Genome-wide association study identifies novel loci predisposing to cutaneous melanoma - PubMed (original) (raw)
. 2011 Dec 15;20(24):5012-23.
doi: 10.1093/hmg/ddr415. Epub 2011 Sep 17.
Li-E Wang, Jeffrey E Lee, Jeffrey E Gershenwald, Wei V Chen, Shenying Fang, Roman Kosoy, Mingfeng Zhang, Abrar A Qureshi, Selina Vattathil, Christopher W Schacherer, Julie M Gardner, Yuling Wang, D Tim Bishop, Jennifer H Barrett; GenoMEL Investigators; Stuart MacGregor, Nicholas K Hayward, Nicholas G Martin, David L Duffy; Q-Mega Investigators; Graham J Mann, Anne Cust, John Hopper; AMFS Investigators; Kevin M Brown, Elizabeth A Grimm, Yaji Xu, Younghun Han, Kaiyan Jing, Caitlin McHugh, Cathy C Laurie, Kim F Doheny, Elizabeth W Pugh, Michael F Seldin, Jiali Han, Qingyi Wei
Collaborators, Affiliations
- PMID: 21926416
- PMCID: PMC3298855
- DOI: 10.1093/hmg/ddr415
Genome-wide association study identifies novel loci predisposing to cutaneous melanoma
Christopher I Amos et al. Hum Mol Genet. 2011.
Abstract
We performed a multistage genome-wide association study of melanoma. In a discovery cohort of 1804 melanoma cases and 1026 controls, we identified loci at chromosomes 15q13.1 (HERC2/OCA2 region) and 16q24.3 (MC1R) regions that reached genome-wide significance within this study and also found strong evidence for genetic effects on susceptibility to melanoma from markers on chromosome 9p21.3 in the p16/ARF region and on chromosome 1q21.3 (ARNT/LASS2/ANXA9 region). The most significant single-nucleotide polymorphisms (SNPs) in the 15q13.1 locus (rs1129038 and rs12913832) lie within a genomic region that has profound effects on eye and skin color; notably, 50% of variability in eye color is associated with variation in the SNP rs12913832. Because eye and skin colors vary across European populations, we further evaluated the associations of the significant SNPs after carefully adjusting for European substructure. We also evaluated the top 10 most significant SNPs by using data from three other genome-wide scans. Additional in silico data provided replication of the findings from the most significant region on chromosome 1q21.3 rs7412746 (P = 6 × 10(-10)). Together, these data identified several candidate genes for additional studies to identify causal variants predisposing to increased risk for developing melanoma.
Figures
Figure 1.
Description of the flow of samples and markers through the genotyping study reported for CM cases and controls from MD Anderson Cancer Center.
Figure 2.
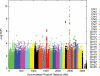
Results from genome-wide association analysis of 818 977 (818 237 + 740 pseudo-autochromosomal) genotypes from 1804 CM cases and 1026 controls, after adjusting for two PCs.
Figure 3.
Association tests for the HERC2/OCA2 region on chromosome 15q13.1. Genotyped SNPs are indicated as diamonds and imputed SNPs as circles. The most significant SNP in the region is HERC2 rs1129038. The strength of the pairwise correlation between the surrounding markers and the most significant SNP is depicted by the size of the symbols: the larger the size, the stronger the LD. Genes in the region are annotated with location, range and orientation.
Figure 4.
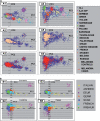
PCA of CM cases and controls showing the relationship among populations of European and Middle-Eastern origin. (A) Analyses of 1804 CM cases and 1026 controls, with an additional 5001 Caucasians of known origin, are shown for PC1 and PC2 (A1, A2, A3), and for PC1 and PC3 (A4, A5, A6). The results are shown with either the known population groups color-coded (A1 and A4) or with controls color-coded (A2 and A5) or cases color-coded (A3 and A6). The population groups are indicated in the legend key. AJA 4GP are Ashkenazi Jewish Americans with four grandparents of Ashkenazi heritage. The group denoted ‘ALL’ are Americans of European ancestry. Results show similarity of PCA loadings for cases and controls in A2–A6. (B) The color code indicates the correspondence of controls or cases to seven different population groups with graphs for PC1 and PC2 (B1 and B3), and PC1 and PC3 (B2 and B4). The population groups were based on correspondence to PCA. The population groups corresponded to the range of the self-identified population group member defined by the first three PCs (see A). Where overlaps were present, the midpoints in the overlapped region were used to define the boundary between the population group assignment. The legend key shows the corresponding population groups with abbreviations for the following groups: Ashkenazi Jewish (ASHKEN), eastern European (EEUR), German (GER), Scandinavian (SCAN) and United Kingdom (UK). Analysis of CM case–control data alone yielded two PCs, whereas analysis including additional European controls yielded the additional third PC, shown in this figure.
Figure 5.
Association tests from four studies and from fixed-effects meta-analysis for a region of chromosome 15 surrounding the HERC2/OCA2 region. The region telomeric to the HERC2 gene contains a region of many complex rearrangements including CHRFAM7A—a rearrangement of nicotinic acetylcholine receptor 7 and FAM7A. The gap in genotyping in the region between 25 168 374 and 25 271 522 base pairs overlays a region with some reported copy number variants in the GABRG3 gene.
Figure 6.
Association tests for a region of chromosome 1 surrounding the LASS2/ANXA9 region. Genotyped SNPs are indicated as diamonds and imputed SNPs as circles. The most significant SNP in the region is rs1722784. The overall structure of the LD with SNPs in this region is reflected by estimated recombination rates. The strength of the pairwise correlation between the surrounding markers and the most significant SNP is depicted by the size of the symbols: the larger the size, the stronger the LD. Genes in the region are annotated with location, range and orientation.
Figure 7.
Association tests from four studies using fixed-effects meta-analysis for a region of chromosome 1 spanning the ARNT/LASS2/ANXA9 region.
References
- Siegel R., Ward E., Brawley O., Jemal A. Cancer statistics, 2011: the impact of eliminating socioeconomic and racial disparities on premature cancer deaths. CA Cancer J. Clin. 2011;61:212–236. -PubMed
- Cust A.E., Schmid H., Maskiell J.A., Jetann J., Ferguson M., Holland E.A., Agha-Hamilton C., Jenkins M.A., Kelly J., Kefford R.F., et al. Population-based, case-control-family design to investigate genetic and environmental influences on melanoma risk: Australian Melanoma Family Study. Am. J. Epidemiol. 2009;170:1541–1554. -PMC -PubMed
- Meyle K.D., Guldberg P. Genetic risk factors for melanoma. Hum. Genet. 2009;126:499–510. -PubMed
Publication types
MeSH terms
Substances
Grants and funding
- P30 CA016672/CA/NCI NIH HHS/United States
- U01 HG004446/HG/NHGRI NIH HHS/United States
- R01 CA137365/CA/NCI NIH HHS/United States
- C8216/A6129/CRUK_/Cancer Research UK/United Kingdom
- P30CA016672/CA/NCI NIH HHS/United States
- R01 CA83115/CA/NCI NIH HHS/United States
- HHSN268200782096C/HG/NHGRI NIH HHS/United States
- C588/A10589/CRUK_/Cancer Research UK/United Kingdom
- HG004446/HG/NHGRI NIH HHS/United States
- ImNIH/Intramural NIH HHS/United States
- R01CA133996/CA/NCI NIH HHS/United States
- R01CA100264/CA/NCI NIH HHS/United States
- R01 CA-83115-01A2/CA/NCI NIH HHS/United States
- R01 CA133996/CA/NCI NIH HHS/United States
- R01 CA083115/CA/NCI NIH HHS/United States
- C588/A4994/CRUK_/Cancer Research UK/United Kingdom
- P50 CA093459/CA/NCI NIH HHS/United States
- 10589/CRUK_/Cancer Research UK/United Kingdom
- R01 CA100264/CA/NCI NIH HHS/United States
- 2P50CA093459/CA/NCI NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical