De novo insertions and deletions of predominantly paternal origin are associated with autism spectrum disorder - PubMed (original) (raw)
doi: 10.1016/j.celrep.2014.08.068. Epub 2014 Oct 2.
Michael F Walker 2, Nicholas J Carriero 3, Michael DiCola 4, A Jeremy Willsey 5, Adam Y Ye 6, Zainulabedin Waqar 7, Luis E Gonzalez 7, John D Overton 8, Stephanie Frahm 4, John F Keaney 3rd 9, Nicole A Teran 7, Jeanselle Dea 2, Jeffrey D Mandell 2, Vanessa Hus Bal 2, Catherine A Sullivan 7, Nicholas M DiLullo 7, Rehab O Khalil 10, Jake Gockley 11, Zafer Yuksel 12, Sinem M Sertel 13, A Gulhan Ercan-Sencicek 14, Abha R Gupta 15, Shrikant M Mane 16, Michael Sheldon 17, Andrew I Brooks 4, Kathryn Roeder 18, Bernie Devlin 19, Matthew W State 20, Liping Wei 21, Stephan J Sanders 22
Affiliations
- PMID: 25284784
- PMCID: PMC4194132
- DOI: 10.1016/j.celrep.2014.08.068
De novo insertions and deletions of predominantly paternal origin are associated with autism spectrum disorder
Shan Dong et al. Cell Rep. 2014.
Abstract
Whole-exome sequencing (WES) studies have demonstrated the contribution of de novo loss-of-function single-nucleotide variants (SNVs) to autism spectrum disorder (ASD). However, challenges in the reliable detection of de novo insertions and deletions (indels) have limited inclusion of these variants in prior analyses. By applying a robust indel detection method to WES data from 787 ASD families (2,963 individuals), we demonstrate that de novo frameshift indels contribute to ASD risk (OR = 1.6; 95% CI = 1.0-2.7; p = 0.03), are more common in female probands (p = 0.02), are enriched among genes encoding FMRP targets (p = 6 × 10(-9)), and arise predominantly on the paternal chromosome (p < 0.001). On the basis of mutation rates in probands versus unaffected siblings, we conclude that de novo frameshift indels contribute to risk in approximately 3% of individuals with ASD. Finally, by observing clustering of mutations in unrelated probands, we uncover two ASD-associated genes: KMT2E (MLL5), a chromatin regulator, and RIMS1, a regulator of synaptic vesicle release.
Copyright © 2014 The Authors. Published by Elsevier Inc. All rights reserved.
Figures
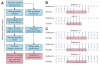
Figure 1. Experimental overview
A) Indels were predicted in 787 families from the SSC using Dindel. Throughout the analytical pipeline, probands and siblings are treated equally to allow accurate assessment of de novo indel burden. Informative SNPs were used to establish the parent-of-origin of de novo indels. B) Alignment errors at the end of reads lead to indels being mis-called as SNVs. C) An indel can be represented in multiple ways in VCF format. See also Table S1.
Figure 2. De novo indel burden and genes with multiple hits
(A) The rate of de novo indels and SNVs is shown for 602 probands (red) and matched unaffected siblings (blue). “All” refers to all RefSeq genes in hg19. “Brain” refers to the subset of genes that are brain-expressed. “Nonsense” refers to single nucleotide substitutions that result in a premature stop codon; “splice-site” refers to single nucleotide substitutions that disrupt the canonical splice-site. Error bars represent the 95% confidence intervals and p-values are calculated with a one-sided paired Wilcoxon test. (B) Two de novo frameshift indels in independent samples are shown in the gene KMT2E. Both indels are likely to induce nonsense-mediated decay (Nagy and Maquat, 1998). (C) Two de novo frameshift indels in independent samples are shown in the gene RIMS1. Both indels are likely to induce nonsense-mediated decay (Nagy and Maquat, 1998). See also Figures S1 and S2.
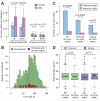
Figure 3. Sex difference, parent-of-origin and parental age
(A) A consistently higher rate of de novo frameshift indels was observed in female probands (pink) compared to male probands (blue), but this difference was not observed in unaffected siblings. “All” describes all de novo frameshift indels; “Brain” includes only those expressed in the brain. Error bars represent the 95% confidence intervals and p-values are calculated with a one-sided paired Wilcoxon test. (B) Histogram of full-scale IQ in all probands (green) and probands with a de novo frameshift indel (red). (C) The majority of de novo indels for which parent-of-origin could be resolved were found to be on the paternal (blue) rather than the maternal (pink) chromosome (p<0.001; Binomial). This result was observed in both probands and siblings separately. (D) No clear relationship between the presence of a de novo indel and increased paternal age was observed for probands (green) or siblings (purple). P-values were estimated with a Poisson regression.
References
Publication types
MeSH terms
Substances
Grants and funding
- R37 MH057881/MH/NIMH NIH HHS/United States
- KL2 TR000140/TR/NCATS NIH HHS/United States
- T32 HD007149/HD/NICHD NIH HHS/United States
- CAPMC/ CIHR/Canada
- HHMI/Howard Hughes Medical Institute/United States
- U01 MH100239/MH/NIMH NIH HHS/United States
- UL1 TR000142/TR/NCATS NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources