High-throughput profiling of off-target DNA cleavage reveals RNA-programmed Cas9 nuclease specificity - PubMed (original) (raw)
High-throughput profiling of off-target DNA cleavage reveals RNA-programmed Cas9 nuclease specificity
Vikram Pattanayak et al. Nat Biotechnol. 2013 Sep.
Abstract
The RNA-programmable Cas9 endonuclease cleaves double-stranded DNA at sites complementary to a 20-base-pair guide RNA. The Cas9 system has been used to modify genomes in multiple cells and organisms, demonstrating its potential as a facile genome-engineering tool. We used in vitro selection and high-throughput sequencing to determine the propensity of eight guide-RNA:Cas9 complexes to cleave each of 10(12) potential off-target DNA sequences. The selection results predicted five off-target sites in the human genome that were confirmed to undergo genome cleavage in HEK293T cells upon expression of one of two guide-RNA:Cas9 complexes. In contrast to previous models, our results show that guide-RNA:Cas9 specificity extends past a 7- to 12-base-pair seed sequence. Our results also suggest a tradeoff between activity and specificity both in vitro and in cells as a shorter, less-active guide RNA is more specific than a longer, more-active guide RNA. High concentrations of guide-RNA:Cas9 complexes can cleave off-target sites containing mutations near or within the PAM that are not cleaved when enzyme concentrations are limiting.
Figures
Figure 1. In vitro selection overview
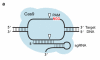
(a) Cas9 complexed with a short guide RNA (sgRNA) recognizes ~20 bases of a target DNA substrate that is complementary to the sgRNA sequence and cleaves both DNA strands. The white triangles represent cleavage locations. (b) A modified version of our previously described in vitro selection was used to comprehensively profile Cas9 specificity. A concatemeric pre-selection DNA library in which each molecule contains one of 10^12 distinct variants of a target DNA sequence (white rectangles) was generated from synthetic DNA oligonucleotides by ligation and rolling-circle amplification. This library was incubated with a Cas9:sgRNA complex of interest. Cleaved library members contain 5′ phosphate groups (green circles) and therefore are substrates for adapter ligation and PCR. The resulting amplicons were subjected to high-throughput DNA sequencing and computational analysis.
Figure 1. In vitro selection overview
(a) Cas9 complexed with a short guide RNA (sgRNA) recognizes ~20 bases of a target DNA substrate that is complementary to the sgRNA sequence and cleaves both DNA strands. The white triangles represent cleavage locations. (b) A modified version of our previously described in vitro selection was used to comprehensively profile Cas9 specificity. A concatemeric pre-selection DNA library in which each molecule contains one of 10^12 distinct variants of a target DNA sequence (white rectangles) was generated from synthetic DNA oligonucleotides by ligation and rolling-circle amplification. This library was incubated with a Cas9:sgRNA complex of interest. Cleaved library members contain 5′ phosphate groups (green circles) and therefore are substrates for adapter ligation and PCR. The resulting amplicons were subjected to high-throughput DNA sequencing and computational analysis.
Figure 2. In vitro selection results for Cas9:CLTA1 sgRNA
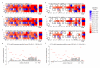
Heat maps show the specificity profiles of Cas9:CLTA1 sgRNA v2.1 under enzyme-limiting conditions (a, b), Cas9:CLTA1 sgRNA v1.0 under enzyme-excess conditions (c, d), and Cas9:CLTA1 sgRNA v2.1 under enzyme-excess conditions (e, f). Heat maps show all post-selection sequences (a, c, e) or only those sequences containing a single mutation in the 20-base pair sgRNA-specified target site and two-base pair PAM (b, d, f). Specificity scores of 1.0 (dark blue) and -1.0 (dark red) corresponds to 100% enrichment for and against, respectively, a particular base pair at a particular position. Black boxes denote the intended target nucleotides. (g) Effect of Cas9:sgRNA concentration on specificity. Positional specificity changes between enzyme-limiting (200 nM DNA, 100 nM Cas9:sgRNA v2.1) and enzyme-excess (200 nM DNA, 1000 nM Cas9:sgRNA v2.1) conditions are shown for CLTA1. Red lines indicate the maximum possible change in positional specificity for a given position. (h) Effect of sgRNA architecture on specificity. Positional specificity changes between sgRNA v1.0 and sgRNA v2.1 under enzyme-excess conditions are shown for CLTA1. Red lines indicate the maximum possible change in positional specificity for a given position. See Supplementary Figures S4-S6, S23, and S24 for corresponding data for CLTA2, CLTA3, and CLTA4.
Comment in
- Staying on target with CRISPR-Cas.
Carroll D. Carroll D. Nat Biotechnol. 2013 Sep;31(9):807-9. doi: 10.1038/nbt.2684. Nat Biotechnol. 2013. PMID: 24022156 No abstract available.
References
Publication types
MeSH terms
Substances
Grants and funding
- HHMI/Howard Hughes Medical Institute/United States
- T32 GM007753/GM/NIGMS NIH HHS/United States
- R01GM073794-05/GM/NIGMS NIH HHS/United States
- R01 GM073794/GM/NIGMS NIH HHS/United States
- T32GM007753/GM/NIGMS NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous