Distinct oncogenic Ras signals characterized by profound differences in flux through the RasGDP/RasGTP cycle - PubMed (original) (raw)
Comment
Distinct oncogenic Ras signals characterized by profound differences in flux through the RasGDP/RasGTP cycle
Marsilius Mues et al. Small GTPases. 2017.
Abstract
T cell acute lymphoblastic leukemia/lymphoma (T-ALL) is an aggressive bone marrow cancer in children and adults, and chemotherapy often fails for relapsing patients. Molecularly targeted therapy is hindered by heterogeneity in T-ALL and mechanistic details of the affected pathways in T-ALL are needed. Deregulation of Ras signals is common in T-ALL. Ras is genetically mutated to a constitutively active form in about 15% of all haematopoietic malignancies, but there is a range of other ways to augment signaling through the Ras pathway. Several groups including our own uncovered that RasGRP1 overexpression leads to T-ALL in mouse models and in pediatric T-ALL patients, and we reported that this Ras guanine nucleotide exchange factor, RasGRP1, cooperates with cytokines to drive leukemogenesis. In our recent study by Ksionda et al. we analyzed the molecular details of cytokine receptor-RasGRP1-Ras signals in T-ALL and compared these to signals from mutated Ras alleles, which yielded several surprising results. Examples are the striking differences in flux through the RasGDP/RasGTP cycle in distinct T-ALL or unexpected differences in wiring of the Ras signaling pathway between T-ALL and normal developing T cells, which we will discuss here.
Keywords: PI3K; Ras; RasGAP; RasGRP1; cancer; heterogeneity; leukemia; mouse models; patients; therapy.
Figures
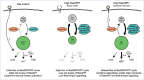
Figure 1.
Schematic comparison of different oncogenic Ras signaling in RasMUT and RasGRP1-overexpressing T-ALL. Left: Leukemic cells with constitutively active Ras display high net levels of active RasGTP and very low nucleotide exchange rates due to the defective GTPase function of Ras. Middle: RasGRP1-overexpressing T-ALL show high levels of flux through the RasGDP/GTP cycle, yet low net levels of RasGTP, presumably due to the high compensatory activity of RasGAP family members. Right: After cytokine activation, RasGRP1-overexpressing T-ALL reveal elevated RasGTP levels and active downstream signaling, primarily through the PI3K-Akt pathway.
Comment on
- RasGRP1 overexpression in T-ALL increases basal nucleotide exchange on Ras rendering the Ras/PI3K/Akt pathway responsive to protumorigenic cytokines.
Ksionda O, Melton AA, Bache J, Tenhagen M, Bakker J, Harvey R, Winter SS, Rubio I, Roose JP. Ksionda O, et al. Oncogene. 2016 Jul 14;35(28):3658-68. doi: 10.1038/onc.2015.431. Epub 2015 Nov 9. Oncogene. 2016. PMID: 26549032 Free PMC article.
References
- Zhang J, Ding L, Holmfeldt L, Wu G, Heatley SL, Payne-Turner D, Easton J, Chen X, Wang J, Rusch M, et al.. The genetic basis of early T-cell precursor acute lymphoblastic leukaemia. Nature 2012; 481:157-63; PMID:22237106; http://dx.doi.org/ 10.1038/nature10725 -DOI -PMC -PubMed
- Pui CH, Relling MV, Downing JR. Acute lymphoblastic leukemia. N Engl J Med 2004; 350:1535-48; PMID:15071128; http://dx.doi.org/ 10.1056/NEJMra023001 -DOI -PubMed
- Pui CH, Evans WE. Treatment of acute lymphoblastic leukemia. N Engl J Med 2006; 354:166-78; PMID:16407512; http://dx.doi.org/ 10.1056/NEJMra052603 -DOI -PubMed
- Thiel E, Kranz BR, Raghavachar A, Bartram CR, Löffler H, Messerer D, Ganser A, Ludwig WD, Büchner T, Hoelzer D. Prethymic phenotype and genotype of pre-T (CD7+/ER-)-cell leukemia and its clinical significance within adult acute lymphoblastic leukemia. Blood 1989; 73:1247-58; PMID:2467704 -PubMed
- Pullen J, Shuster JJ, Link M, Borowitz M, Amylon M, Carroll AJ, Land V, Look AT, McIntyre B, Camitta B. Significance of commonly used prognostic factors differs for children with T cell acute lymphocytic leukemia (ALL), as compared to those with B-precursor ALL. A Pediatric Oncology Group (POG) study. Leukemia 1999; 13:1696-707; PMID:10557041; http://dx.doi.org/ 10.1038/sj.leu.2401555 -DOI -PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources