The genetic determinants of recurrent somatic mutations in 43,693 blood genomes - PubMed (original) (raw)
. 2023 Apr 28;9(17):eabm4945.
doi: 10.1126/sciadv.abm4945. Epub 2023 Apr 26.
Cecelia A Laurie 2, Jai G Broome 2 3, Kent D Taylor 4, Xiuqing Guo 4, Alan R Shuldiner 5, Jeffrey R O'Connell 5, Joshua P Lewis 5, Eric Boerwinkle 6, Kathleen C Barnes 7, Nathalie Chami 8 9, Eimear E Kenny 10, Ruth J F Loos 8 9, Myriam Fornage 11, Susan Redline 12 13, Brian E Cade 12 13 14, Frank D Gilliland 15, Zhanghua Chen 15, W James Gauderman 15, Rajesh Kumar 16 17, Leslie Grammer 17, Robert P Schleimer 17, Bruce M Psaty 18 19 20, Joshua C Bis 18, Jennifer A Brody 18, Edwin K Silverman 21, Jeong H Yun 21, Dandi Qiao 21, Scott T Weiss 12 21, Jessica Lasky-Su 12 21, Dawn L DeMeo 12 21, Nicholette D Palmer 22, Barry I Freedman 23, Donald W Bowden 22, Michael H Cho 24, Ramachandran S Vasan 25, Andrew D Johnson 25 26, Lisa R Yanek 27, Lewis C Becker 27, Sharon Kardia 28, Jiang He 29, Robert Kaplan 30, Susan R Heckbert 19 31, Nicholas L Smith 19 31 32, Kerri L Wiggins 33, Donna K Arnett 34, Marguerite R Irvin 35, Hemant Tiwari 35, Adolfo Correa 36, Laura M Raffield 37, Yan Gao 38, Mariza de Andrade 39, Jerome I Rotter 4, Stephen S Rich 40, Ani W Manichaikul 40, Barbara A Konkle 20, Jill M Johnsen 20 41, Marsha M Wheeler 42, Brian S Custer 43, Ravindranath Duggirala 44 45, Joanne E Curran 44 45, John Blangero 44 45, Hongsheng Gui 46 47, Shujie Xiao 46 47, L Keoki Williams 46 47, Deborah A Meyers 48, Xingnan Li 49, Victor Ortega 50, Stephen McGarvey 51, C Charles Gu 52, Yii-Der Ida Chen 4, Wen-Jane Lee 53, M Benjamin Shoemaker 54, Dawood Darbar 55, Dan Roden 56, Christine Albert 57, Charles Kooperberg 58, Pinkal Desai 59 60, Thomas W Blackwell 1, Goncalo R Abecasis 1 61, Albert V Smith 1, Hyun M Kang 1, Rasika Mathias 27, Pradeep Natarajan 14 62 63, Siddhartha Jaiswal 64, Alexander P Reiner 58 19, Alexander G Bick 65; NHLBI Trans-Omics for Precision Medicine (TOPMed) Consortium
Affiliations
- PMID: 37126548
- PMCID: PMC10132750
- DOI: 10.1126/sciadv.abm4945
The genetic determinants of recurrent somatic mutations in 43,693 blood genomes
Joshua S Weinstock et al. Sci Adv. 2023.
Abstract
Nononcogenic somatic mutations are thought to be uncommon and inconsequential. To test this, we analyzed 43,693 National Heart, Lung and Blood Institute Trans-Omics for Precision Medicine blood whole genomes from 37 cohorts and identified 7131 non-missense somatic mutations that are recurrently mutated in at least 50 individuals. These recurrent non-missense somatic mutations (RNMSMs) are not clearly explained by other clonal phenomena such as clonal hematopoiesis. RNMSM prevalence increased with age, with an average 50-year-old having 27 RNMSMs. Inherited germline variation associated with RNMSM acquisition. These variants were found in genes involved in adaptive immune function, proinflammatory cytokine production, and lymphoid lineage commitment. In addition, the presence of eight specific RNMSMs associated with blood cell traits at effect sizes comparable to Mendelian genetic mutations. Overall, we found that somatic mutations in blood are an unexpectedly common phenomenon with ancestry-specific determinants and human health consequences.
Figures
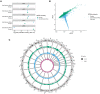
Fig. 1.. Identification of recurrent somatic mutations from TOPMed whole genomes.
(A) Identification of RNMSMs. (B) The allele frequency spectrum of somatic mutations. These represent the frequency of sets of variants with the given allele count. (C) The average number of RNMSMs stratified across age bins with 95% CIs. (D) The histogram of the slopes of the individual RNMSMs with age. (E) Enrichments of chromHMM annotations across blood epigenomes from Roadmap Epigenomics. Enrichments are defined as the log odds ratio (OR) multiplied by the −log10 (P value) from the Fisher’s exact test. Treg, regulatory T cell; TH, T helper cell; IL-17pp, interleukin-17pp; PMA, phorbol 12-myristate 13-acetate; MACS, magnetic-activated cell sorting; Tmem, T memory.
Fig. 2.. Somatic genotype PCs are associated with genetic ancestry.
(A) Global ancestry labels, as estimated from RFMix, are regressed on the first five sPCs in a multinomial regression with a study indicator included as a covariate. The ORs are estimated with East Asian ancestry as the reference level. (B) A scatterplot of sPC4 and sPC5, with colors indicated by the RFMix global ancestry label. (C) A circular genome plot where the angle indicates genomic position, and the radius within a given track indicates the allele frequency, which ranges from 0 to 0.12. A separate track is plotted for the allele frequencies computed separately in European, Sub-Saharan African, and East Asian genomes, respectively, which are colored by the legend indicated in (B). The size of points indicates the fixation index (Fst) estimate, where larger points have larger Fst values.
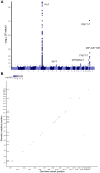
Fig. 3.. Germline determinants of RNMSM burden.
(A) Manhattan plot from GWAS of RNMSM burden, computed using SAIGE. Germline variants included had a minor allele count ≥ 600 and were distinct from the set of RNMSMs. (B) Genetic determinants of individual RNMSMs are primarily in “cis,” where cis is defined as within 2 Mb of the RNMSM. Genomic coordinates of the associated linkage disequilibrium (LD)-clumped germline variants are plotted on the x axis, and coordinates of the RNMSMs are plotted on the y axis. Points falling along the diagonal indicate cis associations. Tick marks along the y axis indicate positions of the RNMSMs from the LD-clumped associations.
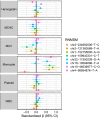
Fig. 4.. RNMSMs associate with blood cell traits.
Associations between eight RNMSMs (q values < 0.05) with harmonized blood cell traits in 2996 individuals. Units correspond to an increase of an SD of the phenotype associated with an increase in the VAF value of an RNMSM by 0.10. WBC, white blood cell; MCHC, mean corpuscular hemoglobin concentration; MCV, mean corpuscular volume.
References
- K. Yizhak, F. Aguet, J. Kim, J. M. Hess, K. Kübler, J. Grimsby, R. Frazer, H. Zhang, N. J. Haradhvala, D. Rosebrock, D. Livitz, X. Li, E. Arich-Landkof, N. Shoresh, C. Stewart, A. V. Segrè, P. A. Branton, P. Polak, K. G. Ardlie, G. Getz, RNA sequence analysis reveals macroscopic somatic clonal expansion across normal tissues. Science 364, eaaw0726 (2019). -PMC -PubMed
- S. Abelson, G. Collord, S. W. K. Ng, O. Weissbrod, N. Mendelson Cohen, E. Niemeyer, N. Barda, P. C. Zuzarte, L. Heisler, Y. Sundaravadanam, R. Luben, S. Hayat, T. T. Wang, Z. Zhao, I. Cirlan, T. J. Pugh, D. Soave, K. Ng, C. Latimer, C. Hardy, K. Raine, D. Jones, D. Hoult, A. Britten, J. D. McPherson, M. Johansson, F. Mbabaali, J. Eagles, J. K. Miller, D. Pasternack, L. Timms, P. Krzyzanowski, P. Awadalla, R. Costa, E. Segal, S. V. Bratman, P. Beer, S. Behjati, I. Martincorena, J. C. Y. Wang, K. M. Bowles, J. R. Quirós, A. Karakatsani, C. La Vecchia, A. Trichopoulou, E. Salamanca-Fernández, J. M. Huerta, A. Barricarte, R. C. Travis, R. Tumino, G. Masala, H. Boeing, S. Panico, R. Kaaks, A. Krämer, S. Sieri, E. Riboli, P. Vineis, M. Foll, J. McKay, S. Polidoro, N. Sala, K.-T. Khaw, R. Vermeulen, P. J. Campbell, E. Papaemmanuil, M. D. Minden, A. Tanay, R. D. Balicer, N. J. Wareham, M. Gerstung, J. E. Dick, P. Brennan, G. S. Vassiliou, L. I. Shlush, Prediction of acute myeloid leukaemia risk in healthy individuals. Nature 559, 400–404 (2018). -PMC -PubMed
- S. Jaiswal, P. Fontanillas, J. Flannick, A. Manning, P. V. Grauman, B. G. Mar, R. Coleman Lindsley, C. H. Mermel, N. Burtt, A. Chavez, J. M. Higgins, V. Moltchanov, F. C. Kuo, M. J. Kluk, B. Henderson, L. Kinnunen, H. A. Koistinen, C. Ladenvall, G. Getz, A. Correa, B. F. Banahan, S. Gabriel, S. Kathiresan, H. M. Stringham, M. I. McCarthy, M. Boehnke, J. Tuomilehto, C. Haiman, L. Groop, G. Atzmon, J. G. Wilson, D. Neuberg, D. Altshuler, B. L. Ebert, Age-related clonal hematopoiesis associated with adverse outcomes. N. Engl. J. Med. 26, 2488–2998 (2014). -PMC -PubMed
- G. Genovese, A. K. Kähler, R. E. Handsaker, J. Lindberg, S. A. Rose, S. F. Bakhoum, K. Chambert, E. Mick, B. M. Neale, M. Fromer, S. M. Purcell, O. Svantesson, M. Landén, M. Höglund, S. Lehmann, S. B. Gabriel, J. L. Moran, E. S. Lander, P. F. Sullivan, P. Sklar, H. Grönberg, C. M. Hultman, S. A. McCarroll, Clonal hematopoiesis and blood-cancer risk inferred from blood DNA sequence. N. Engl. J. Med. 371, 2477–2487 (2014). -PMC -PubMed
MeSH terms
Grants and funding
- R01 HL113323/HL/NHLBI NIH HHS/United States
- HHSN268201800001C/HL/NHLBI NIH HHS/United States
- R01 AI132476/AI/NIAID NIH HHS/United States
- DP2 HL157540/HL/NHLBI NIH HHS/United States
- P01 HL132825/HL/NHLBI NIH HHS/United States
- DP5 OD029586/OD/NIH HHS/United States
- R01 HL120393/HL/NHLBI NIH HHS/United States
- U01 HL120393/HL/NHLBI NIH HHS/United States
- R01 HL153805/HL/NHLBI NIH HHS/United States
- R01 HL117626/HL/NHLBI NIH HHS/United States
- K08 HL146972/HL/NHLBI NIH HHS/United States
- P01 CA196569/CA/NCI NIH HHS/United States