998), and altered a phylogenetically conserved residue (PhyloP score = 2.54). Cardiac abnormalities are present in approximately one third of patients with subterminal chromosome 10q deletions and recently CTBP2 was proposed as a candidate CHD gene.32 The clinical manifestations of our patient, truncus arteriosus and right aortic arch, resemble the phenotypes identified in a _Ctbp2_-null mouse (failure of vascular remodeling and cardiac looping).33 We suggest that CTBP2 sequence analyses in individuals with chromosome 10q deletions may identify additional variants in a subset of patients that modify phenotype.
Correlation of CHD phenotypes and CNVs
The frequency of de novo CNVs was 10% among conotruncal anomalies, 6% among left-sided obstructive lesions and 21% in heterotaxy. We observed a modest trend towards increased extra-cardiac manifestations such as developmental delay in patients with de novo CNVs (Supplemental Table VI). Approximately 31% of all CHD patients studied with SNP arrays or WES had extra-cardiac manifestations, whilst 40% (21/52; OR:1.5, Fisher _p_=0.2) of patients with de novo CNVs had extra-cardiac features. This association has been found in some,34 but not all,20 previous studies, perhaps due to differences in the ages of the CHD patients studied, methods of clinical data collection, and the definition of an extra-cardiac anomaly.
Gene networks impacted by CNVs in CHD
We employed pathway and network analysis with DAVID,35 DAPPLE,36 and WebGestalt,37 using as input four different lists of genes encoded within all de novo CNV loci (Methods and Supplemental table IV). Initial gene lists contained: (1) all genes encoded in a de novo CNV interval; (2) genes previously defined as causative with CNVs intervals plus all genes in novel de novo CNV intervals; (3) only genes contained within novel de novo CNV intervals; (4) all genes contained within de novo CNV intervals that are highly expressed (top 25%) in the developing heart.26
DAVID identified enrichment of a gene pathway implicated in acetylation p<2.3x10−4; phosphoprotein p<3.9x10−4, and G protein-activated inward rectifier potassium channel p<2.5x10−2 (Benjamini-Hochberg corrected). WebGestalt implicated an enrichment of previously identified CHD genes including ELN, NKX2.5, GATA4, and ZEB2 contributing to Gene Ontology processes: anatomical structure formation involved in morphogenesis p<0.03, cardioblast differentiation p<0.03, and septum secundum development p<0.02 (Benjamini-Hochberg corrected).
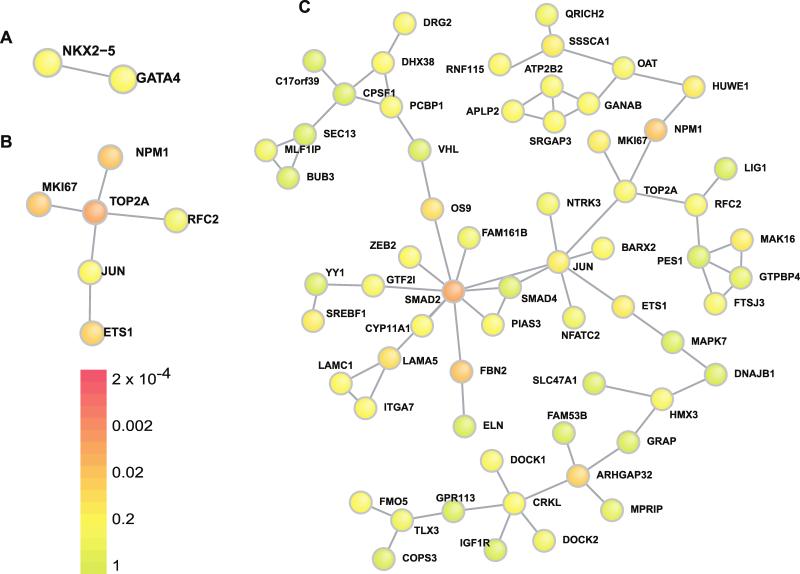
Using DAPPLE, we identified two additional sub-networks of direct protein/protein interactions that were consistently observed across four gene lists. Among genes encoded within CNVs that are highly expressed in the developing heart, a sub-network consisting of NKX2.5 and GATA4 (p<0.1, Figure 2a) and a sub-network consisting of ETS1, JUN, TOP2A, and MKI67 (p<0.01, Figure 2b) were identified. By further expanding the CNV gene lists to include genes with de novo LOF mutations, the ETS1/JUN/TOP2A sub-network was significantly elaborated upon and enriched (p<0.005). Each of these three genes was directly linked through protein-protein interactions to sub-networks containing ≥ 10 additional genes identified in either CNV or WES datasets.26 This entire network incorporated over 60 genes implicated in CHD (Figure 2c). As the ETS1/JUN/TOP2A sub-network was robust to the specific de novo CNV gene list (criteria 2 above) and expanded with the addition of genes containing rare de novo LOF mutations, the data suggest that this sub-network contains genes and pathways involved in CHD.
Figure 2.
Network analysis of CNV loci genes. Two networks of direct protein-protein interactions, (A) NKX2.5/Gata4 and (B) ETS1/JUN/TOP2A, were consistently identified in the DAPPLE de novo CNV loci analysis. P-values from the genes highly expressed in the developing heart, the most restrictive gene set list, are presented here. (C) The ETS1/JUN/TOP2A network was significantly elaborated upon by incorporating genes with deleterious de novo point mutations and indels in the WES exome sequencing analysis in addition to the CNV loci. Of note, two probands had de novo ETS1 variants (one CNV and one frameshift), two probands had de novo SMAD2 variants (a splice site mutation and a highly conserved missense variant) and two probands had de novo ELN variants (both Williams syndrome CNVs).
DISCUSSION
We report whole-genome CNV analyses using complementary detection technologies in a large cohort of CHD patients. CNV detection in WES has been investigated in schizophrenia38 and autism,39 but array-based and sequence-based strategies have not previously been directly compared, and our data highlight the differences between array-based and sequence-based strategies to detect de novo CNVs. By defining small CNVs with high resolution and integrating these findings with WES data that identified rare deleterious mutations, we identified novel de novo CNVs and genes involved in the pathogenesis of CHD. We show that 9.8% (53/538) of CHD patients without a previously identified genetic etiology have rare de novo CNVs. We previously demonstrated that 10% of CHD patients in our cohort have de novo single nucleotide or small insertion/deletion mutations in genes highly expressed in the developing heart that are likely to be damaging.26 None of the CHD patients with rare de novo CNVs reported here carry these variants. Even if all the de novo CNVs and de novo predicted pathogenic sequence variants we have identified were causative, we do not yet know the etiology for the majority of CHD subjects in our study.
Our detection rate of approximately 10% de novo CNVs in CHD patients is equivalent to previous studies, despite identifying small CNVs. Had we not excluded patients with known pathogenic CNVs identified through clinical care, we expect that de novo CNVs would have been identified in approximately 15% of CHD patients, based on the prevalence of common de novo CNVs in CHD (e.g., 7% of TOF with chromosome 22q11 deletions, and 1% of TOF to 1q21 CNVs). In our study, these CNV loci accounted for <1% of CHD probands
Despite these exclusion criteria, we identified a four-fold increased frequency of rare de novo CNVs relative to the background frequencies of 1.2% (detected by SNP arrays) and 1.8% (detected by WES) of de novo CNVs in controls (_p_=7 x 10−5, _p_=4 x10−4 respectively). Even after excluding previously defined CNVs, we still observed an approximate two-fold increase in novel rare de novo CNVs (_p_=0.02).
Since the odds ratio of de novo CNVs in cases vs controls was 3.5-4.6, we estimate that between 50-70% of de novo CNVs observed in cases may be disease causing. The possibility exists that a higher percentage of de novo CNVs increase the risk of CHD but may not be sufficient to cause CHD without other contributing genetic or environmental factors. Additionally, subtle anatomic defects in the heart may not have been diagnosed in the control group since controls were not systematically examined by echocardiogram. Overall, our evidence suggests a model in which de novo CNVs contribute to CHD
The comparison of dense array-based platforms and WES analyses to detect independently validated CNVs indicate that each strategy identifies only ~70% of the CNVs that should be within the detection limitations of each technology. As such, these two CNV methodologies provide substantial complementary information. An important corollary to this conclusion is that previously published CNV analyses in human disease may have significantly underestimated the burden conveyed by these structural variants.
Amongst all confirmed de novo CNVs, 61% (41) were deletions and 39% (26) were duplications. The proportion of these classes of CNVs are not significantly different; whether or not the trend toward more CNV deletions in CHD is biologically meaningful or reflects greater sensitivity to detect deletions by these methods will require further analyses. De novo CNVs ranged in size from less than 1 kb to 12.8 Mb, with a median size of 110 kb. Thus, half of the independently confirmed CNVs were smaller than the reported detection limit of most prior studies. For example, four CHD patients had 200-kb de novo CNVs on chromosome 15. While the pathogenicity of the identified CNVs remains to be determined, we propose that the smaller CNVs involving fewer genes are particularly valuable in defining specific candidate CHD genes in comparison to larger CNVs that typically include many more candidates. The ability to reliably detect small CNVs is helpful, particularly if they fall within large CNVs previously identified and define a critical interval of overlap. For example, we identified one de novo CNV that only affected JUN and another that only altered TOP2A, two genes that were implicated by network analyses as interacting with transcription factors SMAD2, SMAD4 and ETS1, molecules that play important roles in cardiovascular development.
Although there is considerable complexity in CHD phenotypes, we observed no significant difference in the frequencies of rare de novo CNVs among distinct CHD sub-classifications. While CHD patients with CNVs in our cohort were more likely to have extra-cardiac phenotypes (OR: 1.5), this trend fell short of significance. Whether this finding reflects shared developmental biologic pathways among different organ systems or the possibility that CNVs perturb multiple genes that individually contribute to organ system development is unknown.
We identified several de novo CNVs that impacted established CHD genes including GATA4 and GJA5. We also identified a CHD patient with a deletion of chromosome 5q34-q35.2, encompassing NKX2-5. LOF NKX2-5 mutations are an established cause of CHD, and CNVs encompassing NKX2-5 have been previously recognized in CHD.
We identified recurrent de novo CNVs involving deletions or duplications at chromosome 15q11.2. As the proximal region of chromosome 15 is meiotically unstable due to the segmental duplications that serve as breakpoint hotspots, recurrent de novo events at this locus might reflect locus genomic instability. However, the excess burden of de novo CNVs at this locus in CHD patients compared to controls (Fisher _p_=0.007) suggests otherwise. The report of an excess burden of inherited deletions in CHD patients at this locus3 lends further evidence for pathogenicity although this study lacked information on inheritance. As CNVs at chromosome15q11.2 CNV exhibit incomplete penetrance for both neuropsychiatric and CHD phenotypes, genes affected by this could participate in inherited and sporadic CHD.
Chromosome 15q11.2 deletions and duplications are implicated in neurodevelopmental disorders including schizophrenia, intellectual disability and autism.43-45 That chromosome 15q11.2 CNVs are also associated with CHD adds to a growing list of loci (22q11,46 1q21, 7q11.23,48 16p11.2, and 16p13.11) that link cardiac malformations and neurocognitive disorders. These (and other) genetic loci may explain in part the significant co-expression of heart and brain developmental phenotypes in many children.
By integrating CNV and sequencing data from WES, we also identified candidate genes within CNV regions that may cause dominant or recessive forms of CHD. We present the first human ETS1 LOF mutation that likely contributes to Jacobsen syndrome. We also identified a rare inherited and predicted deleterious CTBP2 missense variant that is hemizygous due to a de novo CNV deletion, associated with a CHD phenotype comparable to that observed in _Ctbp2-_null mice. Continued integration of CNV and sequence data should enable more comprehensive assessments of genetic causes of disease. The current study provides suggestive data, and sequencing large cohorts of CHD patients for mutations in these two genes will be necessary to unambiguously prove the role of these genes in CHD.
Network analyses by DAPPLE was more successful in elucidating novel network biology than DAVID and WebGestalt, which rely heavily on previously annotated gene sets and are challenged by the addition of unrelated genes encoded with CNV intervals along with pathogenic genes. If pathogenic CNVs on average contain one main causal gene and approximately five unrelated genes, then we might expect DAVID and WebGestalt to be less informative for CNV network analyses.51 Conversely, DAPPLE, based on proteome-wide protein-protein interaction data rather than previously curated gene lists, calculates p-values through within-degree node-label permutation, which is more permissive to background noise.36
DAPPLE network analysis reinforced the central role of transcriptional regulation in congenital heart disease. The identification of one network, including NKX2.5/GATA4, provided a robust positive control as protein-protein interactions and substantial contributions by these molecules to CHD are previously described. Direct protein-protein interactions between ETS1/JUN/TOP2A have also been reported,54-56 but this network has not been previously implicated in CHD. In an expanded network analysis of these molecules that included rare LOF mutations identified from exome sequencing, JUN was linked to SMAD2 and SMAD4, molecules that participate in cardiac development through the TGF-β signaling pathway.57-60
We focused our current analysis solely on rare de novo CNVs. As the etiology of CHDs is known to be polygenic, and incomplete penetrance of genes for CHD has been previously described, future analyses of rare inherited CNVs may expand these findings.
The novel de novo CNVs we report should be considered provisional pending replication in independent studies. Replication of the overall effect and the magnitude of the risk of these identified variants is needed. While it is not yet possible to draw a conclusion about whether any particular de novo CNV is causal, the identification of additional CNVs and mutations in specific genes within the CNV intervals will be required to validate the new loci identified here.
In summary, integration of high resolution complementary platforms for CNV and sequence data on large numbers of patients with CHD has proven valuable to define the underlying genomic architecture of CHD and expand the genes and networks involved in cardiac development and is likely applicable to the study of other diseases.
Supplementary Material
304458R2 Acknowledgement Permissions
304458R2 Online Data Supplement
CircRes_CIRCRES-2014-304458.xml
Novelty and Significance.
What Is Known?
- Congenital heart disease (CHD) is amoung the most common birth defects.
- Many genomic loci are implicated in CHD, but most cases are of unknown etiology.
What New Information Does This Article Contribute?
- To determine impact of de novo CNVs in CHD, we found an increased de novo copy number variation (CNV) frequency in CHD families compared to healthy control families (termed burden).
- We found true recurrent de novos to find and define significant CHD CNV genes.
- We performed network analysis to assess biological gene function of single de novo CNVs.
CHD is amoung the most common birth defects. Many genomic loci are implicated in CHD, but most cases are of unknown etiology. A significant increase in CNV burden was observed when comparing CHD trios with healthy trios, using either SNP array (_p_=7x10−5, Odds Ratio (OR)=4.6) or WES data (_p_=6x10−4, OR=3.5) and remained after removing 16% of de novo CNV loci previously reported as pathogenic (_p_=0.02, OR=2.7). We observed recurrent de novo CNVs on 15q11.2 encompassing CYFIP1, NIPA1, and NIPA2 and single de novo CNVs encompassing DUSP1, JUN, JUP, MED15, MED9, PTPRE SREBF1, TOP2A, and ZEB2, genes that interact with established CHD proteins NKX2-5 and GATA4. Integrating de novo variants in WES and CNV data suggests that ETS1 is the pathogenic gene altered by 11q24.2-q25 deletions in Jacobsen syndrome and that CTBP2 is the pathogenic gene in 10q sub-telomeric deletions. This is the first large cohort study of CHD families using WES and dense state of the art SNP arrays for integrative de novo CNV discovery. The new loci implicated here provide novel diagnostic markers for early detection of CHD and novel therapeutic drug targets.
Table 1.
Confirmed rare de novo CNVs in discovery cohort. Genomic coordinates refer to hg19.
| ID | Chr | Start | End | Band | CNV1 | Syndrome/ gene | AnalysisObserved2 | Cardiac Lesion:(diagnosis)3 | ParentOrigin | Extra-cardiac | Ngenes | Size(kb) |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1-01401 | 1 | 59247993 | 59251097 | p32.1 | 1 | JUN | A | LVOT (HLHS) | - | - | 1 | 3.1 |
| 1-03171 | 1 | 145586403 | 145799634 | q21.1 | 3 | 1q21.1 dup/ GJA54 | A E | CTD (TOF/APVS) | - | - | 7 | 213.2 |
| 1-01036 | 1 | 146631133 | 147416212 | q21.1 | 3 | 1q21.1 dup/ GJA54 | E | CTD (TOF) | M | - | 15 | 785.1 |
| 1-01486 | 1 | 194201171 | 194304070 | q24.2-q25 | 3 | CDC73 | A | LVOT (HLHS) | - | Yes | 0 | 102.9 |
| 1-01518 | 1 | 248750565 | 248795110 | q44 | 3 | OR2T10,OR2T11 | A | LVOT (HLHS) | - | - | 2 | 44.5 |
| 1-01536 | 2 | 70168995 | 70359345 | p13.3 | 1 | PCBP1 | A | CTD (TOF/PA) | - | - | 5 | 190.4 |
| 1-01401 | 2 | 102493466 | 103001458 | q11.2-q12.1 | 1 | MAP4K4 | E | LVOT (HLHS) | - | - | 6 | 508.0 |
| 1-01401 | 2 | 145155868 | 145274931 | q22.3 | 1 | Mowat-Wilson/ ZEB24 | E | LVOT (HLHS) | - | - | 1 | 119.1 |
| 1-00762 | 3 | 60661 | 11712230 | p26.1 | 3 | ARL8B,ARPC4,CAMK1,CAV3,CRBN,EMC3,ITPR1,SEC13,SETD5,VGLL4 | A | ASD/PS (ASD) | - | Yes | 103 | 11651.6 |
| 1-01049 | 3 | 15637812 | 15643461 | p25.1 | 3 | BTD,HACL1 | E | CTD (TOF) | - | - | 2 | 5.6 |
| 1-01045 | 3 | 47780965 | 48309270 | p21.31 | 3 | CDC25A,DHX30,MAP4,SMARCC1 | A | LVOT (HLHS) | - | - | 14 | 528.3 |
| 1-02093 | 3 | 197143652 | 197186111 | q29 | 3 | BDH1 | A | CTD (TOF/PA) | - | Yes | 0 | 42.5 |
| 1-00771 | 4 | 185603346 | 185638397 | q34.1 | 1 | CENPU,PRIMPOL | E | CTD (DTGA/VSD) | P | Yes | 2 | 35.1 |
| 1-00789 | 5 | 136464 | 232969 | p15.33 | 3 | CCDC127,LRRC14B,PLEKHG4B,SDHA | A | CTD (TOF) | - | - | 4 | 96.5 |
| 1-00113 | 5 | 133706994 | 133730455 | q31.1 | 1 | UBE2B | A | CTD (TOF/PA) | - | Yes | 1 | 23.5 |
| 1-00296 | 5 | 166386727 | 173073664 | q34-q35.2 | 1 | NKX2.54 | A | CTD (TOF) | M | Yes | 53 | 6686.9 |
| 1-01916 | 6 | 36646788 | 36651971 | p21.2 | 1 | CDKN1A | A | HTX (HTX) | - | - | 1 | 5.2 |
| 1-01049 | 6 | 43484783 | 43485159 | p21.1 | 3 | POLR1C | E | CTD (TOF) | - | - | 1 | 0.4 |
| 1-00096 | 7 | 50179707 | 50191153 | p12.2 | 1 | C7orf72 | E | CTD (TOF/PA) | - | Yes | 1 | 11.4 |
| 1-00800 | 7 | 72719386 | 74138603 | q11.23 | 1 | Williams syndrome_4_ | A | CTD (VSD/PS) | P | Yes | 34 | 1419.2 |
| 1-00540 | 7 | 72721123 | 74140708 | q11.23 | 1 | Williams syndrome_4_ | A | LVOT (ASD) | M | Yes | 34 | 1419.6 |
| 1-00977 | 7 | 138258252 | 143807632 | q24-q25 | 1 | C7orf55,FAM115A,LUC7L2,MKRN1,NDUFB2,UBN2,ZC3HAV1L,ZYX | E | CTD (TOF) | - | - | 175 | 5549.4 |
| 1-01995 | 7 | 142334207 | 142460871 | q34 | 1 | MTRNR2L6,PRSS1 | E | CTD (TOF) | M | - | 15 | 126.7 |
| 1-01562 | 8 | 8067768 | 12530976 | p22.1-p23.1 | 3 | GATA44 | A | CTD (TOF) | - | - | 75 | 4463.2 |
| 1-02625 | 8 | 8102183 | 12190106 | p23.1 | 3 | GATA44 | A | LVOT (CoA) | M | Yes | 62 | 4087.9 |
| 1-00566 | 8 | 11606428 | 11710963 | p23.1 | 1 | GATA44 | A E | CTD (TOF) | - | - | 6 | 104.5 |
| 1-00948 | 8 | 119053343 | 119064098 | q24.1 | 1 | EXT1 | A | LVOT (CoA) | P | Yes | 1 | 10.8 |
| 1-02360 | 9 | 5302500 | 5337760 | p24.1 | 3 | RLN1,RLN2 | A | CTD (ASD) | - | Yes | 3 | 35.3 |
| 1-01852 | 11 | 34458230 | 34460862 | p13 | 1 | CAT | A | CTD (VSD) | - | - | 1 | 2.6 |
| 1-00565 | 11 | 42968283 | 42970488 | p12 | 3 | HNRNPKP3 | A | LVOTO (ASD) | - | - | 0 | 2.2 |
| 1-01536 | 11 | 65157239 | 65408708 | q13.1 | 1 | EHBP1L1,LTBP3,MAP3K11,PCNXL3,SCYL1,SSSCA1 | A | CTD (TOF/PA) | - | - | 14 | 251.5 |
| 1-00230 | 11 | 86939592 | 87025456 | q14.2 | 1 | TMEM135 | A E | LVOT (ASD) | P | Yes | 1 | 85.9 |
| 1-01486 | 11 | 125641368 | 134943190 | q24.2-q25 | 1 | Jacobsen / ETS14 | A E | LVOT (HLHS) | P | Yes | 73 | 9301.8 |
| 1-00795 | 11 | 134598043 | 134617838 | q25 | 3 | LOC283177 | A | CTD (VSD) | M | - | 0 | 19.8 |
| 1-00124 | 12 | 8003758 | 8123306 | p13.31 | 3 | SLC2A14,SLC2A3 | A | LVOT (As/HLHS) | - | - | 3 | 119.5 |
| 1-00050 | 12 | 52845952 | 52862783 | q13.13 | 1 | KRT6C | A | LVOT (HLHS) | - | - | 1 | 16.8 |
| 1-02411 | 14 | 58860893 | 58881694 | q23.1 | 1 | TIMM9,TOMM20L | A | CTD (TOF) | - | - | 2 | 20.8 |
| 1-01049 | 14 | 74551632 | 74551731 | q24.3 | 3 | LIN52 | E | CTD (TOF) | - | - | 1 | 0.1 |
| 1-00192 | 15 | 22296985 | 23161330 | q11.2 | 3 | 1 MB from PW / CYFIP14 | A | LVOT (CoA) | - | - | 20 | 864.3 |
| 1-00315 | 15 | 22750305 | 23140114 | q11.2 | 3 | 1 MB from PW / CYFIP14 | A | LVOT (CoA) | M | - | 5 | 389.8 |
| 1-01396 | 15 | 22750305 | 23228712 | q11.2 | 1 | 1 MB from PW / CYFIP14 | A E | CTD (TOF/PA) | P | - | 6 | 478.4 |
| 1-00243 | 15 | 22835893 | 23062345 | q11.2 | 1 | 1 MB from PW / CYFIP14 | E | LVOT (CoA) | P | Yes | 4 | 226.5 |
| 1-01994 | 15 | 28389771 | 28446734 | q13.2 | 1 | HERC2 | E | LVOT (ASD) | P | - | 1 | 57.0 |
| 1-01696 | 15 | 44833588 | 44856873 | q21.1 | 1 | EIF3J,SPG11 | A E | CTD (TriAtresia/DTGA) | - | - | 2 | 23.3 |
| 1-01941 | 15 | 88761539 | 88779300 | q25.3 | 3 | NTRK3 | A | CTD (TOF/DTGA) | P | - | 1 | 17.8 |
| 1-01427 | 17 | 21562473 | 22252439 | p11.2 | 1 | FAM27L,FLJ36000,MTRNR2L1 | A | HTX (HTX) | - | Yes | 7 | 690.0 |
| 1-00561 | 17 | 27962393 | 28099002 | q11.2 | 1 | SSH2 | A | LVOT (ASD) | - | Yes | 3 | 136.6 |
| 1-01995 | 17 | 38544624 | 38548586 | q21.1 | 1 | TOP2A | A E | CTD (TOF) | - | - | 1 | 4.0 |
| 1-01049 | 17 | 39845210 | 39846477 | q21.2 | 3 | EIF1 | E | CTD (TOF) | - | - | 2 | 1.3 |
| 1-01588 | 18 | 65138642 | 78015180 | q22.1-q23 | 1 | NFATC14 | A | LVOT (CoA) | - | Yes | 58 | 12876.5 |
| 1-02170 | 19 | 20601006 | 20717536 | p12 | 1 | ZNF826P | A | CTD (TOF) | - | Yes | 1 | 116.5 |
| 1-00174 | 19 | 40515744 | 40681387 | q13.2 | 1 | ZNF546,ZNF780A,ZNF780B | A | CTD (TOF/PA) | - | Yes | 4 | 165.6 |
| 1-01536 | 19 | 47792293 | 47905132 | q13.33 | 1 | C5AR1,C5AR2,DHX34 | A | CTD (TOF/PA) | - | - | 3 | 112.8 |
| 1-00730 | 20 | 14529657 | 14583899 | p12.2 | 1 | MACROD2,MACROD2-IT1 | A | CTD (DTGA) | - | - | 2 | 54.2 |
| 1-01194 | 22 | 18844632 | 21500000 | q11.2 | 1 | DiGeorge / TBX14 | A | CTD (VSD) | P | Yes | 80 | 2655.4 |
| 1-00113 | 22 | 18886915 | 22000000 | q11.2 | 1 | DiGeorge / TBX14 | A E | CTD (TOF/PA) | P | Yes | 96 | 3113.1 |
| 1-01836 | 22 | 19020529 | 21380382 | q11.2 | 1 | DiGeorge / TBX14 | A E | CTD (TOF) | M | - | 70 | 2359.9 |
| 1-00988 | 22 | 20733495 | 21464479 | q11.2 | 1 | DiGeorge / TBX14 | A | CTD (HLHS/HTX) | M | Yes | 31 | 731.0 |
| 1-02133 | 22 | 25661725 | 25919492 | q11.23 | 3 | 22q11 distal duplication_4_ | A | CTD (TOF) | - | - | 4 | 257.8 |
| 1-00425 | 22 | 36038076 | 36149338 | q12.3 | 1 | APOL5,APOL6,RBFOX2 | A E | LVOT (HLHS) | - | - | 4 | 111.3 |
| 1-01427 | 22 | 42522638 | 42531210 | q13.2 | 3 | CYP2D6 | A | HTX (HTX) | - | Yes | 2 | 8.6 |
| 1-01941 | X | 23003525 | 23086619 | p22.11 | 3 | DDX53,RP11-40F8.2 | A | CTD (TOF/DTGA) | - | - | 1 | 83.1 |
| 1-00197 | X | 148685645 | 148693146 | q28 | 3 | TMEM185A | E | LVOT (HLHS) | - | Yes | 1 | 7.5 |
ACKNOWLEDGMENTS
The authors are grateful to the patients and families who participated in this research. We thank the following team members for contributions to patient recruitment: D. Awad, K. Celis, D. Etwaru, J. Kline, R. Korsin, A. Lanz, E. Marquez, J. K. Sond, A. Wilpers, R. Yee (Columbia Medical School); A. Roberts, K. Boardman, J. Geva, J. Gorham, B. McDonough, A. Monafo, J. Stryker (Harvard Medical School); N. Cross (Yale School of Medicine); S. M. Edman, J. L. Garbarini, J. E. Tusi, S. H. Woyciechowski (Children's Hospital of Philadelphia); R. Kim, J. Ellashek and N. Tran (Children's Hospital of Los Angeles); K. Flack (University College London); A. Romano, D.Gruber, N. Stellato (Steve and Alexandra Cohen Children's Medical Center of New York); D. Guevara, A. Julian, M. Mac Neal, C. Mintz (Icahn School of Medicine at Mount Sinai); and G. Porter and E. Taillie (University of Rochester School of Medicine and Dentistry). We also thank V. Spotlow, P. Candrea, K. Pavlik and M. Sotiropoulos for their expert production of exome sequences, and we thank M. Lemma, C. Kim, F. G. Otieno, M. Khan and K. Thomas for their expert production of genome wide genotypes.
We are grateful to all of the families at the participating SFARI Simplex Collection (SSC) sites, as well as the principal investigators (A. Beaudet, R. Bernier, J. Constantino, E. Cook, E. Fombonne, D. Geschwind, D. Grice, A. Klin, D. Ledbetter, C. Lord, C. Martin, D. Martin, R. Maxim, J. Miles, O. Ousley, K. Pelphrey, B. Peterson, J. Piggot, C. Saulnier, M. State, W. Stone, J. Sutcliffe, C. Walsh, E. Wijsman) and the coordinators and staff at the SSC clinical sites.
The authors thank New England Research Institutes (NERI) S. Tennstedt, B. Williams, D. Nash, J. Barenholtz, K. Cucchi, K. Dandreo, S. Yates, T. Hamza, and C. Taglienti.
SOURCES OF FUNDING
This work was supported by the National Institutes of Health (NIH) National Heart, Lung, and Blood Institute (NHLBI) Pediatric Cardiac Genomics Consortium (U01-HL098188, U01-HL098147, U01-HL098153, U01-HL098163, U01-HL098123, U01-HL098162) and in part by the Simons Foundation for Autism Research and the NIH Centers for Mendelian Genomics (5U54HG006504).
Nonstandard Abbreviations and Acronyms
CTD
conotruncal defect
LVOT
Left Ventricular Outflow Tract Obstruction
TA
truncus arteriosus
TOF
tetralogy of Fallot
HLH
hypoplastic left heart syndrome
APVS
Absent pulmonary valve syndrome
ASD
Atrial septal defect
CoA
Coarctation of the Aorta
DTGA
dextro-Transposition of the great arteries
HTX
Heterotaxy
PA
Pulmonary Atresia
PS
Pulmonary Stenosis
TriAtresia
Tricuspid atresia
VSD
Ventricular Septal Defect
Footnotes
C.S., E.G., B.D.G., R.L., J.S., H.H., and W.K.C. are co-senior authors.
REFERENCES
- 1.Hoffman JI, Kaplan S. The incidence of congenital heart disease. J Am Coll Cardiol. 2002;39:1890–1900. doi: 10.1016/s0735-1097(02)01886-7. [DOI] [PubMed] [Google Scholar]
- 2.Tennant PW, Pearce MS, Bythell M, Rankin J. 20-year survival of children born with congenital anomalies: A population-based study. Lancet. 2010;375:649–656. doi: 10.1016/S0140-6736(09)61922-X. [DOI] [PubMed] [Google Scholar]
- 3.Soemedi R, Wilson IJ, Bentham J, Darlay R, Topf A, Zelenika D, Cosgrove C, Setchfield K, Thornborough C, Granados-Riveron J, Blue GM, Breckpot J, Hellens S, Zwolinkski S, Glen E, Mamasoula C, Rahman TJ, Hall D, Rauch A, Devriendt K, Gewillig M, J OS, Winlaw DS, Bu'Lock F, Brook JD, Bhattacharya S, Lathrop M, Santibanez-Koref M, Cordell HJ, Goodship JA, Keavney BD. Contribution of global rare copy-number variants to the risk of sporadic congenital heart disease. Am J Hum Genet. 2012;91:489–501. doi: 10.1016/j.ajhg.2012.08.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Fahed AC, Gelb BD, Seidman JG, Seidman CE. Genetics of congenital heart disease: The glass half empty. Circ Res. 2013;112:707–720. doi: 10.1161/CIRCRESAHA.112.300853. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Greenway SC, Pereira AC, Lin JC, DePalma SR, Israel SJ, Mesquita SM, Ergul E, Conta JH, Korn JM, McCarroll SA, Gorham JM, Gabriel S, Altshuler DM, Quintanilla-Dieck Mde L, Artunduaga MA, Eavey RD, Plenge RM, Shadick NA, Weinblatt ME, De Jager PL, Hafler DA, Breitbart RE, Seidman JG, Seidman CE. De novo copy number variants identify new genes and loci in isolated sporadic tetralogy of fallot. Nat Genet. 2009;41:931–935. doi: 10.1038/ng.415. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Silversides CK, Lionel AC, Costain G, Merico D, Migita O, Liu B, Yuen T, Rickaby J, Thiruvahindrapuram B, Marshall CR, Scherer SW, Bassett AS. Rare copy number variations in adults with tetralogy of fallot implicate novel risk gene pathways. PLoS Genet. 2012;8:e1002843. doi: 10.1371/journal.pgen.1002843. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Erdogan F, Larsen LA, Zhang L, Tumer Z, Tommerup N, Chen W, Jacobsen JR, Schubert M, Jurkatis J, Tzschach A, Ropers HH, Ullmann R. High frequency of submicroscopic genomic aberrations detected by tiling path array comparative genome hybridisation in patients with isolated congenital heart disease. J Med Genet. 2008;45:704–709. doi: 10.1136/jmg.2008.058776. [DOI] [PubMed] [Google Scholar]
- 8.Goldmuntz E, Paluru P, Glessner J, Hakonarson H, Biegel JA, White PS, Gai X, Shaikh TH. Microdeletions and microduplications in patients with congenital heart disease and multiple congenital anomalies. Congenit Heart Dis. 2011;6:592–602. doi: 10.1111/j.1747-0803.2011.00582.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Luo C, Yang YF, Yin BL, Chen JL, Huang C, Zhang WZ, Wang J, Zhang H, Yang JF, Tan ZP. Microduplication of 3p25.2 encompassing raf1 associated with congenital heart disease suggestive of noonan syndrome. Am J Med Genet A. 2012;158A:1918–1923. doi: 10.1002/ajmg.a.35471. [DOI] [PubMed] [Google Scholar]
- 10.Priest JR, Girirajan S, Vu TH, Olson A, Eichler EE, Portman MA. Rare copy number variants in isolated sporadic and syndromic atrioventricular septal defects. Am J Med Genet A. 2012;158A:1279–1284. doi: 10.1002/ajmg.a.35315. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Tomita-Mitchell A, Maslen CL, Morris CD, Garg V, Goldmuntz E. Gata4 sequence variants in patients with congenital heart disease. J Med Genet. 2007;44:779–783. doi: 10.1136/jmg.2007.052183. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Christiansen J, Dyck JD, Elyas BG, Lilley M, Bamforth JS, Hicks M, Sprysak KA, Tomaszewski R, Haase SM, Vicen-Wyhony LM, Somerville MJ. Chromosome 1q21.1 contiguous gene deletion is associated with congenital heart disease. Circ Res. 2004;94:1429–1435. doi: 10.1161/01.RES.0000130528.72330.5c. [DOI] [PubMed] [Google Scholar]
- 13.Carey AS, Liang L, Edwards J, Brandt T, Mei H, Sharp AJ, Hsu DT, Newburger JW, Ohye RG, Chung WK, Russell MW, Rosenfeld JA, Shaffer LG, Parides MK, Edelmann L, Gelb BD. Effect of copy number variants on outcomes for infants with single ventricle heart defects. Circ Cardiovasc Genet. 2013;6:444–451. doi: 10.1161/CIRCGENETICS.113.000189. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Breckpot J, Thienpont B, Peeters H, de Ravel T, Singer A, Rayyan M, Allegaert K, Vanhole C, Eyskens B, Vermeesch JR, Gewillig M, Devriendt K. Array comparative genomic hybridization as a diagnostic tool for syndromic heart defects. J Pediatr. 2010;156:810–817. 817, e811–817, e814. doi: 10.1016/j.jpeds.2009.11.049. [DOI] [PubMed] [Google Scholar]
- 15.Erdogan F, Larsen LA, Zhang L, Tumer Z, Tommerup N, Chen W, Jacobsen JR, Schubert M, Jurkatis J, Tzschach A, Ropers HH, Ullmann R. High frequency of submicroscopic genomic aberrations detected by tiling path array comparative genome hybridisation in patients with isolated congenital heart disease. J Med Genet. 2008;45:704–709. doi: 10.1136/jmg.2008.058776. [DOI] [PubMed] [Google Scholar]
- 16.Hitz MP, Lemieux-Perreault LP, Marshall C, Feroz-Zada Y, Davies R, Yang SW, Lionel AC, D'Amours G, Lemyre E, Cullum R, Bigras JL, Thibeault M, Chetaille P, Montpetit A, Khairy P, Overduin B, Klaassen S, Hoodless P, Awadalla P, Hussin J, Idaghdour Y, Nemer M, Stewart AF, Boerkoel C, Scherer SW, Richter A, Dube MP, Andelfinger G. Rare copy number variants contribute to congenital left-sided heart disease. PLoS Genet. 2012;8:e1002903. doi: 10.1371/journal.pgen.1002903. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Payne AR, Chang SW, Koenig SN, Zinn AR, Garg V. Submicroscopic chromosomal copy number variations identified in children with hypoplastic left heart syndrome. Pediatr Cardiol. 2012;33:757–763. doi: 10.1007/s00246-012-0208-9. [DOI] [PubMed] [Google Scholar]
- 18.Richards AA, Santos LJ, Nichols HA, Crider BP, Elder FF, Hauser NS, Zinn AR, Garg V. Cryptic chromosomal abnormalities identified in children with congenital heart disease. Pediatr Res. 2008;64:358–363. doi: 10.1203/PDR.0b013e31818095d0. [DOI] [PubMed] [Google Scholar]
- 19.Thienpont B, Mertens L, de Ravel T, Eyskens B, Boshoff D, Maas N, Fryns JP, Gewillig M, Vermeesch JR, Devriendt K. Submicroscopic chromosomal imbalances detected by array-cgh are a frequent cause of congenital heart defects in selected patients. Eur Heart J. 2007;28:2778–2784. doi: 10.1093/eurheartj/ehl560. [DOI] [PubMed] [Google Scholar]
- 20.Warburton D, Ronemus M, Kline J, Jobanputra V, Williams I, Anyane-Yeboa K, Chung W, Yu L, Wong N, Awad D, Yu CY, Leotta A, Kendall J, Yamrom B, Lee YH, Wigler M, Levy D. The contribution of de novo and rare inherited copy number changes to congenital heart disease in an unselected sample of children with conotruncal defects or hypoplastic left heart disease. Hum Genet. 2014;133:11–27. doi: 10.1007/s00439-013-1353-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Goldmuntz E, Clark BJ, Mitchell LE, Jawad AF, Cuneo BF, Reed L, McDonald-McGinn D, Chien P, Feuer J, Zackai EH, Emanuel BS, Driscoll DA. Frequency of 22q11 deletions in patients with conotruncal defects. J Am Coll Cardiol. 1998;32:492–498. doi: 10.1016/s0735-1097(98)00259-9. [DOI] [PubMed] [Google Scholar]
- 22.Rauch R, Hofbeck M, Zweier C, Koch A, Zink S, Trautmann U, Hoyer J, Kaulitz R, Singer H, Rauch A. Comprehensive genotype-phenotype analysis in 230 patients with tetralogy of fallot. J Med Genet. 2010;47:321–331. doi: 10.1136/jmg.2009.070391. [DOI] [PubMed] [Google Scholar]
- 23.Chieffo C, Garvey N, Gong W, Roe B, Zhang G, Silver L, Emanuel BS, Budarf ML. Isolation and characterization of a gene from the digeorge chromosomal region homologous to the mouse tbx1 gene. Genomics. 1997;43:267–277. doi: 10.1006/geno.1997.4829. [DOI] [PubMed] [Google Scholar]
- 24.Pediatric Cardiac Genomics C, Gelb B, Brueckner M, Chung W, Goldmuntz E, Kaltman J, Kaski JP, Kim R, Kline J, Mercer-Rosa L, Porter G, Roberts A, Rosenberg E, Seiden H, Seidman C, Sleeper L, Tennstedt S, Kaltman J, Schramm C, Burns K, Pearson G, Rosenberg E. The congenital heart disease genetic network study: Rationale, design, and early results. Circ Res. 2013;112:698–706. doi: 10.1161/CIRCRESAHA.111.300297. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Sanders SJ, Ercan-Sencicek AG, Hus V, Luo R, Murtha MT, Moreno-De-Luca D, Chu SH, Moreau MP, Gupta AR, Thomson SA, Mason CE, Bilguvar K, Celestino-Soper PB, Choi M, Crawford EL, Davis L, Wright NR, Dhodapkar RM, DiCola M, DiLullo NM, Fernandez TV, Fielding-Singh V, Fishman DO, Frahm S, Garagaloyan R, Goh GS, Kammela S, Klei L, Lowe JK, Lund SC, McGrew AD, Meyer KA, Moffat WJ, Murdoch JD, O'Roak BJ, Ober GT, Pottenger RS, Raubeson MJ, Song Y, Wang Q, Yaspan BL, Yu TW, Yurkiewicz IR, Beaudet AL, Cantor RM, Curland M, Grice DE, Gunel M, Lifton RP, Mane SM, Martin DM, Shaw CA, Sheldon M, Tischfield JA, Walsh CA, Morrow EM, Ledbetter DH, Fombonne E, Lord C, Martin CL, Brooks AI, Sutcliffe JS, Cook EH, Jr., Geschwind D, Roeder K, Devlin B, State MW. Multiple recurrent de novo cnvs, including duplications of the 7q11.23 williams syndrome region, are strongly associated with autism. Neuron. 2011;70:863–885. doi: 10.1016/j.neuron.2011.05.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Zaidi S, Choi M, Wakimoto H, Ma L, Jiang J, Overton JD, Romano-Adesman A, Bjornson RD, Breitbart RE, Brown KK, Carriero NJ, Cheung YH, Deanfield J, DePalma S, Fakhro KA, Glessner J, Hakonarson H, Italia MJ, Kaltman JR, Kaski J, Kim R, Kline JK, Lee T, Leipzig J, Lopez A, Mane SM, Mitchell LE, Newburger JW, Parfenov M, Pe'er I, Porter G, Roberts AE, Sachidanandam R, Sanders SJ, Seiden HS, State MW, Subramanian S, Tikhonova IR, Wang W, Warburton D, White PS, Williams IA, Zhao H, Seidman JG, Brueckner M, Chung WK, Gelb BD, Goldmuntz E, Seidman CE, Lifton RP. De novo mutations in histone-modifying genes in congenital heart disease. Nature. 2013;498:220–223. doi: 10.1038/nature12141. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.De Rubeis S, Pasciuto E, Li KW, Fernandez E, Di Marino D, Buzzi A, Ostroff LE, Klann E, Zwartkruis FJ, Komiyama NH, Grant SG, Poujol C, Choquet D, Achsel T, Posthuma D, Smit AB, Bagni C. Cyfip1 coordinates mrna translation and cytoskeleton remodeling to ensure proper dendritic spine formation. Neuron. 2013;79:1169–1182. doi: 10.1016/j.neuron.2013.06.039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Meng Q, Rayala SK, Gururaj AE, Talukder AH, O'Malley BW, Kumar R. Signaling-dependent and coordinated regulation of transcription, splicing, and translation resides in a single coregulator, pcbp1. Proc Natl Acad Sci U S A. 2007;104:5866–5871. doi: 10.1073/pnas.0701065104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Ye M, Coldren C, Liang X, Mattina T, Goldmuntz E, Benson DW, Ivy D, Perryman MB, Garrett-Sinha LA, Grossfeld P. Deletion of ets-1, a gene in the jacobsen syndrome critical region, causes ventricular septal defects and abnormal ventricular morphology in mice. Hum Mol Genet. 2010;19:648–656. doi: 10.1093/hmg/ddp532. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Exome variant server, nhlbi go exome sequencing project (esp) seattle, wa: [january 2014]. (url: Http://evs.gs.washington.edu/evs/) [Google Scholar]
- 31.Genomes Project C, Abecasis GR, Auton A, Brooks LD, DePristo MA, Durbin RM, Handsaker RE, Kang HM, Marth GT, McVean GA. An integrated map of genetic variation from 1,092 human genomes. Nature. 2012;491:56–65. doi: 10.1038/nature11632. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Courtens W, Wuyts W, Rooms L, Pera SB, Wauters J. A subterminal deletion of the long arm of chromosome 10: A clinical report and review. Am J Med Genet A. 2006;140:402–409. doi: 10.1002/ajmg.a.31053. [DOI] [PubMed] [Google Scholar]
- 33.Hildebrand JD, Soriano P. Overlapping and unique roles for c-terminal binding protein 1 (ctbp1) and ctbp2 during mouse development. Mol Cell Biol. 2002;22:5296–5307. doi: 10.1128/MCB.22.15.5296-5307.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Breckpot J, Tranchevent LC, Thienpont B, Bauters M, Troost E, Gewillig M, Vermeesch JR, Moreau Y, Devriendt K, Van Esch H. Bmpr1a is a candidate gene for congenital heart defects associated with the recurrent 10q22q23 deletion syndrome. Eur J Med Genet. 2012;55:12–16. doi: 10.1016/j.ejmg.2011.10.003. [DOI] [PubMed] [Google Scholar]
- 35.Huang da W, Sherman BT, Tan Q, Kir J, Liu D, Bryant D, Guo Y, Stephens R, Baseler MW, Lane HC, Lempicki RA. David bioinformatics resources: Expanded annotation database and novel algorithms to better extract biology from large gene lists. Nucleic Acids Res. 2007;35:W169–175. doi: 10.1093/nar/gkm415. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Rossin EJ, Lage K, Raychaudhuri S, Xavier RJ, Tatar D, Benita Y. International Inflammatory Bowel Disease Genetics C, Cotsapas C, Daly MJ. Proteins encoded in genomic regions associated with immune-mediated disease physically interact and suggest underlying biology. PLoS Genet. 2011;7:e1001273. doi: 10.1371/journal.pgen.1001273. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Wang J, Duncan D, Shi Z, Zhang B. Web-based gene set analysis toolkit (webgestalt): Update 2013. Nucleic Acids Res. 2013;41:W77–83. doi: 10.1093/nar/gkt439. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Fromer M, Moran JL, Chambert K, Banks E, Bergen SE, Ruderfer DM, Handsaker RE, McCarroll SA, O'Donovan MC, Owen MJ, Kirov G, Sullivan PF, Hultman CM, Sklar P, Purcell SM. Discovery and statistical genotyping of copy-number variation from whole-exome sequencing depth. Am J Hum Genet. 2012;91:597–607. doi: 10.1016/j.ajhg.2012.08.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Poultney CS, Goldberg AP, Drapeau E, Kou Y, Harony-Nicolas H, Kajiwara Y, De Rubeis S, Durand S, Stevens C, Rehnstrom K, Palotie A, Daly MJ, Ma'ayan A, Fromer M, Buxbaum JD. Identification of small exonic cnv from whole-exome sequence data and application to autism spectrum disorder. Am J Hum Genet. 2013;93:607–619. doi: 10.1016/j.ajhg.2013.09.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Schott JJ, Benson DW, Basson CT, Pease W, Silberbach GM, Moak JP, Maron BJ, Seidman CE, Seidman JG. Congenital heart disease caused by mutations in the transcription factor nkx2-5. Science. 1998;281:108–111. doi: 10.1126/science.281.5373.108. [DOI] [PubMed] [Google Scholar]
- 41.Baekvad-Hansen M, Tumer Z, Delicado A, Erdogan F, Tommerup N, Larsen LA. Delineation of a 2.2 mb microdeletion at 5q35 associated with microcephaly and congenital heart disease. Am J Med Genet A. 2006;140:427–433. doi: 10.1002/ajmg.a.31087. [DOI] [PubMed] [Google Scholar]
- 42.Buysse K, Crepel A, Menten B, Pattyn F, Antonacci F, Veltman JA, Larsen LA, Tumer Z, de Klein A, van de Laar I, Devriendt K, Mortier G, Speleman F. Mapping of 5q35 chromosomal rearrangements within a genomically unstable region. J Med Genet. 2008;45:672–678. doi: 10.1136/jmg.2008.058883. [DOI] [PubMed] [Google Scholar]
- 43.Kirov G, Grozeva D, Norton N, Ivanov D, Mantripragada KK, Holmans P, International Schizophrenia C, Wellcome Trust Case Control C. Craddock N, Owen MJ, O'Donovan MC. Support for the involvement of large copy number variants in the pathogenesis of schizophrenia. Hum Mol Genet. 2009;18:1497–1503. doi: 10.1093/hmg/ddp043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Stefansson H, Meyer-Lindenberg A, Steinberg S, Magnusdottir B, Morgen K, Arnarsdottir S, Bjornsdottir G, Walters GB, Jonsdottir GA, Doyle OM, Tost H, Grimm O, Kristjansdottir S, Snorrason H, Davidsdottir SR, Gudmundsson LJ, Jonsson GF, Stefansdottir B, Helgadottir I, Haraldsson M, Jonsdottir B, Thygesen JH, Schwarz AJ, Didriksen M, Stensbol TB, Brammer M, Kapur S, Halldorsson JG, Hreidarsson S, Saemundsen E, Sigurdsson E, Stefansson K. Cnvs conferring risk of autism or schizophrenia affect cognition in controls. Nature. 2014;505:361–366. doi: 10.1038/nature12818. [DOI] [PubMed] [Google Scholar]
- 45.Stefansson H, Rujescu D, Cichon S, Pietilainen OP, Ingason A, Steinberg S, Fossdal R, Sigurdsson E, Sigmundsson T, Buizer-Voskamp JE, Hansen T, Jakobsen KD, Muglia P, Francks C, Matthews PM, Gylfason A, Halldorsson BV, Gudbjartsson D, Thorgeirsson TE, Sigurdsson A, Jonasdottir A, Jonasdottir A, Bjornsson A, Mattiasdottir S, Blondal T, Haraldsson M, Magnusdottir BB, Giegling I, Moller HJ, Hartmann A, Shianna KV, Ge D, Need AC, Crombie C, Fraser G, Walker N, Lonnqvist J, Suvisaari J, Tuulio-Henriksson A, Paunio T, Toulopoulou T, Bramon E, Di Forti M, Murray R, Ruggeri M, Vassos E, Tosato S, Walshe M, Li T, Vasilescu C, Muhleisen TW, Wang AG, Ullum H, Djurovic S, Melle I, Olesen J, Kiemeney LA, Franke B, Group. Sabatti C, Freimer NB, Gulcher JR, Thorsteinsdottir U, Kong A, Andreassen OA, Ophoff RA, Georgi A, Rietschel M, Werge T, Petursson H, Goldstein DB, Nothen MM, Peltonen L, Collier DA, St Clair D, Stefansson K. Large recurrent microdeletions associated with schizophrenia. Nature. 2008;455:232–236. doi: 10.1038/nature07229. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Kobrynski LJ, Sullivan KE. Velocardiofacial syndrome, digeorge syndrome: The chromosome 22q11.2 deletion syndromes. Lancet. 2007;370:1443–1452. doi: 10.1016/S0140-6736(07)61601-8. [DOI] [PubMed] [Google Scholar]
- 47.Mefford HC, Sharp AJ, Baker C, Itsara A, Jiang Z, Buysse K, Huang S, Maloney VK, Crolla JA, Baralle D, Collins A, Mercer C, Norga K, de Ravel T, Devriendt K, Bongers EM, de Leeuw N, Reardon W, Gimelli S, Bena F, Hennekam RC, Male A, Gaunt L, Clayton-Smith J, Simonic I, Park SM, Mehta SG, Nik-Zainal S, Woods CG, Firth HV, Parkin G, Fichera M, Reitano S, Lo Giudice M, Li KE, Casuga I, Broomer A, Conrad B, Schwerzmann M, Raber L, Gallati S, Striano P, Coppola A, Tolmie JL, Tobias ES, Lilley C, Armengol L, Spysschaert Y, Verloo P, De Coene A, Goossens L, Mortier G, Speleman F, van Binsbergen E, Nelen MR, Hochstenbach R, Poot M, Gallagher L, Gill M, McClellan J, King MC, Regan R, Skinner C, Stevenson RE, Antonarakis SE, Chen C, Estivill X, Menten B, Gimelli G, Gribble S, Schwartz S, Sutcliffe JS, Walsh T, Knight SJ, Sebat J, Romano C, Schwartz CE, Veltman JA, de Vries BB, Vermeesch JR, Barber JC, Willatt L, Tassabehji M, Eichler EE. Recurrent rearrangements of chromosome 1q21.1 and variable pediatric phenotypes. N Engl J Med. 2008;359:1685–1699. doi: 10.1056/NEJMoa0805384. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Golzio C, Katsanis N. Genetic architecture of reciprocal cnvs. Curr Opin Genet Dev. 2013;23:240–248. doi: 10.1016/j.gde.2013.04.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Ghebranious N, Giampietro PF, Wesbrook FP, Rezkalla SH. A novel microdeletion at 16p11.2 harbors candidate genes for aortic valve development, seizure disorder, and mild mental retardation. Am J Med Genet A. 2007;143A:1462–1471. doi: 10.1002/ajmg.a.31837. [DOI] [PubMed] [Google Scholar]
- 50.Golzio C, Willer J, Talkowski ME, Oh EC, Taniguchi Y, Jacquemont S, Reymond A, Sun M, Sawa A, Gusella JF, Kamiya A, Beckmann JS, Katsanis N. Kctd13 is a major driver of mirrored neuroanatomical phenotypes of the 16p11.2 copy number variant. Nature. 2012;485:363–367. doi: 10.1038/nature11091. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Huang da W, Sherman BT, Lempicki RA. Bioinformatics enrichment tools: Paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 2009;37:1–13. doi: 10.1093/nar/gkn923. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Schlesinger J, Schueler M, Grunert M, Fischer JJ, Zhang Q, Krueger T, Lange M, Tonjes M, Dunkel I, Sperling SR. The cardiac transcription network modulated by gata4, mef2a, nkx2.5, srf, histone modifications, and micrornas. PLoS Genet. 2011;7:e1001313. doi: 10.1371/journal.pgen.1001313. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Stennard FA, Costa MW, Elliott DA, Rankin S, Haast SJ, Lai D, McDonald LP, Niederreither K, Dolle P, Bruneau BG, Zorn AM, Harvey RP. Cardiac t-box factor tbx20 directly interacts with nkx2-5, gata4, and gata5 in regulation of gene expression in the developing heart. Dev Biol. 2003;262:206–224. doi: 10.1016/s0012-1606(03)00385-3. [DOI] [PubMed] [Google Scholar]
- 54.Kroll DJ, Sullivan DM, Gutierrez-Hartmann A, Hoeffler JP. Modification of DNA topoisomerase ii activity via direct interactions with the cyclic adenosine-3′,5′-monophosphate response element-binding protein and related transcription factors. Mol Endocrinol. 1993;7:305–318. doi: 10.1210/mend.7.3.8387155. [DOI] [PubMed] [Google Scholar]
- 55.Logan SK, Garabedian MJ, Campbell CE, Werb Z. Synergistic transcriptional activation of the tissue inhibitor of metalloproteinases-1 promoter via functional interaction of ap-1 and ets-1 transcription factors. J Biol Chem. 1996;271:774–782. doi: 10.1074/jbc.271.2.774. [DOI] [PubMed] [Google Scholar]
- 56.Miyamoto-Sato E, Fujimori S, Ishizaka M, Hirai N, Masuoka K, Saito R, Ozawa Y, Hino K, Washio T, Tomita M, Yamashita T, Oshikubo T, Akasaka H, Sugiyama J, Matsumoto Y, Yanagawa H. A comprehensive resource of interacting protein regions for refining human transcription factor networks. PLoS One. 2010;5:e9289. doi: 10.1371/journal.pone.0009289. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Caputo V, Cianetti L, Niceta M, Carta C, Ciolfi A, Bocchinfuso G, Carrani E, Dentici ML, Biamino E, Belligni E, Garavelli L, Boccone L, Melis D, Andria G, Gelb BD, Stella L, Silengo M, Dallapiccola B, Tartaglia M. A restricted spectrum of mutations in the smad4 tumor-suppressor gene underlies myhre syndrome. Am J Hum Genet. 2012;90:161–169. doi: 10.1016/j.ajhg.2011.12.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Chen CR, Kang Y, Siegel PM, Massague J. E2f4/5 and p107 as smad cofactors linking the tgfbeta receptor to c-myc repression. Cell. 2002;110:19–32. doi: 10.1016/s0092-8674(02)00801-2. [DOI] [PubMed] [Google Scholar]
- 59.Macias-Silva M, Abdollah S, Hoodless PA, Pirone R, Attisano L, Wrana JL. Madr2 is a substrate of the tgfbeta receptor and its phosphorylation is required for nuclear accumulation and signaling. Cell. 1996;87:1215–1224. doi: 10.1016/s0092-8674(00)81817-6. [DOI] [PubMed] [Google Scholar]
- 60.Waldrip WR, Bikoff EK, Hoodless PA, Wrana JL, Robertson EJ. Smad2 signaling in extraembryonic tissues determines anterior-posterior polarity of the early mouse embryo. Cell. 1998;92:797–808. doi: 10.1016/s0092-8674(00)81407-5. [DOI] [PubMed] [Google Scholar]
- 61.Fischbach GD, Lord C. The simons simplex collection: A resource for identification of autism genetic risk factors. Neuron. 2010;68:192–195. doi: 10.1016/j.neuron.2010.10.006. [DOI] [PubMed] [Google Scholar]
- 62.Sanders SJ, Murtha MT, Gupta AR, Murdoch JD, Raubeson MJ, Willsey AJ, Ercan-Sencicek AG, DiLullo NM, Parikshak NN, Stein JL, Walker MF, Ober GT, Teran NA, Song Y, El-Fishawy P, Murtha RC, Choi M, Overton JD, Bjornson RD, Carriero NJ, Meyer KA, Bilguvar K, Mane SM, Sestan N, Lifton RP, Gunel M, Roeder K, Geschwind DH, Devlin B, State MW. De novo mutations revealed by whole-exome sequencing are strongly associated with autism. Nature. 2012;485:237–241. doi: 10.1038/nature10945. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Glessner JT, Li J, Hakonarson H. Parsecnv integrative copy number variation association software with quality tracking. Nucleic Acids Res. 2013;41:e64. doi: 10.1093/nar/gks1346. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Wang K, Li M, Hadley D, Liu R, Glessner J, Grant SF, Hakonarson H, Bucan M. Penncnv: An integrated hidden markov model designed for high-resolution copy number variation detection in whole-genome snp genotyping data. Genome Res. 2007;17:1665–1674. doi: 10.1101/gr.6861907. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Colella S, Yau C, Taylor JM, Mirza G, Butler H, Clouston P, Bassett AS, Seller A, Holmes CC, Ragoussis J. Quantisnp: An objective bayes hidden-markov model to detect and accurately map copy number variation using snp genotyping data. Nucleic Acids Res. 2007;35:2013–2025. doi: 10.1093/nar/gkm076. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.van Karnebeek CD, Hennekam RC. Associations between chromosomal anomalies and congenital heart defects: A database search. Am J Med Genet. 1999;84:158–166. doi: 10.1002/(sici)1096-8628(19990521)84:2<158::aid-ajmg13>3.0.co;2-5. [DOI] [PubMed] [Google Scholar]
- 67.Li H, Durbin R. Fast and accurate long-read alignment with burrows-wheeler transform. Bioinformatics. 2010;26:589–595. doi: 10.1093/bioinformatics/btp698. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Cox AJ. Eland: Efficient large-scale alignment of nucleotide databases. Illumina; San Diego: 2007. [Google Scholar]
- 69.DePristo MA, Banks E, Poplin R, Garimella KV, Maguire JR, Hartl C, Philippakis AA, del Angel G, Rivas MA, Hanna M, McKenna A, Fennell TJ, Kernytsky AM, Sivachenko AY, Cibulskis K, Gabriel SB, Altshuler D, Daly MJ. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat Genet. 2011;43:491–498. doi: 10.1038/ng.806. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Cingolani P, Platts A, Wang le L, Coon M, Nguyen T, Wang L, Land SJ, Lu X, Ruden DM. A program for annotating and predicting the effects of single nucleotide polymorphisms, snpeff: Snps in the genome of drosophila melanogaster strain w1118; iso-2; iso-3. Fly (Austin) 2012;6:80–92. doi: 10.4161/fly.19695. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Pinheiro LB, Coleman VA, Hindson CM, Herrmann J, Hindson BJ, Bhat S, Emslie KR. Evaluation of a droplet digital polymerase chain reaction format for DNA copy number quantification. Anal Chem. 2012;84:1003–1011. doi: 10.1021/ac202578x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Iossifov I, Ronemus M, Levy D, Wang Z, Hakker I, Rosenbaum J, Yamrom B, Lee YH, Narzisi G, Leotta A, Kendall J, Grabowska E, Ma B, Marks S, Rodgers L, Stepansky A, Troge J, Andrews P, Bekritsky M, Pradhan K, Ghiban E, Kramer M, Parla J, Demeter R, Fulton LL, Fulton RS, Magrini VJ, Ye K, Darnell JC, Darnell RB, Mardis ER, Wilson RK, Schatz MC, McCombie WR, Wigler M. De novo gene disruptions in children on the autistic spectrum. Neuron. 2012;74:285–299. doi: 10.1016/j.neuron.2012.04.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
304458R2 Acknowledgement Permissions
304458R2 Online Data Supplement
CircRes_CIRCRES-2014-304458.xml