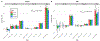Assessing the contribution of rare variants to complex trait heritability from whole-genome sequence data - PubMed (original) (raw)
. 2022 Mar;54(3):263-273.
doi: 10.1038/s41588-021-00997-7. Epub 2022 Mar 7.
Deepti Jain 2, Zhili Zheng 3; TOPMed Anthropometry Working Group; NHLBI Trans-Omics for Precision Medicine (TOPMed) Consortium; L Adrienne Cupples 4 5, Aladdin H Shadyab 6, Barbara McKnight 2, Benjamin M Shoemaker 7, Braxton D Mitchell 8 9, Bruce M Psaty 10, Charles Kooperberg 11, Ching-Ti Liu 4, Christine M Albert 12 13 14, Dan Roden 15, Daniel I Chasman 14, Dawood Darbar 16, Donald M Lloyd-Jones 17, Donna K Arnett 18, Elizabeth A Regan 19, Eric Boerwinkle 20, Jerome I Rotter 21, Jeffrey R O'Connell 8, Lisa R Yanek 22, Mariza de Andrade 23, Matthew A Allison 24, Merry-Lynn N McDonald 25, Mina K Chung 26, Myriam Fornage 27, Nathalie Chami 28 29, Nicholas L Smith 30 31 32, Patrick T Ellinor 12 33, Ramachandran S Vasan 5 34 35, Rasika A Mathias 36, Ruth J F Loos 28 29, Stephen S Rich 37, Steven A Lubitz 33 38, Susan R Heckbert 31 32, Susan Redline 39 40 41, Xiuqing Guo 21, Y -D Ida Chen 21, Cecelia A Laurie 2, Ryan D Hernandez 42 43, Stephen T McGarvey 44, Michael E Goddard 45 46, Cathy C Laurie 2, Kari E North 47, Leslie A Lange 48, Bruce S Weir 2, Loic Yengo 3, Jian Yang # 49 50, Peter M Visscher # 51 52
Collaborators, Affiliations
- PMID: 35256806
- PMCID: PMC9119698
- DOI: 10.1038/s41588-021-00997-7
Assessing the contribution of rare variants to complex trait heritability from whole-genome sequence data
Pierrick Wainschtein et al. Nat Genet. 2022 Mar.
Abstract
Analyses of data from genome-wide association studies on unrelated individuals have shown that, for human traits and diseases, approximately one-third to two-thirds of heritability is captured by common SNPs. However, it is not known whether the remaining heritability is due to the imperfect tagging of causal variants by common SNPs, in particular whether the causal variants are rare, or whether it is overestimated due to bias in inference from pedigree data. Here we estimated heritability for height and body mass index (BMI) from whole-genome sequence data on 25,465 unrelated individuals of European ancestry. The estimated heritability was 0.68 (standard error 0.10) for height and 0.30 (standard error 0.10) for body mass index. Low minor allele frequency variants in low linkage disequilibrium (LD) with neighboring variants were enriched for heritability, to a greater extent for protein-altering variants, consistent with negative selection. Our results imply that rare variants, in particular those in regions of low linkage disequilibrium, are a major source of the still missing heritability of complex traits and disease.
© 2022. The Author(s), under exclusive licence to Springer Nature America, Inc.
Conflict of interest statement
The remaining authors declare no competing interests.
Figures
Figure 1:
GREML-LDMS estimates with 8 bins (2 LD bins for each of the 4 MAF bins) correcting for 20 PCs (calculated from LD-pruned HM3 SNPs) after imputing SNPs from Illumina InfiniumCore24, GSA 24 and Affymetrix Axiom arrays using Haplotype Reference Consortium reference panels for N=25,465 samples. (A) Estimates of hG+IMP2 for height are between 0.50-0.56 (SE 0.06-0.07). (B) Estimates for BMI are between 0.16-0.21 (SE 0.07). The large SEs of the estimates for variants with MAF between 0.0001 to 0.001 can be explained by the large number of imputed variants in this MAF bin because the sampling variance of a SNP-based heritability estimate is proportional to the effective number of independent variants . Between ~19.0M and ~20.0M variants in total are included in the analysis. The number of variants in each of the 4 MAF bins (twice the number in each LD bin) can be found in Supplementary Figure 8.
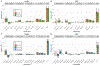
Figure 2:
GREML-LDMS of height and BMI for N=25,465 samples using 3 or 4 LD groups for each MAF bin correcting for 48/160/320 PCs computed from WGS variants. Each variant was allocated in a tertile or a quartile according to its LD score. (A) Estimates using 3 LD bins for height: 0.68 (SE 0.09 – 0.10). (B) Estimates using 3 LD bins for BMI: 0.30 – 0.32 (SE 0.10). (C) Estimates using 4 LD bins for height: 0.67 – 0.68 (SE 0.10). (D) Estimates using 4 LD bins for BMI: 0.28 – 0.30 (SE 0.10).
Figure 3:
Variance explained per variant (the estimate of genetic variance divided by the number of variants in each bin) from GREML-LDMS with the low-LD and low-MAF (< 0.1) variants partitioned into 2 distinct categories according to the SnpEff putative effect of the variant (protein-altering or non-protein-altering), correcting for 48 PCs from WGS variants for N=25,465 samples. There is a total of 11 genetic components in this analysis. There is an apparent enrichment of heritability in the protein-altering groupings (low LD) over non-protein-altering (low LD) or high LD variants for height (A) as well as for BMI, although the standard errors for this trait are large (B).
Comment in
- Discovering missing heritability in whole-genome sequencing data.
Young AI. Young AI. Nat Genet. 2022 Mar;54(3):224-226. doi: 10.1038/s41588-022-01012-3. Nat Genet. 2022. PMID: 35273401 No abstract available.
References
- Lynch M & Walsh B Genetics and analysis of quantitative traits. (Sinauer, 1998).
- Fisher RA XV.—The Correlation between Relatives on the Supposition of Mendelian Inheritance. Transactions of the Royal Society of Edinburgh 52, 399–433, doi: 10.1017/s0080456800012163 (1918). -DOI
Methods References
Publication types
MeSH terms
Grants and funding
- N01 HC085080/HL/NHLBI NIH HHS/United States
- P30 ES010126/ES/NIEHS NIH HHS/United States
- K01 AG059898/AG/NIA NIH HHS/United States
- U01 HL080295/HL/NHLBI NIH HHS/United States
- HHSN268200800007C/HL/NHLBI NIH HHS/United States
- R01 DK110113/DK/NIDDK NIH HHS/United States
- IK6 BX006185/BX/BLRD VA/United States
- R35 HL135818/HL/NHLBI NIH HHS/United States
- K08 HL141601/HL/NHLBI NIH HHS/United States
- R01 HL111314/HL/NHLBI NIH HHS/United States
- N01 HC085079/HL/NHLBI NIH HHS/United States
- R01 GM075091/GM/NIGMS NIH HHS/United States
- R01 DK124097/DK/NIDDK NIH HHS/United States
- R01 HL158071/HL/NHLBI NIH HHS/United States
- R01 AG023629/AG/NIA NIH HHS/United States
- U54 GM115428/GM/NIGMS NIH HHS/United States
- N01 HC085081/HL/NHLBI NIH HHS/United States
- R01 HL059367/HL/NHLBI NIH HHS/United States
- U01 HL130114/HL/NHLBI NIH HHS/United States
- HHSN268201200036C/HL/NHLBI NIH HHS/United States
- HHSN268201800001C/HL/NHLBI NIH HHS/United States
- R01 MH100141/MH/NIMH NIH HHS/United States
- T32 CA154274/CA/NCI NIH HHS/United States
LinkOut - more resources
Full Text Sources
Research Materials