Genomic insights into rapid speciation within the world's largest tree genus Syzygium - PubMed (original) (raw)
. 2022 Sep 12;13(1):5031.
doi: 10.1038/s41467-022-32637-x.
Yee Wen Low # 1 2 3, Crystal M Tomlin # 6, Joffre Ali Ahmad 7, Wisnu H Ardi 8, Kate Armstrong 9, Parusuraman Athen 10, Ahmad Berhaman 11, Ruth E Bone 12, Martin Cheek 12, Nicholas R W Cho 4, Le Min Choo 10, Ian D Cowie 13, Darren Crayn 14, Steven J Fleck 6, Andrew J Ford 15, Paul I Forster 16, Deden Girmansyah 17, David J Goyder 12, Bruce Gray 14, Charlie D Heatubun 12 18 19, Ali Ibrahim 10, Bazilah Ibrahim 10, Himesh D Jayasinghe 20 21, Muhammad Ariffin Kalat 7, Hashendra S Kathriarachchi 20, Endang Kintamani 17, Sin Lan Koh 10, Joseph T K Lai 22, Serena M L Lee 10, Paul K F Leong 10, Wei Hao Lim 10, Shawn K Y Lum 23, Ridha Mahyuni 17, William J F McDonald 16, Faizah Metali 24, Wendy A Mustaqim 25, Akiyo Naiki 26, Kang Min Ngo 23, Matti Niissalo 10, Subhani Ranasinghe 27, Rimi Repin 28, Himmah Rustiami 17, Victor I Simbiak 19, Rahayu S Sukri 24, Siti Sunarti 17, Liam A Trethowan 12, Anna Trias-Blasi 12, Thais N C Vasconcelos 12 29, Jimmy F Wanma 19, Pudji Widodo 30, Douglas Siril A Wijesundara 21, Stuart Worboys 14, Jing Wei Yap 31, Kien Thai Yong 32, Gillian S W Khew 10 4, Jarkko Salojärvi 4 5, Todd P Michael 33, David J Middleton 10, David F R P Burslem 34, Charlotte Lindqvist 35 36, Eve J Lucas 37, Victor A Albert 38 39
Affiliations
- PMID: 36097018
- PMCID: PMC9468008
- DOI: 10.1038/s41467-022-32637-x
Genomic insights into rapid speciation within the world's largest tree genus Syzygium
Yee Wen Low et al. Nat Commun. 2022.
Abstract
Species radiations, despite immense phenotypic variation, can be difficult to resolve phylogenetically when genetic change poorly matches the rapidity of diversification. Genomic potential furnished by palaeopolyploidy, and relative roles for adaptation, random drift and hybridisation in the apportionment of genetic variation, remain poorly understood factors. Here, we study these aspects in a model radiation, Syzygium, the most species-rich tree genus worldwide. Genomes of 182 distinct species and 58 unidentified taxa are compared against a chromosome-level reference genome of the sea apple, Syzygium grande. We show that while Syzygium shares an ancient genome doubling event with other Myrtales, little evidence exists for recent polyploidy events. Phylogenomics confirms that Syzygium originated in Australia-New Guinea and diversified in multiple migrations, eastward to the Pacific and westward to India and Africa, in bursts of speciation visible as poorly resolved branches on phylogenies. Furthermore, some sublineages demonstrate genomic clines that recapitulate cladogenetic events, suggesting that stepwise geographic speciation, a neutral process, has been important in Syzygium diversification.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures
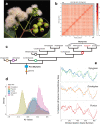
Fig. 1. Assembly and structural evolution of the Syzygium grande reference genome.
a S. grande inflorescence, flowers and fruits; the latter evoke the common name “sea apple”; b HiC contact map for the scaffolded genome, showing 11 assembled chromosomes; c Phylogeny of major lineages of Myrtales, following Maurin et al. . Genera of Myrtaceae used in genome structural and phylogenetic analyses are also depicted. Punica (Lythraceae) was also examined for structural evolution. Open circles represent the multiple, independent polyploidy events predicted by the 1KP study; our results here suggest instead a single Pan-Myrtales whole genome duplication (blue rectangle) which followed the gamma hexaploidy (orange triangle) present in all core eudicots. d Synonymous substitution rate density plots for internal polyploid paralogs within Syzygium, Eucalyptus, Punica, Populus and Vitis. Modal peaks in these three Myrtales species suggest a single underlying polyploidy event. Ks asymmetries were calibrated using the gamma event present in each species. Both histograms and smoothed curves are shown. e Fractionation bias mappings of Myrtales chromosomal scaffolds, 2 each (different colours), onto Vitis vinifera chromosome 2 show similar patterns for all three Myrtales species (excluding cases of chromosomal rearrangements among the three, which are discernible as different scaffold colour switchings compared to the Vitis chromosome). _X_-axis shows the percent retention of fractionated gene pairs following polyploidisation; _Y_-axis shows the position of gene pairs along the Vitis chromosome. Photograph credit: WHL (a). Source data are provided as a Source Data file.
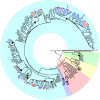
Fig. 2. Phylogenetic tree based on single nucleotide polymorphisms among all 292 resequenced Myrtaceae accessions.
Black circles represent the at-least 12 independent invasions of Sunda from Sahul. The green circle represents migration from Sunda to the Indian subcontinent, and the purple circle denotes further migration from there to Africa. Blue versus red circles at the leaves of the tree represent Syzygium accessions from Bukit Timah Nature Reserve and Danum Valley Conservation area, respectively. Background colours represent the recognised subgenera, (clockwise from the root, excluding the outgroup taxa) Syzygium subg. Sequestratum (green), S. subg. Perikion (yellow), S. subg. Acmena (red), the S. rugosum clade (purple) and S. subg. Syzygium (cyan).
Fig. 3. Principal component analysis and phylogenetic reconstructions of single nucleotide polymorphism variation within Syzygium.
a A NeighborNet phylogenetic network shows considerable character discordance among genome-wide SNPs that may be indicative of incomplete lineage sorting. This discordance is particularly noteworthy at the highly webbed base of the Syzygium grande group (close-up view in square inset; see also the labelled network in Supplementary Fig. 89). b PCA of principal components 1 and 2 of Syzygium subg. Syzygium individuals. Clinal patterns are readily observed; the Syzygium grande group (centred around 0.10 on PC2) is comprised of a medium-blue paraphyletic grade subtending a pink terminal lineage, as shown in (c), the RAxML SNP tree, which is colour-coded following the PCA. Shading of two groups on (a) matches the colour coding on the tree as well as the colours and symbols on the PCA plot. Small matching symbols in shaded areas are shown for clarity. Edge with two horizontal lines at tip represents the outgroup taxon, Syzygium rugosum, clipped for length. Source data are provided as a Source Data file.
Fig. 4. Genomic palaeodemography of Syzygium accessions from the S. grande group.
Pairwise Sequentially Markovian Coalescent analyses, coloured by groups/clades in Fig. 3. Source data are provided as a Source Data file.
Fig. 5. Reproductive trait diversity in the genus Syzygium, as examined to reconstruct ancestral states using Mesquite.
a Free petals (Syzygium pendens); b Calyptrate calyx (Syzygium paradoxum); c Pseudocalyptrate corolla (Syzygium adelphicum); d Fruits maturing green (Syzygium cf. dyerianum); e Pendulous inflorescence or infructescence (Syzygium boonjee). Photograph credits: YWL (a)–(e).
References
- Gittenberger E. What about non-adaptive radiation? Biol. J. Linn. Soc. 1991;43:263–272. doi: 10.1111/j.1095-8312.1991.tb00598.x. -DOI
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources