Significance of Heparanase in Cancer and Inflammation (original) (raw)
Abstract
Heparan sulfate proteoglycans (HSPGs) are primary components at the interface between virtually every eukaryotic cell and its extracellular matrix. HSPGs not only provide a storage depot for heparin-binding molecules in the cell microenvironment, but also decisively regulate their accessibility, function and mode of action. As such, they are intimately involved in modulating cell invasion and signaling loops that are critical for tumor growth, inflammation and kidney function. In a series of studies performed since the cloning of the human heparanase gene, we and others have demonstrated that heparanase, the sole heparan sulfate degrading endoglycosidase, is causally involved in cancer progression, inflammation and diabetic nephropathy and hence is a valid target for drug development. Heparanase is causally involved in inflammation and accelerates colon tumorigenesis associated with inflammatory bowel disease. Notably, heparanase stimulates macrophage activation, while macrophages induce production and activation of latent heparanase contributed by the colon epithelium, together generating a vicious cycle that powers colitis and the associated tumorigenesis. Heparanase also plays a decisive role in the pathogenesis of diabetic nephropathy, degrading heparan sulfate in the glomerular basement membrane and ultimately leading to proteinuria and kidney dysfunction. Notably, clinically relevant doses of ionizing radiation (IR) upregulate heparanase expression and thereby augment the metastatic potential of pancreatic carcinoma. Thus, combining radiotherapy with heparanase inhibition is an effective strategy to prevent tumor resistance and dissemination in IR-treated pancreatic cancer patients. Also, accumulating evidence indicate that peptides derived from human heparanase elicit a potent anti-tumor immune response, suggesting that heparanase represents a promising target antigen for immunotherapeutic approaches against a broad variety of tumours. Oligosaccharide-based compounds that inhibit heparanase enzymatic activity were developed, aiming primarily at halting tumor growth, metastasis and angiogenesis. Some of these compounds are being evaluated in clinical trials, targeting both the tumor and tumor microenvironment.
Keywords: Extracellular matrix, Tumor microenvironment, Inflammation, Tumor associated macrophages, Heparan sulfate, Heparanase, Metastasis, Colitis, Colon carcinoma
Preface
The extracellular matrix (ECM) is a heterogeneous mixture of proteins and polysaccharides that surrounds cells, providing physical support for cellular organization into tissues and organs. Traditionally, the ECM was regarded as an inert scaffold providing a structural framework for cells to form tissues and organs. Back in 1979, we were among the first to realize that the ECM plays an active role in orchestrating cellular responses to both normal and pathological situations [1, 2]. The emerging notion was one of active interplay between cells and ECM where the cells synthesize matrix components which in turn dictate and regulate cell shape and function [1, 2]. The ECM network of proteins, glycoproteins and proteoglycans provides adherent cells with structural support and biochemical cues that regulate cell fate and function. We developed a straightforward approach to coat plastic surfaces with ECM deposited by cultured endothelial cells and demonstrated that this naturally produced ECM closely resembles the subendothelial basement membrane in vivo [2, 3]. This ECM and the more commonly used 3-dimensional tumor-derived basement membrane-like substrate (Matrigel) [4] are being applied to sustain cell proliferation, differentiation and survival in vitro_,_ retaining the in vivo characteristics [5]. The ECM/Matrigel system is also widely used to study tumor cell invasion and vascular sprouting. In subsequent studies we have demonstrated that the ECM and basement membrane provide a storage depot for FGF2 and thereby regulates its bioavailability [6]. By now, the function of ECM as a reservoir for bioactive molecules is well recognized and highly important to the current appreciation of the tumor microenvironment and its significance in cancer progression and treatment.
Apart of studies on the biology of the ECM in general, our research has been focused on heparan sulfate (HS) glycosaminoglycan (GAG), one of the most important components of the ECM, basement membranes and cell surface molecules, shown to have a pronounced effect on fundamental biological processes, ranging from development and formation of blood vessels to cell invasion, inflammation and viral infection [7]. GAGs are linear polysaccharides consisting of a repeating disaccharide generally of an acetylated amino sugar alternating with uronic acid. Notably, while 4 and 20 building blocks make nucleic acids and proteins, respectively, 32 disaccharide building blocks make up these complex, highly acidic and information dense biopolymers. The chemical heterogeneity and structural complexity of GAGs make investigations of these molecules most challenging, raising fundamental questions as to how topological positioning and function of cells and tissues are regulated by GAGs. The biosynthesis of HS GAG takes place in the Golgi system and has been studied in great detail. Briefly, the polysaccharide chains are modified at various positions by sulfation, epimerization and N-acetylation, yielding clusters of sulfated disaccharides separated by low or non-sulfated regions [8, 9]. The sulfated saccharide domains provide numerous docking sites for a multitude of protein ligands, ensuring that a wide variety of bioactive molecules (i.e., cytokines, chemokines, growth factors, enzymes, protease inhibitors, ECM proteins) bind to the cell surface and ECM [10–12] and thereby function in the control of normal and pathological processes, among which are morphogenesis, tissue repair, inflammation, vascularization, and cancer metastasis [8–10, 12]. Unlike the well resolved biosynthetic pathway, the mode of HS breakdown is less characterized. Enzymatic activity capable of cleaving glucuronidic linkages and releasing polysaccharide chains resistant to further degradation by the enzyme was first identified by Ogren and Lindahl [13]. The physiological function of this activity was initially implicated in degradation of macromolecular heparin to physiologically active fragments [13, 14]. Subsequent studies revealed that the same enzyme (heparanase) is critically involved in various pathologies such as cancer progression [15], chronic inflammation [16, 17] and kidney dysfunction [18]. The present review summarizes our long term and ongoing research on the biology of the heparanase enzyme emphasizing its function in the tumor microenvironment and involvement in cancer and inflammation.
Heparan Sulfate Proteoglycans (HSPGs)
Proteoglycans are composed of a core protein to which GAGs side chains are covalently attached. Units of N-acetylglucosamine and glucuronic/iduronic acid form heparan sulfate (HS). HS biosynthesis is initiated by formation of a polysaccharide-protein linkage region, attaching four sugar units to a serine residue in the core protein. This tetrasaccharide sequence is extended by alternating addition of _N_-acetylglucosamine (GlcNAc) and _D_-glucuronic acid (GlcA) residues, forming repeating disaccharides of (GlcAß1,4-GlcNAcα1,4)n. During this polymerization process, the repeating disaccharide units are modified by a series of reactions in a sequential mode [19]. The first modification is _N_-deacetylation/_N_-sulfation of GlcNAc units, followed by C5-epimerization of GlcA to _L_-iduronic acid (IdoA) residues, and _O_-sulfation at C2 of IdoA and C6 of GlcNS residues. Tissue/cell specific regulation during HS biosynthesis [20, 21] dictates subtle and selective modulation of interactions between HS and various proteins [22, 23]. Two main types of cell-surface HS proteoglycans (HSPGs) core proteins have been identified: the transmembrane syndecan with four isoforms, carrying HS near their extracellular tips and occasionally also chondroitin sulfate chains near the cell surface [10], and the glycosylphosphatidyl inositol (GPI)-linked glypican with six isoforms, carrying several HS side chains near the plasma membrane and often an additional chain near the tip of its ectodomain [24]. Two major types of ECM-bound HSPGs are found: agrin, abundant in most basement membranes, primarily in the synaptic region [25]; and perlecan, endowed with a widespread tissue distribution and a very complex modular structure [26].
From mice to worms, embryos that lack HS die during gastrulation [27], suggesting a critical developmental role for HSPGs. HSPGs function is not limited to developmental processes but play key roles in numerous biological settings, including cytoskeleton organization, cell-cell and cell-ECM interactions [8, 28, 29]. HSPGs exert their multiple functional repertoires via several distinct mechanisms that combine structural, biochemical and regulatory aspects. By interacting with other macromolecules such as laminin, fibronectin, and collagens I and IV, HSPGs contribute to the structural integrity, self-assembly and insolubility of the ECM and basement membrane, thus intimately modulating cell-ECM interactions [10, 30, 31]. Accumulating evidence indicate that HSPGs act to inhibit cellular invasion by promoting tight cell-cell and cell-ECM interactions, and by maintaining the structural integrity and self assembly of the ECM [32, 33]. Notably, one of the characteristics of malignant transformation is down regulation of GAGs biosynthesis, especially of the HS chains [32, 33]. Low levels of cell surface HS also correlate with high metastatic capacity of many tumors. Biochemically, HSPGs often facilitate the biological activity of bound ligands by actively participating in receptor-ligand complex formation [34]. In other cases, HSPGs mediate cellular uptake and catabolism of selected ligands [34], and/or sequester polypeptides to the ECM and cell surface, generally as an inactive reservoir [6, 35–38]. Cleavage of HSPGs would ultimately release these proteins and convert them into bioactive mediators, ensuring rapid tissue response to local or systemic cues.
Mammalian Heparanase
Heparanase is an endo-β-glucuronidase that cleaves HS side chains presumably at sites of low sulfation, releasing saccharide products with appreciable size (4–7 kDa) that can still associate with protein ligands and facilitate their biological potency. Mammalian cells express primarily a single dominant functional heparanase enzyme (heparanase-1) [39, 40]. A second heparanase (heparanase-2) has been cloned and sequenced but has not been shown to have HS degrading activity [41, 42]. For simplification, throughout this review we will refer to heparanase-1 as heparanase. Enzymatic degradation of HS leads to disassembly of the ECM and is therefore involved in fundamental biological phenomena associated with tissue remodeling and cell migration, including inflammation, angiogenesis and metastasis [15, 39, 40]. Normally, heparanase is found mainly in platelets, mast cells, placental trophoblasts, keratinocytes and leukocytes. Heparanase released from activated platelets and cells of the immune system facilitates extravasation of inflammatory and tumor cells [43]. It also stimulates endothelial mitogenesis, primarily through release of HS-bound growth factors (i.e., FGF, HGF, VEGF) residing in the ECM [44].
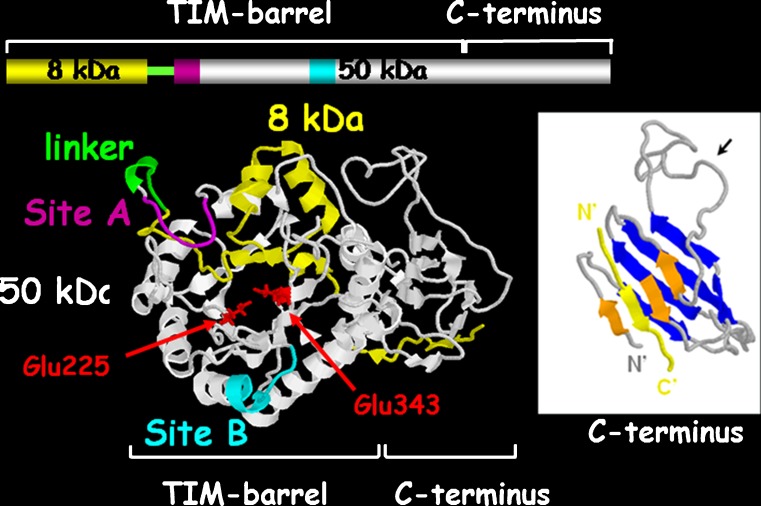
The heparanase mRNA encodes a 61.2 kDa protein with 543 amino acids. This pro-enzyme is post translationally cleaved into 8 and 50 kDa subunits that non-covalently associate to form the active heparanase [45, 46] (Fig. 1). Heterodimer formation is essential for heparanase enzymatic activity [45, 47]. The heparanase structure delineates a TIM-barrel fold harboring the enzyme’ active site and a C-terminus domain that is critical for heparanase secretion and signaling function [48] (Fig. 1). Site-directed mutagenesis revealed that similar to other glycosyl hydrolases, heparanase has a common catalytic mechanism that involves two conserved acidic residues, a putative proton donor at Glu225 and a nucleophile at Glu343 [49] (Fig. 1). Cellular processing of the latent 65 kDa pro-heparanase into its active 8 + 50 kDa heterodimer involves removal a 6 kDa linker segment and is inhibited by a cell permeable inhibitor of cathepsin L [50]. Moreover, multiple site-directed mutagenesis and cathepsin L gene silencing and knockout experiments indicate that cathepsin L is the predominant enzyme responsible for processing and activation of pro-heparanase [51]. Applying a structural model, it has been demonstrated that the linker segment, or even a small 1 kDa portion at its C-terminus, render the active site inaccessible to the HS substrate [51].
Fig. 1.
Predicted model of the active heparanase heterodimer showing the 50 + 8 kDa heparanase subunits, TIM-barrel and C-terminus domains, active site (Glu225 & E343, Red), and heparin binding domains (sites A and B). Right: Predicted structure of the C-terminus domain
Heparanase in Cancer
In early studies heparanase activity was shown to be associated with the metastatic potential of tumor-derived cells such as B16 melanoma [52] and T-lymphoma [53]. These observations gained substantial support when specific molecular probes became available shortly after cloning of the heparanase gene. Both over-expression [54, 55] and silencing [55, 56] of the heparanase gene clearly indicate that heparanase not only enhances cell dissemination, but also promotes the establishment of a vascular network that accelerates primary tumor growth and provides a gateway for invading metastatic cells [40, 57]. These studies enabled researchers to critically approve the notion that HS cleavage by heparanase is required for structural remodeling of the ECM underlying tumors and endothelial cells, thereby facilitating cell invasion, and providing a proof-of-concept for the pro-metastatic and pro-angiogenic capacity of heparanase. The clinical significance of the enzyme in tumor progression emerged from a systematic evaluation of heparanase expression in primary human tumors. Immunohistochemistry, in situ hybridization, RT-PCR and real time-PCR analyses revealed that heparanase is up-regulated in essentially all major types of human cancer, namely carcinomas, sarcomas and hematological malignancies [40, 48, 57, 58] (Table 1). Notably, increased heparanase levels were most often associated with reduced patients’ survival post operation (Fig. 2), increased tumor metastasis and higher microvessel density [40, 57, 59] (Table 1), thus critically supporting the intimate involvement of heparanase in tumor progression and encouraging the development of heparanase inhibitors as anti-cancer therapeutics [59–63].
Table 1.
Correlation between heparanase expression and key clinical parameters in human cancer
| Author (Ref.) | Carcinoma | Patient number | % Positive | Positive correlation with | ||
|---|---|---|---|---|---|---|
| Gohji et al., 2001 [197] | Bladder | 40 | 85 (17/20) | aMVD | Metastasis | Survival |
| Gohji et al., 2001 [198] | Bladder | 67 | 48 (32/67) | Metastasis | ||
| Maxhimer et al., 2002 [66] | Breast | 53 | 36 (19/53) | Tumor size | ||
| Shinyo et al., 2003 [199] | Cervical | 92 | 49 (45/92) | MVD | Survival | |
| Friedman et al., 2000 [178] | Colon | 17 | 100 | |||
| Nobuhisa et al., 2005 [200] | Colon | 54 | 69 (37/54) | Metastasis | Survival | |
| Sato et al., 2004 [201] | Colorectal | 130 | 25 (33/130) | MVD | Metastasis | Survival |
| Watanabe et al., 2003 [202] | Endometrial | 40 | 50 (20/40) | MVD | ||
| Endo et al., 2001 [203] | Gastric | 63 | 49 (31/63) | Metastasis | ||
| Tang et al., 2002 [68] | Gastric | 116 | 83 (96/116) | Tumor size | Metastasis | Survival |
| Takaoka et al., 2003 [204] | Gastric | 44 | 80 (35/44) | Metastasis | Survival | |
| Doweck et al., 2006 [64] | Head & Neck | 74 | 86 (64/74) | Tumor size | Survival | |
| El-Assal et al., 2001 [65] | Hepatocellular | 55 | 47 (26/55) | MVD | Tumor size | |
| Takahasi et al., 2004 [205] | Lung | 76 | Metastasis | Survival | ||
| Cohen et al., 2008 [206] | Lung | 114 | 75 (85/114) | Metastsis | Survival | |
| Kelly et al., 2003 [207] | Multiple Myloma | 100 | 86 (86/100) | MVD | ||
| Bar-Sela et al., 2006 [208] | Nasopharyngeal | 46 | 35 (16/46) | Survival | ||
| Zheng et al., 2009 [209] | Neuroblastoma | 42 | 62 (26/42) | Survival | ||
| Koliopanos et al., 2001 [146] | Pancreatic | 33 | 75 (25/33) | Survival | ||
| Kim et al., 2002 [210] | Pancreatic | 89 | 78 (69/89) | Survival | ||
| Rohloff et al., 2002 [148] | Pancreatic | 50 | 76 (38/50) | Metastasis | Survival | |
| Mikami et al., 2008 [211] | Renal | 70 | Metastasis | Survival | ||
| Ben-Izhak et al., 2006 [212] | bSalivary gland | 60 | 70 (42/60) | Survival | ||
| Shafat et al., 2010 [58] | Ewing’s Sarcoma | 69 | 100 | Tumor size | ||
| Nagler et al., 2007 [67] | Tongue | 60 | 92 (55/60) | Tumor size | Survival |
Fig. 2.
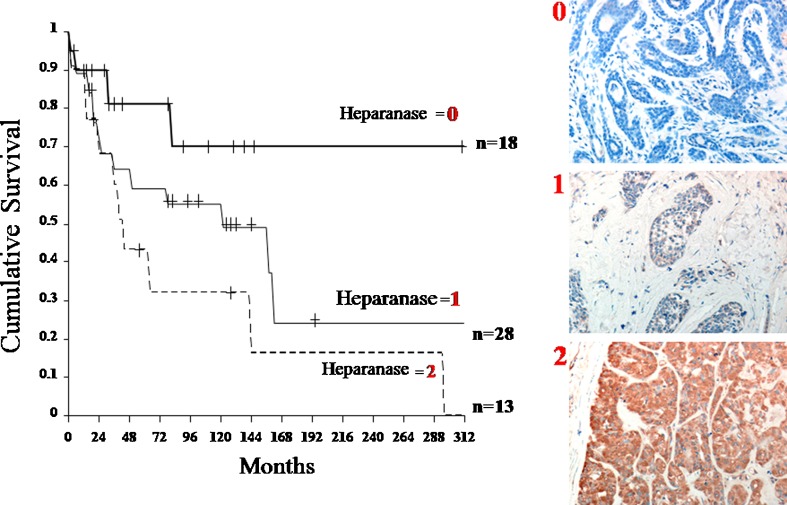
Overall survival curves of patients with salivary gland tumors according to heparanase immunostaining levels. Right: Immunohistochemical staining of heparanase in patients with salivary gland tumors. Formalin-fixed paraffin-embedded sections of salivary gland tumors were subjected to immunostaining of heparanase, applying anti-heparanase pAb 733. Staining was graded as 0 (negative), 1 (weak) and 2 (strong). Left: Kaplan-Meier analysis showed poor survival of patients with positive (score 1 & 2) heparanase expression, compared with patients who were diagnosed as heparanase-negative (score 0). After 300 months (25 years) of follow-up, 0% of heparanase-positive patients survived compared with 70% of patients with no detectable heparanase expression. [Reproduced from Ben-Izhak et al., (2006) Neoplasia8, 879–884]
Importantly, heparanase up-regulation in human tumors (i.e., head & neck, tongue, hepatocellular, breast and gastric carcinomas) is associated with large tumors [64–68] (Table 1). Likewise, heparanase over-expression enhanced [54, 69–72], while local delivery of anti-heparanase siRNA inhibited [55] the progression of tumor xenografts, altogether implying that heparanase function is not limited to tumor metastasis but is also engaged in accelerated vascularization and growth of the primary lesion [54] (Table 1). Evidence indicates that heparanase not only assists in the breakdown of ECM but also is involved in regulating the bioavailability and activity of growth factors and cytokines. Briefly, as discussed above, various heparin-binding growth factors are sequestered by HS in the ECM, providing a localized, readily accessible depot, protected from proteolytic degradation [6, 73], yet available to activate cells after being released by heparanase. Local release and activation of tissue-specific growth factors is clearly involved in creating a favorable ‘soil’ for growth of the primary tumor as well as in dictating the organ selectivity of metastasis.
Other ECM-Degrading Enzymes
Tumor cell invasion and spread through the blood and lymphatics is the hallmark of malignant disease and the greatest impediment to cancer cure. Metastasis is a multistage process that requires cancer cells to escape from the primary tumor, survive in the circulation, seed at distant sites and grow. Each of these processes involves rate-limiting steps that are influenced by the malignant and non-malignant cells of the tumor microenvironment [74–76]. Numerous studies have shown that metastases formation depends on the ability of tumor cells to invade blood vessel walls and tissue barriers in a process involving enzymes capable of digesting ECM components. Attention focused on serine (i.e., plasminogen activators) and cysteine (i.e., cathepsins) proteases as well as matrix metalloproteinases (MMPs) [77]. These enzymes, whose substrates include major components of the ECM are often upregulated in metastatic cancers [77, 78]. It was originally thought that their role was simply to break down tissue barriers, enabling tumor cells to invade through stroma and blood vessels at primary and secondary sites. Subsequent studies revealed that MMPs also participate in angiogenesis [79] and their substrate repertoire goes far beyond ECM elements [77, 80, 81]. While MMPs attracted much attention, other proteases constitute the tumor milieu. A large family of proteases consists of cysteine proteases named cathepsins. Like MMPs, some cathepsins are often upregulated in cancer and, once secreted or localized to the cell surface, can degrade components of the ECM [82–84]. In addition, as discussed above, cathepsin L is held responsible for the processing and activation of heparanase, found to be strongly implicated in cell dissemination associated with tumor metastasis, angiogenesis, and inflammation.
Notably, proteases can contribute to the sustained growth of established tumor foci by cleavage of the ectodomain of membrane-bound pro-forms of growth factors, releasing peptides that are mitogens for tumor cells and/or vascular endothelial cells [77]. On the other hand, some ECM degradation products (i.e., tumstatin, endostatin) suppress endothelial cell proliferation and thereby inhibit tumor angiogenesis [85], further emphasizing the significance of the tumor microenvironment in the control of cell growth and function. Moreover, some ECM-degrading enzymes (i.e., cathepsins, plasminogen activators, MMPs, heparanase) are also engaged in multiple signaling pathways, primarily by means of non-enzymatic activities that affect both the tumor cells and the tumor microenvironment [40, 77]. Both normal and tumor cells appear to use the same set of ECM-degrading enzymes and molecular machinery to perform their biological functions. Thus, the normal physiological functions of proteases and heparanase in embryonic morphogenesis, wound healing, tissue repair and inflammation have been effectively ‘hijacked’ by tumor cells.
Heparanase-Protease Cooperation
A cross-talk between heparanase and MMPs has been demonstrated. Thus, enhanced expression of heparanase leads to increased levels of MMP-9, while heparanase gene silencing resulted in reduced MMP-9 activity [86]. Moreover, not only MMP-9 but also urokinase-type plasminogen activator (uPA) and its receptor (uPAR), molecular determinants responsible for MMP-9 activation, are up-regulated by heparanase [86]. These findings provided the first evidence for cooperation between heparanase and MMPs in regulating HSPGs on the cell surface and likely in the ECM, and are supported by our generation and characterization of heparanase knockout (KO) mice. Despite the complete lack of heparanase gene expression and enzymatic activity, heparanase-KO mice develop normally, are fertile, and exhibit no apparent anatomical or functional abnormalities [87]. Notably, heparanase deficiency was accompanied by a marked elevation of MMP family members such as MMP-2, MMP-9, and MMP-14, in an organ-dependent manner, suggesting that MMPs provide tissue-specific compensation for heparanase deficiency [87]. These and other results [86] suggest that heparanase is intimately engaged in the regulation of gene transcription and acts as a master regulator of protease expression, mediating gene induction or repression depending on the biological setting. These effects of heparanase are attributed to its nuclear uptake, degradation of nuclear HS and the associated activation of histone deacetylase (HDAC) activity [88].
Inflammation–A Major Determinant of the Tumor Microenvironment and Cancer Progression
According to Paget’s ‘seed and soil’ hypothesis [89], metastatic spread cannot simply be explained by blood vessels anatomy, but is rather directed in a specific manner toward a ‘congenital soil’ [89]. The ‘seed and soil’ hypothesis is now widely accepted [90], and the terms ‘seed’ and ‘soil’ have been characterized to some extent at the cellular and molecular levels. The ‘soil’ is defined as biologically unique microenvironment that enables the ‘seed’ (metastatic cancer cell) to strike roots, germinate, and grow (i.e., proliferate). The ‘congenital soil’ can be made, for example, of endothelial cells expressing specific adhesion molecules in different vascular beds [91], or of tumor cells expressing receptors that respond to specific ligands, leading to their traffic and arrest in a specific tissue. The CXCR4 receptor, for example, plays a role in the bone marrow, lymph nodes, and pulmonary homing of metastases from breast tumor that expresses its ligand, CXCL12 (SDF-1) [90, 92]. In addition, emerging evidence indicates that bone marrow-derived hematopoietic progenitor cells positive for vascular endothelial growth factor receptor (VEGFR) arrive at a distant organ, alter the tissue microenvironment and promote the recruitment of metastatic tumor cells [92, 93]. Accordingly, treatment with neutralizing VEGFR during the early phases of primary tumor development or following the formation of the premetastatic niche inhibits metastasis [94].
While the ‘soil’ is still insufficiently characterized, the ‘seed’ (i.e., metastatic cancer cell) has been studied in far greater detail. It is generally accepted that in addition to the dynamic adhesive and motility capabilities intrinsic to tumor cells, tumors are supported continuously by host (stromal) cells which are recruited to the neoplastic lesion and affect all stages of tumor progression and metastasis. The tumor microenvironment includes epithelial cells, fibroblasts, endothelial cells, infiltrated leukocytes, and bone marrow-derived (stem) cells of the above lineages. Endothelial cells lining blood and lymph vessels are major component of the tumor microenvironment, and anti-angiogenesis therapy, targeting vascular endothelial growth factor (VEGF) or its receptor (VEGFR) is implemented clinically [95]. Cancer-associated fibroblasts (CAF) comprise another important component of the tumor microenvironment [96, 97]. CAF are phenotypically and functionally distinct from their normal counterparts in their increased rate of proliferation and expression of ECM components and growth factors [98–100]. By targeting HPV early region that includes the E6/E7 oncogenes to the skin tissue (K14-HPV16), it was demonstrated that CAF are pro-inflammatory and enhance tumor growth, angiogenesis, and recruitment of macrophages [101]. Thus, CAF functionally link major components of the tumor microenvironment (macrophages, endothelial cells) in a cross-talk that enhances tumor development and cell dissemination.
The last decade critically revealed the decisive role of inflammatory responses in different stages of tumor development and metastasis [102]. The most frequently found immune cells within the tumor microenvironment are tumor-associated macrophages [102, 103]. Notably, high density of these cells correlates with poor prognosis, while removal of macrophages almost completely ablated metastasis in a mouse of breast cancer [103–106]. In a recent study (discussed in detail below) we have demonstrated the importance of mucosal macrophages and heparanase in sustaining immune-epithelial crosstalk underlying colitis-associated colon tumorigenesis [16].
Heparanase and Heparan Sulfate in Inflammation
Upregulation of heparanase was reported in different inflammatory conditions, often associated with degradation of HS and release of chemokines anchored within the ECM network and cell surfaces. Moreover, remodelling of the ECM facilitates transmigration of inflammatory cells towards the injury sites. Prior to cloning of the heparanase gene, heparanase activity originating in activated cells of the immune system (T-lymphocytes, neutrophils) has been found to contribute to their ability to penetrate blood vessels and accumulate in target organs [43]. In a more recent study we have demonstrated that up-regulation of heparanase, locally expressed (i.e., by vascular endothelium, skin keratinocytes) at the site of inflammation, is an essential step of delayed type hypersensitivity [107]. Degradation of HS in the subendothelial basement membrane resulted in vascular leakage, a hallmark of delayed type hypersensitivity skin reactions [107]. Upregulation of heparanase has also been found in colonic epithelium of patients with inflammatory bowel disease (IBD) [16, 108] (Fig. 3), and in skin lesions of psoriasis patients (our unpublished results). Notably, heparanase staining was primarily detected in epithelial rather than immune cells, altogether indicating that immunocytes are not the primary source of the enzyme in inflammation. Heparanase activity was also found to be dramatically elevated in sinovial fluid from rheumatoid arthritis patients [17], suggesting an important role for heparanase in promoting joint destruction and indicating heparanase as an attractive target for the treatment of rheumatoid arthritis [17].
Fig. 3.
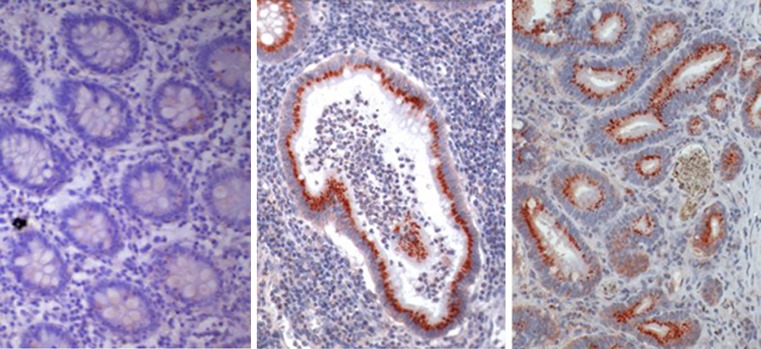
Heparanase expression in acute and chronic phases of UC. Tissue specimens derived from normal colon tissue (left), UC patients in acute (middle) and chronic (right) phases of the disease were stained with anti-heparanase antibody (reddish staining). Photographs are representative of control (n = 29) and UC (n = 10) samples (original magnification, ×200). [Reproduced from Lerner et al., (2011) J Clin Invest 121:1709–1721]
Heparanase Powers a Chronic Inflammatory Circuit that Promotes Colitis-Associated Tumorigenesis
The most feared long-term complication of IBD (in particular, UC) is colon carcinoma, as patients with ulcerative colitis (UC) have a risk of colorectal cancer which is an order of magnitude higher than the normal population [109]. In fact, colon carcinoma represents a paradigm for the association between inflammation and cancer [110]. While significant progress has been made in deciphering the role of inflammatory cytokines (TNFα, IL-1β, IL-6) and their downstream transcription factors (NFκB, STAT3) in tumor-stimulating cross-talk between immune and epithelial cells [110–112], little is known about the role of ECM-degrading enzymes in this cross-talk. Based on the preferential expression of heparanase in chronically inflamed colonic epithelium (Fig. 3) and increased incidence of colon cancer in colitis patients, we hypothesized that stimulation of heparanase expression plays an important role in the pathogenesis of UC, representing a mechanistic link between inflammation and cancer. Utilizing UC tissue specimens, along with a mouse model of colitis-associated cancer induced by the carcinogen azoxymethane (AOM), followed by the inflammatory agent dextran sodium sulfate (DSS) [113], we found that heparanase is constantly overexpressed by the colonic epithelium in UC and experimental colitis during both the acute and chronic phases of the disease. Moreover, heparanase over-expression preserves a chronic inflammatory conditions in DSS colitis and thus creates a tumor-promoting microenvironment characterized by enhanced NF-κB signaling [111, 114], STAT 3 induction [115], and increased vascularization. Furthermore, we identified a novel biological mechanism contributing to chronic colitis and the associated colon tumorigenesis. This mechanism involves a self-sustained cycle through which heparanase of epithelial origin, acting synergistically with the local flora and cytokine milieu, facilitates abnormal activation of innate immune cells (i.e., macrophages) which, in turn, stimulate further production of the enzyme by the colonic epithelium. Moreover, in chronic colitis activated macrophages represent a primary source of cathepsin L responsible for proteolytic activation of latent heparanase [16]. A detailed analysis of the functional importance of heparanase in modulating inflammatory responses and sustaining immune-epithelial crosstalk underlying the pathogenesis of chronic colitis-associated tumorigenesis, is presented below.
Immunohistochemical staining revealed that while heparanase expression was practically not detected in healthy colon epithelium, in UC specimens the epithelial lining showed a marked expression of heparanase in both the acute (Fig. 3, middle) and chronic (Fig. 3, right) phases of the disease. In all phases, inflammatory cells in the involved areas, showed little or no heparanase expression. We investigated the tempo-spatial pattern of heparanase expression in experimental DSS-induced colitis. DSS causes an acute inflammatory reaction and ulceration in the colon [116] and when three cycles of DSS administration are applied, acute inflammation is followed by chronic colitis [116]. Moreover, three cycles of DSS subsequent to a single pretreatment with the chemical carcinogen AOM result in development of colon tumors in ~100% of the treated mice [113], representing a well established model of colitis-associated cancer. A marked increase in heparanase mRNA, protein and enzymatic activity was readily detected in the colon during all phases of DSS-induced colitis. In accordance, increased levels of cathepsin L mRNA and protein were detected in the mouse colonic tissue at corresponding time points [16].
Heparanase Over-Expression Preserves a Chronic Inflammatory Condition and Increases the Incidence and Severity of Colitis-Associated Tumors
Unlike ulcerative colitis (UC) patients, in which increased heparanase expression is constantly preserved (Fig. 3), in DSS colitis, heparanase levels gradually decreased during the chronic phase. Thus, to assess the precise role of heparanase in colitis-related tumorigenesis, we utilized heparanase transgenic (_Hpa_-tg) mice previously shown to express elevated levels of heparanase in colonic epithelium, but not in splenocytes [117], closely resembling heparanase expression pattern in UC patients. Applying the AOM-DSS protocol, we noted a marked increase in tumor incidence, vascularization and severity in _Hpa_-tg mice vs. their wt littermates. Moreover, when three DSS cycles were administered alone, without AOM pretreatment, none of the wt mice developed colonic tumors, whereas in 90% of the _Hpa_-tg mice colonic tumors were easily seen, emphasizing the essential role of heparanase in this experimental system [16].
In contrast to the acute phase, heparanase over-expression profoundly affected the chronic phase of DSS-induced colitis, as demonstrated by microscopic and biochemical analyses of inflammatory phenotypes expressed by _Hpa_-tg vs. wt mice on experimental day 80 (1 month after cessation of the last DSS cycle). Altogether, our results indicate that the constantly elevated levels of heparanase in _Hpa_-tg mice (resembling the observations in UC patients, Fig. 3) markedly affect the chronic phase of DSS colitis and create a tumor-promoting inflammatory microenvironment primarily via increased vascularization, enhanced NF-κB signaling, augmented levels of TNFα, and STAT 3 activation [16].
Macrophages Represent Both a Cellular Target for Heparanase Action and Regulators of Heparanase Expression/Activity in Chronic Colitis
Macrophages are known to have a dual role in inflammation. In the scenario of inflammation resolution, macrophages perform phagocytosis and produce anti-inflammatory cytokines, thereby preventing inflammatory responses from lasting too long. However, if inflammation resolution is deregulated, macrophage response switches to the pattern of chronic inflammation. Recruitment and activation of macrophages within the intestinal mucosa play a key role in the pathogenesis of both human UC [118, 119] and murine DSS colitis [120, 121]. Moreover, activated macrophages are candidate cells linking between inflammation and cancer [106, 122]. Notably, the tumor promoting cytokines IL-1, IL-6 and TNFα [111, 112] are produced by activated macrophages and, along with macrophage-derived growth factors and reactive oxygen species, foster tumor initiation [119, 122]. This notion led us to examine the involvement of macrophages in our system. In fact, we detected increased macrophage infiltration in the colon of _Hpa_-tg vs. wt mice on day 80 of the chronic DSS colitis model [16]. Macrophages infiltrating the _Hpa_-tg colon manifested a striking pro-inflammatory profile, evidenced by increased number of TNFα expressing macrophages and elevated NFκB signaling. These findings led us to assume that heparanase over-expression directly affects macrophage activation. To recapitulate conditions occurring in UC (i.e., heparanase-rich environment and abundant microbial flora), we isolated mouse peritoneal macrophages and stimulated them with lipopolysacharide (LPS) in the absence or presence of recombinant active heparanase. Pretreatment with heparanase strongly sensitized macrophages to activation by LPS, as indicated by a marked increase in TNFα, IL-6 and IL-12p35 [16], all macrophage-derived cytokines known to be induced by TLR-4 signaling and tightly involved in the pathogenesis of UC [119]. These findings are in agreement with previous reports showing that intact extracellular HS inhibits LPS-mediated TLR4 signaling and macrophage activation, and that its removal relieves this inhibition [123]. Given that one of the unique aspects of colorectal cancer development is the involvement of lumenal flora and TLR signaling [124, 125], the observed ability of heparanase to sensitize macrophages to LPS activation is of particular significance in light of the increased epithelial permeability to lumenal microbial products, characteristic of UC [126]. Altogether, the above data suggest that up-regulated heparanase enables enhanced activation of macrophages, reprogramming their response from resolution of inflammation to unresolved chronic colitis.
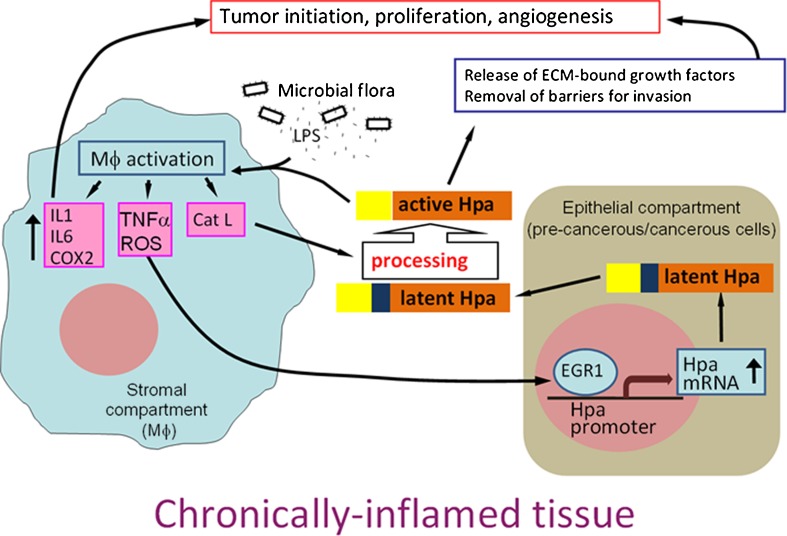
On the other hand, we found that activated macrophages are capable of inducing heparanase expression in colonic epithelial cells, most likely through TNFα-mediated stimulation of Egr1, a powerful inducer of heparanase transcription in colonic tumor cells [127] and DSS colitis [16]. Notably, TNFα was shown to stimulate Early growth response 1 (Egr1 transcription factor regulating heparanase promoter activity) expression in IBD patients [128]. In addition, due to their unique ability to secrete mature cathepsin L and allow extracellular accumulation of the active enzyme [129], activated macrophages appear to be responsible for proteolytic activation of latent proheparanase in colitis. Thus, macrophages not only represent a cellular target for heparanase action, but also decisively upregulate heparanase in chronic colitis, both at the transcriptional and posttranslational levels. These results demonstrate and highlight the cooperation between two cellular compartments (i.e., colon epithelium and activated macrophages) in heparanase activation during inflammation-associated colon carcinogenesis. Collectively, our results [16] indicate that heparanase generates a self-sustaining connection between chronic colitis and tumorigenesis (Fig. 4). Briefly, macrophages activated by influx of the luminal flora secrete TNFα and stimulate production of heparanase by the colon epithelium. The secreted 65 kDa latent heparanase is processed into its active form by cathepsin L, supplied by the activated macrophages. Enzymatically-active heparanase sensitizes macrophages to further activation by microbial flora, thus preventing inflammation resolution, switching macrophage responses to the chronic inflammation pattern and creating tumor-inducing inflammatory environment [16]. In addition, high heparanase levels support tumor progression via stimulation of angiogenesis, release of ECM-bound growth factors and bioactive HS fragments and removal of extracellular barriers for invasion (Fig. 4). Importantntly, as presented in Fig. 4, the above described self-sustaining tumor promoting inflammatory circuit may occur in other organs where inflammation plays a significant role in tumorigenesis and preferential expression of heparanase has been reported during cancer progression (Fig. 4).
Fig. 4.
A model of heparanase-driven vicious cycle that may power chronic inflammation and the associated tumorigenesis. Increased levels of TNFα secreted by activated macrophages induce heparanase expression in the epithelial compartment via Egr1-dependent mechanism. Secreted latent heparanase is processed into its active form by cathepsin (supplied by the macrophages), and in turn sensitizes macrophages to further activation (i.e., in case of colitis by luminal flora) thus preventing inflammation resolution, switching macrophage responses to the chronic inflammation pattern and creating tumor-inducing inflammatory environment. In addition, heparanase promotes tumor progression via stimulation of angiogenesis, release of ECM-bound growth factors and bioactive HS fragments, and removal of extracellular barriers for invasion. [Reproduced in part from Lerner et al., (2011) J Clin Invest 121:1709–1721]
The newly identified heparanase-powered vicious cycle may explain a yet poorly understood “multiplier effect” in IBD inflammation, in which even a small initial elevation in ‘initiating’ inflammatory stimuli gives rise to large increases in downstream cytokines [130]. Thus, disruption of the heparanase-driven chronic inflammatory circuit is highly relevant to the design of therapeutic interventions in colitis and the associated cancer.
Role of Heparanase in Radiation Enhanced Invasiveness of Pancreatic Carcinoma
Pancreatic cancer is one of the most aggressive neoplasm with an extremely low 5-year survival rate [131–133]. Currently, pancreaticoduodenectomy is the only curative form of treatment, however ~90% of pancreatic cancer patients miss the opportunity for complete surgical resection at the time of diagnosis [131, 134]. Thus, radiotherapy remains a major component of treatment modalities for controlling pancreatic tumor progression [135]. However, pancreatic cancer often shows resistance to ionoizing radiation (IR), and randomized trials could not demonstrate benefit from radiation, revealing rather conflicting results [136, 137]. Accumulating preclinical and clinical data suggest that IR may stimulate tumor aggressiveness [138–144], although the identity of downstream effectors acting at the cell or tissue levels and responsible for this effect, remains poorly investigated. Our recent results indicate that IR augments heparanase expression and thereby aggressiveness of pancreatic carcinoma both in vitro and in vivo [145]. Causal involvement of heparanase in pancreatic carcinoma progression is well-documented. There was a 30-fold increase in heparanase mRNA in pancreatic cancer tissue samples, in comparison to normal pancreatic tissue [146]. Moreover, elevated levels of the enzyme have been found in body fluids of patients with active pancreatic cancer disease [147] as compared to healthy donors. Pancreatic cancer patients whose tumors exhibit high levels of the heparanase mRNA had a significantly shorter postoperative survival time than patients whose tumors contained relatively low levels of heparanase [146, 148]. A recent finding that heparanase positivity is a highly significant independent variable for pancreatic adenocarcinoma dedifferentiation and lymph node metastasis further demonstrate a crucial role of the enzyme in the aggressiveness of pancreatic cancer [149].
We have demonstrated that clinically relevant doses of IR augment invasive ability of pancreatic carcinoma cells in vitro and in vivo through upregulation of heparanase expression, and revealed that the molecular mechanism responsible for IR-induced heparanase transcription involves the Early growth response 1 (Egr1) transcription factor [145]. As a transcriptional regulator, Egr1 can both induce and repress the expression of its target genes, including heparanase [127, 150, 151]. Egr1 was previously shown to activate heparanase expression in T lymphocytes, prostate, breast, and colon carcinomas, but to inhibit its transcription in melanoma cells [127]. Transactivation studies using Egr1 expression vector, co-transfected with a reporter construct encoding for LUC under the heparanase promoter, showed that in pancreatic carcinoma cells Egr1 acts to repress heparanase transcription. In fact, IR treatment of PANC1 pancreatic carcinoma cells resulted initially in a transient increase in Egr1, followed by profound and continuous decline in Egr1 levels, as compared to its basal levels in untreated PANC1 cells [145]. Moreover, ChIP analysis revealed a marked decrease in occupancy of the heparanase promoter by Egr1 following IR treatment. We have found that pancreatic carcinoma cells express high basal levels of Egr1 and of the NGFI-A/Egr1-binding protein NAB2. NAB2 is a transcriptional co-repressor that directly interacts with Egr1 and represses activation of its target promoters [152]. It is therefore plausible that in the presence of both Egr1 and NAB2 the heparanase promoter is repressed in pancreatic cells. However, following IR-associated temporal decrease in Egr1, this repression is no longer operative, allowing for activation of the heparanase promoter.
Combination of radiotherapy with drugs that inhibit IR-induced tumor aggressiveness may be an attractive strategy to diminish adverse pro-metastatic action while retaining the therapeutic benefit of radiation, thus reducing resistance of pancreatic cancer to treatment. We have demonstrated that compound SST0001 (Sigma-Tau Research Switzerland, SA), a specific potent heparin-based inhibitor of heparanase enzymatic activity which lacks anti-coagulant activity [59, 153–155], attenuated radiation-induced invasiveness in vitro [145] in an orthotopic model of pancreatic cancer. Thus, spread of orthotopically growing pancreatic tumors was significantly reduced in mice treated with a combination of SST0001 and IR, as compared with either modality alone [145]. Taken together, our results support the combination of radiotherapy with a specific heparanase inhibitor as an effective strategy to prevent tumor resistance and progression, observed in many IR-treated pancreatic cancer patients. Notably, inducibility of heparanase by radiation appears not to be limited to pancreatic tissue, as it was recently demonstrated that heparanase is upregulated by IR in liver [156] and by ultraviolet B radiation in human skin [157].
Heparanase, a Promising Antigen for Tumor Immunotherapy
Heparanase expression at the early stages of tumor initiation and progression, and by the majority of tumor cells, can be utilized to turn the immune system against the very same cells. Accumulating evidence indicate that peptides derived from human heparanase can elicit a potent anti-tumor immune response, leading to lysis of heparanase-positive human gastric (KATO III), colon (SW480), and breast (MCF-7) carcinoma cells, as well as hepatoma (HepG2) and sarcoma (U-2 OS) cells [158–160]. In contrast, no killing effect was noted towards autologous lymphocytes [158–160]. Notably, the development of tumor xenografts produced by B16 melanoma cells was markedly reduced in mice immunized with peptides derived from mouse heparanase (i.e., aa 398–405; 519–526) compared to a control peptide in both immunoprotection and immunotherapy approaches [158, 161]. To increase the immunogenicity of these peptide vaccines, Wang et al. designed four-branched multiple antigenic peptides based on HLA-A2-restricted cytotoxic T lymphocyte (CTL) epitopes of human heparanase. These multiple antigenic peptides vaccines were capable of inducing HLA-A2-restricted and heparanase-specific CTL in vitro and in mice, exerting no effect on autologous lymphocytes and dendritic cells. Moreover, compared to their corresponding linear peptides, heparanase multiple antigenic peptide vaccines elicited a much stronger lysis of tumor cells by activating CD8+ T lymphocytes and increasing the release of IFN-γ. Given the broad spectrum, high effectiveness, high specificity and safety, it was concluded that multiple antigenic peptide vaccines based on CTL epitopes of human heparanase can be used as potent immunogens for tumor immunotherapy [161].
Beckhove et al. elucidated the role of the bone marrow as an important organ for the priming and memory formation of T cell mediated anti-tumor immune response [162–165]. Upon reactivation in vitro and adoptive transfer, tumor antigen-reactive bone marrow memory T cells from tumour patients infiltrated autologous tumors and mediated their complete rejections in xenotransplant mouse models, suggesting their therapeutic potential [162, 163, 166, 167]. These results demonstrate a selective homing of memory T cells from the bone marrow into human tumors suggesting that tumor rejection is based on the recognition of tumor-associated antigens on tumor cells and dendritic cells by specifically activated, central and effector memory T cells [162, 168]. Moreover, it has been demonstrated in a clinical study that adoptive transfer of ex vivo reactivated pre-existing bone marrow memory T cells provides a therapeutic option for the treatment of advanced metastasized breast cancer patients [169]. Of high significance is the evaluation of tumour antigens that are major target structures of spontaneous effector and regulatory T cell responses since the repertoire of such pre-existing tumour specific T cells may have a major influence on the immunological and clinical efficiency of immunotherapeutic strategies. Importantly, HLA-I restricted epitopes from heparanase were characterized as target antigens of spontaneous CD8 T cell responses in breast cancer patients [170]. Briefly, high levels of heparanase specific T lymphocytes were identified in breast cancer patients using heparanase peptide-MHC class I tetramers. Moreover, in a high proportion of these patients, memory T cell responses to heparanase derived HLA-A2 restricted peptides, were demonstrated, leading to generation of anti-tumor cytotoxic T lymphocytes. Thus, heparanase emerged as a new metastasis-associated antigen recognized in breast cancer patients by spontaneously induced memory T lymphocytes [170], indicating that heparanase is an attractive tumor-associated antigen in cancer patients. In addition, the immune system appears capable of sensing ECM degradation and responding to heparan sulfate degradation fragments released by heparanase from the ECM. Thus, heparanase activity might directly activate antigen presenting cells and allow for tumor antigen specific T cell responses in the absence of foreign inflammatory stimuli – a fundamental new mechanism for induction of anti tumor immune responses. Applying newly synthesized long peptides of heparanase that can be processed by dendritic cells, heparanase ranked among the most frequently recognized tumour antigens in patients with pancreatic, colorectal or breast cancer [171]. Together with its selective expression on a small subpopulation of highly malignant metastasizing tumour cells, heparanase provides a unique opportunity for a tumour type independent, metastasis-selective tumour immunotherapy. Taken together, heparanase appears to represent a highly promising target antigen for immunotherapeutic approaches against a broad variety of tumours. While causing the generation of high frequencies of specific CD4 and CD8 memory T cells, heparanase, in contrast to most other tumour antigens (i.e., CEA or MUC1), did not induce spontaneous regulatory T cell responses [171]. Anti-heparanase immunotherapy is thus expected to be prolonged and more efficient due to the absence of T-suppressor cells. These results provide a basis for a vaccination treatment approach that is currently being tested in advanced metastasized breast cancer patients [172]. In fact, studies are underway (Beckhove et al., DKFZ, Heidelberg) to characterize the most promising heparanase-derived long peptides and generate these for clinical use in an immunotherapeutic trial of adoptive T cell transfer of heparanase-reactive, ex vivo activated T cells.
Conclusions and Perspective
Ten years following its cloning, the repertoire of heparanase functions is only starting to be revealed. From activity mainly implicated in cell invasion associated with tumor metastasis, heparanase has turned into a multifaceted protein that appears to participate in essentially all major aspects of tumor progression [39, 48, 173]. Genetic tools (i.e., heparanase knock-out and over-expressing mice [87, 117]) are now available and being utilized to examine the possibility that heparanase is not only intrinsic to tumor cell invasion and angiogenesis, but also modifies, enzymatically and/or by virtue of its pro-adhesion function [173, 174], the site of cell seeding and hence participates in establishing a ‘congenital soil’ for successful homing and colonization of disseminated tumor cells.
Evidence now supports a concept by which growth of the primary tumor is fueled by circulating metastatic tumor cells [175, 176]. According to this notion, tumor cells are present in the circulation in large numbers even at the early stages of cancer and long before metastatic growth at distant sites can be detected [176]. These cells can reinfiltrate and promote growth and angiogenesis of the primary tumor [175]. The possible involvement of heparanase in tumor self-seeding is supported by the timing of its induction during tumorigenesis and its pro-metastatic function. Using the RIP-Tag2 tumor model, it was demonstrated that heparanase mRNA and protein are elevated upon the transition from normal to angiogenic islets, followed by a further increase when solid tumors were detected [177]. Furthermore, heparanase expression is elevated already at the early stages of human neoplasia. In the colon, heparanase gene and protein are expressed already at the stage of adenoma [178], and during esophageal carcinogenesis heparanase expression is induced in Barrett’s epithelium, an early event that predisposes patients to formation of dysplasia which may progress to adenocarcinoma [179]. Tumor self-seeding also facilitates the recruitment of stromal components. The possible involvement of heparanase in this aspect is supported by its pro-angiogenic and pro-inflammatory effects.
There is growing evidence that heparanase upregulates expression of genes that participate in creating aggressive behavior of tumors. These include VEGF, MMP-9, uPA/uPAR and tissue factor and likely other effectors that condition the tumor microenvironment to promote an aggressive cancer phenotype [47, 86, 180]. Although HS has been associated with several functional roles within the nucleus, its inhibition of gene transcription via inhibition of topoisomerase I and histone H3 acetyltransferase (HAT) activity are particularly intriguing [181, 182]. HATs regulate gene expression by catalyzing acetylation of the N-terminal region of histones, thereby modifying chromatin structure in a manner that facilitates transcriptional activation. It was recently shown that both heparin and HS can act as potent inhibitors of p300 and pCAF HAT activities [181]. Notably, a marked reduction in the level of nuclear syndecan-1 was found following upregulation of heparanase in myeloma cells [88], possibly associated with nuclear localization of the enzyme [183–185]. This could lead to increased histone acetylation with an associated increase in gene transcription. In fact, heparanase-mediated loss of nuclear syndecan-1 enhances HAT activity to promote expression of genes that drive an aggressive tumor phenotype [186]. Heparanase regulation of gene expression may thus be related to its ability to inhibit accumulation of HSPGs within the nucleus. Hence, strategies to enhance nuclear HS levels may prove effective in blocking at least some of the heparanase-mediated effects that promote tumor growth and metastasis.
While most attention was addressed in recent years to heparanase function in tumor biology, emerging evidence indicate that heparanase is also engaged in several other pathological disorders. A most interesting example is the apparent role of heparanase in glomerular diseases [187, 188]. HSPGs are important constituent of the glomerular basement membrane and its permselective properties. Loss of HSPGs was observed in several experimental and human glomerulopathies, including diabetic nephropathy, minimal change disease, and membranous glomerulophathy [187]. In addition, expression of heparanase was up-regulated in the course of these diseases [189, 190], likely destructing the permselective properties of HS. Notably, PI-88 (a heparanase inhibitor) was effective as an antiproteinuric drug in experimental nephropathy model [191].
Upregulation of heparanase was reported in different inflammatory conditions such as inflammatory bowel disease [16], rheumatoid arthritis [17], and psoriasis (Lerner et al., our unpublished results), often associated with degradation of HS and release of chemokines anchored within the ECM network and cell surfaces. We have demonstrated that heparanase of epithelial origin induces abnormal activation of innate immune cells (i.e., macrophages) which, in turn, stimulate further production of the enzyme by the epithelial compartment, thereby preserving chronic inflammation and creating a tumor-promoting microenvironment [16]. Mechanistic features of this vicious cycle await further investigation.
Novel heparanase inhibitors such as non-anticoagulant glycol-split heparin (SST0001) [59, 192], PG545 [195] or rationally designed structure-based compounds are hoped to enter the clinic and relief patients’ condition. Compound SST0001 effectively inhibited myeloma growth in vivo, even when confronted with an aggressively growing tumor within human bone [192]. SST0001 also diminishes heparanase-induced shedding of syndecan-1, known to be a potent promoter of myeloma growth, indicating that SST0001 inhibits myeloma growth and angiogenesis via disruption of the heparanase/syndecan-1 axis [192]. Notably, inhibition of cathepsin L by serpin was markedly augmented by heparin and HS [193], suggesting that compound SST0001 will not only inhibit heparanase activity but also heparanase processing through cathepsin L inhibition [194]. A highly promising compound is PG545, a synthetic, fully sulfated HS mimetic that has recently entered Phase I trials for advanced cancer [195]. PG545 exhibits preclinical anti-tumor and anti-metastatic efficacy with a pharmacokinetic profile that supports less frequent dosing compared with other HS mimetics [195]. Ectopic miR-1258 was recently reported to suppress heparanase expression and activity, resulting in inhibition of experimental breast cancer brain metastasis [196]. Clearly, heparanase-inhibiting compounds are broad acting, targeting both the tumor and the tumor microenvironment (i.e., inflammation, angiogenesis) [194].
Although much has been learned in the last decade, the repertoire of heparanase functions in health and disease is only starting to emerge. Clearly, from activity implicated mainly in cell invasion associated with tumor metastasis, heparanase has turned into a multifaceted protein that appears to participate in essentially all major aspects of tumor progression, inflammation and kidney dysfunction.
Acknowledgements
We thank Prof. Benito Casu (‘Ronzoni’ Institute, Milan, Italy) and Prof. Ralph Sanderson (University of Alabama at Birmingham) for their continuous support and active collaboration. This work was supported by National Institutes of Health (NIH) grants CA106456 (IV) and CA138535 (RDS), the Israel Science Foundation (549/06); the Ministry of Science & Technology of the State of Israel and the German Cancer Research Center (DKFZ), the Juvenile Diabetes Research Foundation (JDRF 1-2006-695 and 38-2009-635) and by a research contract from Sigma-Tau Research Switzerland S.A. I. Vlodavsky is a Research Professor of the ICRF. We gratefully acknowledge the contribution, motivation and assistance of the research teams in the Hadassah-Hebrew University Medical Center (Jerusalem, Israel) and the Cancer and Vascular Biology Research Center of the Rappaport Faculty of Medicine (Technion, Haifa).
Conflict of Interest
The authors declare that they have no conflict of interest.
Glossary
ECM
Extracellular matrix
HS
Heparan sulfate
HSPGs
Heparan sulfate proteoglycans
GAG
Glycosaminoglycan
MMP
Matrix metalloproteinase
VEGF
Vascular endothelial growth factor
CAF
Cancer-associated fibroblasts
UC
Ulcerative colitis
IBD
Inflammatory bowel disease
AOM
Azoxymethane
DSS
Dextran sodium sulfate
IR
Ionoizing radiation
Egr1
Early growth response 1
CTL
Cytotoxic T lymphocyte
HAT
Histone acetyltransferase
References
- 1.Vlodavsky I, Gospodarowicz D. Respective roles of laminin and fibronectin in adhesion of human carcinoma and sarcoma cells. Nature. 1981;289:304–306. doi: 10.1038/289304a0. [DOI] [PubMed] [Google Scholar]
- 2.Vlodavsky I, Lui GM, Gospodarowicz D. Morphological appearance, growth behavior and migratory activity of human tumor cells maintained on extracellular matrix versus plastic. Cell. 1980;19:607–616. doi: 10.1016/s0092-8674(80)80037-7. [DOI] [PubMed] [Google Scholar]
- 3.Vlodavsky I (2001) Preparation of extracellular matrices produced by cultured corneal endothelial and PF-HR9 endodermal cells. Curr Protoc Cell Biol. Chapter 10:Unit 10.4 [DOI] [PubMed]
- 4.Kleinman HK, Martin GR. Matrigel: basement membrane matrix with biological activity. Semin Cancer Biol. 2005;15:378–386. doi: 10.1016/j.semcancer.2005.05.004. [DOI] [PubMed] [Google Scholar]
- 5.Xu R, Boudreau A, Bissell MJ. Tissue architecture and function: dynamic reciprocity via extra- and intra-cellular matrices. Cancer Metastasis Rev. 2009;28:167–176. doi: 10.1007/s10555-008-9178-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Folkman J, Klagsbrun M, Sasse J, Wadzinski M, Ingber D, Vlodavsky I. A heparin-binding angiogenic protein–basic fibroblast growth factor–is stored within basement membrane. Am J Pathol. 1988;130:393–400. [PMC free article] [PubMed] [Google Scholar]
- 7.Theocharis AD, Skandalis SS, Tzanakakis GN, Karamanos NK. Proteoglycans in health and disease: novel roles for proteoglycans in malignancy and their pharmacological targeting. FEBS J. 2010;277:3904–3923. doi: 10.1111/j.1742-4658.2010.07800.x. [DOI] [PubMed] [Google Scholar]
- 8.Iozzo RV, San Antonio JD. Heparan sulfate proteoglycans: heavy hitters in the angiogenesis arena. J Clin Invest. 2001;108:349–355. doi: 10.1172/JCI13738. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Kjellen L, Lindahl U. Proteoglycans: structures and interactions. Annu Rev Biochem. 1991;60:443–475. doi: 10.1146/annurev.bi.60.070191.002303. [DOI] [PubMed] [Google Scholar]
- 10.Bernfield M, Gotte M, Park PW, et al. Functions of cell surface heparan sulfate proteoglycans. Annu Rev Biochem. 1999;68:729–777. doi: 10.1146/annurev.biochem.68.1.729. [DOI] [PubMed] [Google Scholar]
- 11.Capila I, Linhardt RJ. Heparin-protein interactions. Angew Chem Int Ed Engl. 2002;41:391–412. doi: 10.1002/1521-3773(20020201)41:3<390::aid-anie390>3.0.co;2-b. [DOI] [PubMed] [Google Scholar]
- 12.Lindahl U, Li JP. Interactions between heparan sulfate and proteins-design and functional implications. Int Rev Cell Mol Biol. 2009;276:105–159. doi: 10.1016/S1937-6448(09)76003-4. [DOI] [PubMed] [Google Scholar]
- 13.Ogren S, Lindahl U. Cleavage of macromolecular heparin by an enzyme from mouse mastocytoma. J Biol Chem. 1975;250:2690–2697. [PubMed] [Google Scholar]
- 14.Thunberg L, Backstrom G, Wasteson A, Robinson HC, Ogren S, Lindahl U. Enzymatic depolymerization of heparin-related polysaccharides. Substrate specificities of mouse mastocytoma and human platelet endo-beta-D-glucuronidases. J Biol Chem. 1982;257:10278–10282. [PubMed] [Google Scholar]
- 15.Parish CR, Freeman C, Hulett MD. Heparanase: a key enzyme involved in cell invasion. Biochim Biophys Acta. 2001;1471:M99–M108. doi: 10.1016/s0304-419x(01)00017-8. [DOI] [PubMed] [Google Scholar]
- 16.Lerner I, Hermano E, Zcharia E, et al. Heparanase powers a chronic inflammatory circuit that promotes colitis-associated tumorigenesis in mice. J Clin Invest. 2011;121:1709–1721. doi: 10.1172/JCI43792. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Li RW, Freeman C, Yu D, et al. Dramatic regulation of heparanase activity and angiogenesis gene expression in synovium from patients with rheumatoid arthritis. Arthritis Rheum. 2008;58:1590–1600. doi: 10.1002/art.23489. [DOI] [PubMed] [Google Scholar]
- 18.Hoven MJ, Rops AL, Bakker MA, et al. Increased expression of heparanase in overt diabetic nephropathy. Kidney Int. 2006;70:2100–2108. doi: 10.1038/sj.ki.5001985. [DOI] [PubMed] [Google Scholar]
- 19.Lindahl U. Biosynthesis of heparin. Biochem Soc Trans. 1990;18:803–805. doi: 10.1042/bst0180803. [DOI] [PubMed] [Google Scholar]
- 20.Kreuger J, Spillmann D, Li JP, Lindahl U. Interactions between heparan sulfate and proteins: the concept of specificity. J Cell Biol. 2006;174:323–327. doi: 10.1083/jcb.200604035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Ledin J, Staatz W, Li JP, et al. Heparan sulfate structure in mice with genetically modified heparan sulfate production. J Biol Chem. 2004;279:42732–42741. doi: 10.1074/jbc.M405382200. [DOI] [PubMed] [Google Scholar]
- 22.Gray E, Mulloy B, Barrowcliffe TW. Heparin and low-molecular-weight heparin. Thromb Haemost. 2008;99:807–818. doi: 10.1160/TH08-01-0032. [DOI] [PubMed] [Google Scholar]
- 23.Lindahl U. Heparan sulfate-protein interactions–a concept for drug design? Thromb Haemost. 2007;98:109–115. [PubMed] [Google Scholar]
- 24.Fransson LA, Belting M, Cheng F, Jonsson M, Mani K, Sandgren S. Novel aspects of glypican glycobiology. Cell Mol Life Sci. 2004;61:1016–1024. doi: 10.1007/s00018-004-3445-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Cole GJ, Halfter W. Agrin: an extracellular matrix heparan sulfate proteoglycan involved in cell interactions and synaptogenesis. Perspect Dev Neurobiol. 1996;3:359–371. [PubMed] [Google Scholar]
- 26.Iozzo RV. Matrix proteoglycans: from molecular design to cellular function. Annu Rev Biochem. 1998;67:609–652. doi: 10.1146/annurev.biochem.67.1.609. [DOI] [PubMed] [Google Scholar]
- 27.Kramer KL, Yost HJ. Heparan sulfate core proteins in cell-cell signaling. Annu Rev Genet. 2003;37:461–484. doi: 10.1146/annurev.genet.37.061103.090226. [DOI] [PubMed] [Google Scholar]
- 28.Sasisekharan R, Shriver Z, Venkataraman G, Narayanasami U. Roles of heparan-sulphate glycosaminoglycans in cancer. Nat Rev Cancer. 2002;2:521–528. doi: 10.1038/nrc842. [DOI] [PubMed] [Google Scholar]
- 29.Simons M, Horowitz A. Syndecan-4-mediated signalling. Cell Signal. 2001;13:855–862. doi: 10.1016/s0898-6568(01)00190-5. [DOI] [PubMed] [Google Scholar]
- 30.Timpl R, Brown JC. Supramolecular assembly of basement membranes. Bioessays. 1996;18:123–132. doi: 10.1002/bies.950180208. [DOI] [PubMed] [Google Scholar]
- 31.Hacker U, Nybakken K, Perrimon N. Heparan sulphate proteoglycans: the sweet side of development. NatRev Mol Cell Biol. 2005;6:530–541. doi: 10.1038/nrm1681. [DOI] [PubMed] [Google Scholar]
- 32.Sanderson RD. Heparan sulfate proteoglycans in invasion and metastasis. Semin Cell Devel Biol. 2001;12:89–98. doi: 10.1006/scdb.2000.0241. [DOI] [PubMed] [Google Scholar]
- 33.Timar J, Lapis K, Dudas J, Sebestyen A, Kopper L, Kovalszky I. Proteoglycans and tumor progression: Janus-faced molecules with contradictory functions in cancer. Semin Cancer Biol. 2002;12:173–186. doi: 10.1016/S1044-579X(02)00021-4. [DOI] [PubMed] [Google Scholar]
- 34.Belting M. Heparan sulfate proteoglycan as a plasma membrane carrier. Trends Biochem Sci. 2003;28:145–151. doi: 10.1016/S0968-0004(03)00031-8. [DOI] [PubMed] [Google Scholar]
- 35.Bashkin P, Doctrow S, Klagsbrun M, Svahn CM, Folkman J, Vlodavsky I. Basic fibroblast growth factor binds to subendothelial extracellular matrix and is released by heparitinase and heparin-like molecules. Biochemistry. 1989;28:1737–1743. doi: 10.1021/bi00430a047. [DOI] [PubMed] [Google Scholar]
- 36.Patel VN, Knox SM, Likar KM, et al. Heparanase cleavage of perlecan heparan sulfate modulates FGF10 activity during ex vivo submandibular gland branching morphogenesis. Development. 2007;134:4177–4186. doi: 10.1242/dev.011171. [DOI] [PubMed] [Google Scholar]
- 37.Vlodavsky I, Bar-Shavit R, Ishai-Michaeli R, Bashkin P, Fuks Z. Extracellular sequestration and release of fibroblast growth factor: a regulatory mechanism? Trends Biochem Sci. 1991;16:268–271. doi: 10.1016/0968-0004(91)90102-2. [DOI] [PubMed] [Google Scholar]
- 38.Vlodavsky I, Miao HQ, Medalion B, Danagher P, Ron D. Involvement of heparan sulfate and related molecules in sequestration and growth promoting activity of fibroblast growth factor. Cancer Metastasis Rev. 1996;15:177–186. doi: 10.1007/BF00437470. [DOI] [PubMed] [Google Scholar]
- 39.Barash U, Cohen-Kaplan V, Dowek I, Sanderson RD, Ilan N, Vlodavsky I. Proteoglycans in health and disease: new concepts for heparanase function in tumor progression and metastasis. FEBS J. 2010;277:3890–3903. doi: 10.1111/j.1742-4658.2010.07799.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Ilan N, Elkin M, Vlodavsky I. Regulation, function and clinical significance of heparanase in cancer metastasis and angiogenesis. Int J Biochem Cell Biol. 2006;38:2018–2039. doi: 10.1016/j.biocel.2006.06.004. [DOI] [PubMed] [Google Scholar]
- 41.Levy-Adam F, Feld S, Cohen-Kaplan V, et al. Heparanase 2 interacts with heparan sulfate with high affinity and inhibits heparanase activity. J Biol Chem. 2010;285:28010–28019. doi: 10.1074/jbc.M110.116384. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.McKenzie E, Tyson K, Stamps A, et al. Cloning and expression profiling of Hpa2, a novel mammalian heparanase family member. Biochem Biophys Res Commun. 2000;276:1170–1177. doi: 10.1006/bbrc.2000.3586. [DOI] [PubMed] [Google Scholar]
- 43.Vlodavsky I, Eldor A, Haimovitz-Friedman A, et al. Expression of heparanase by platelets and circulating cells of the immune system: possible involvement in diapedesis and extravasation. Invasion Metastasis. 1992;12:112–127. [PubMed] [Google Scholar]
- 44.Elkin M, Ilan N, Ishai-Michaeli R, et al. Heparanase as mediator of angiogenesis: mode of action. FASEB J. 2001;15:1661–1663. doi: 10.1096/fj.00-0895fje. [DOI] [PubMed] [Google Scholar]
- 45.Levy-Adam F, Miao HQ, Heinrikson RL, Vlodavsky I, Ilan N. Heterodimer formation is essential for heparanase enzymatic activity. Biochem Biophys Res Commun. 2003;308:885–891. doi: 10.1016/s0006-291x(03)01478-5. [DOI] [PubMed] [Google Scholar]
- 46.McKenzie E, Young K, Hircock M, et al. Biochemical characterization of the active heterodimer form of human heparanase (Hpa1) protein expressed in insect cells. Biochem J. 2003;373:423–435. doi: 10.1042/BJ20030318. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Nardella C, Lahm A, Pallaoro M, Brunetti M, Vannini A, Steinkuhler C. Mechanism of activation of human heparanase investigated by protein engineering. Biochemistry. 2004;43:1862–1873. doi: 10.1021/bi030203a. [DOI] [PubMed] [Google Scholar]
- 48.Fux L, Ilan N, Sanderson RD, Vlodavsky I. Heparanase: busy at the cell surface. Trends Biochem Sci. 2009;34:511–519. doi: 10.1016/j.tibs.2009.06.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Hulett MD, Hornby JR, Ohms SJ, et al. Identification of active-site residues of the pro-metastatic endoglycosidase heparanase. Biochemistry. 2000;39:15659–15667. doi: 10.1021/bi002080p. [DOI] [PubMed] [Google Scholar]
- 50.Abboud-Jarrous G, Rangini-Guetta Z, Aingorn H, et al. Site-directed mutagenesis, proteolytic cleavage, and activation of human proheparanase. J Biol Chem. 2005;280:13568–13575. doi: 10.1074/jbc.M413370200. [DOI] [PubMed] [Google Scholar]
- 51.Abboud-Jarrous G, Atzmon R, Peretz T, et al. Cathepsin L is responsible for processing and activation of proheparanase through multiple cleavages of a linker segment. J Biol Chem. 2008;283:18167–18176. doi: 10.1074/jbc.M801327200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Nakajima M, Irimura T, DiFerrante D, DiFerrante N, Nicolson GL. Heparan sulfate degradation: relation to tumor invasion and metastatic properties of Mouse B 16 Melanoma sublines. Science. 1983;220:611–613. doi: 10.1126/science.6220468. [DOI] [PubMed] [Google Scholar]
- 53.Vlodavsky I, Fuks Z, Bar-Ner M, Ariav Y, Schirrmacher V. Lymphoma cells mediated degradation of sulfated proteoglycans in the subendothelial extracellular matrix: relation to tumor cell metastasis. Cancer Res. 1983;43:2704–2711. [PubMed] [Google Scholar]
- 54.Cohen I, Pappo O, Elkin M, et al. Heparanase promotes growth, angiogenesis and survival of primary breast tumors. Int J Cancer. 2006;118:1609–1617. doi: 10.1002/ijc.21552. [DOI] [PubMed] [Google Scholar]
- 55.Lerner I, Baraz L, Pikarsky E, et al. Function of heparanase in prostate tumorigenesis: potential for therapy. Clin Cancer Res. 2008;14:668–676. doi: 10.1158/1078-0432.CCR-07-1866. [DOI] [PubMed] [Google Scholar]
- 56.Edovitsky E, Elkin M, Zcharia E, Peretz T, Vlodavsky I. Heparanase gene silencing, tumor invasiveness, angiogenesis, and metastasis. J Natl Cancer Inst. 2004;96:1219–1230. doi: 10.1093/jnci/djh230. [DOI] [PubMed] [Google Scholar]
- 57.Vreys V, David G. Mammalian heparanase: what is the message? J Cell Mol Med. 2007;11:427–452. doi: 10.1111/j.1582-4934.2007.00039.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Shafat I, Ben-Arush MW, Issakov J, Meller I, Naroditsky I, Tortoteto M, Cassinelli G, Lanzi C, Pisano C, Ilan N, Vlodavsky I, Zunino F (2011) J cell mol med (PMID: 21029368) [DOI] [PMC free article] [PubMed]
- 59.Vlodavsky I, Ilan N, Naggi A, Casu B. Heparanase: structure, biological functions, and inhibition by heparin-derived mimetics of heparan sulfate. Curr Pharm Des. 2007;13:2057–2073. doi: 10.2174/138161207781039742. [DOI] [PubMed] [Google Scholar]
- 60.Casu B, Guerrini M, Guglieri S, et al. Undersulfated and glycol-split heparins endowed with antiangiogenic activity. J Med Chem. 2004;47:838–848. doi: 10.1021/jm030893g. [DOI] [PubMed] [Google Scholar]
- 61.Ferro V, Hammond E, Fairweather JK. The development of inhibitors of heparanase, a key enzyme involved in tumour metastasis, angiogenesis and inflammation. Mini Rev Med Chem. 2004;4:693–702. doi: 10.2174/1389557043403729. [DOI] [PubMed] [Google Scholar]
- 62.McKenzie EA. Heparanase: a target for drug discovery in cancer and inflammation. Br J Pharmacol. 2007;151:1–14. doi: 10.1038/sj.bjp.0707182. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Miao HQ, Liu H, Navarro E, Kussie P, Zhu Z. Development of heparanase inhibitors for anti-cancer therapy. Curr Med Chem. 2006;13:2101–2111. doi: 10.2174/092986706777935230. [DOI] [PubMed] [Google Scholar]
- 64.Doweck I, Kaplan-Cohen V, Naroditsky I, Sabo E, Ilan N, Vlodavsky I. Neoplasia. 2006;8:1055–1061. doi: 10.1593/neo.06577. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.El-Assal ON, Yamanoi A, Ono T, Kohno H, Nagasue N. Clin Cancer Res. 2001;7(5):1299–1305. [PubMed] [Google Scholar]
- 66.Maxhimer JB, Quiros RM, Stewart R, Dowlatshahi K, Gattuso P, Fan M, Prinz RA, Xu X. Surgery. 2002;132:326–333. doi: 10.1067/msy.2002.125719. [DOI] [PubMed] [Google Scholar]
- 67.Nagler R, Ben-Izhak O, Cohen-Kaplan V, Shafat I, Vlodavsky I, Akrish S, Ilan N. Cancer. 2007;110:2732–2739. doi: 10.1002/cncr.23095. [DOI] [PubMed] [Google Scholar]
- 68.Tang W, Nakamura Y, Tsujimoto M, Sato M, Wang X, Kurozumi K, Nakahara M, Nakao K, Nakamura M, Mori I, Kakudo K. Mod Pathol. 2002;15:593–598. doi: 10.1038/modpathol.3880571. [DOI] [PubMed] [Google Scholar]
- 69.Barash U, Cohen-Kaplan V, Arvatz G, et al. A novel human heparanase splice variant, T5, endowed with protumorigenic characteristics. FASEB J. 2010;24:1239–1248. doi: 10.1096/fj.09-147074. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Doviner V, Maly B, Kaplan V, et al. Spatial and temporal heparanase expression in colon mucosa throughout the adenoma-carcinoma sequence. Mod Pathol. 2006;19:878–888. doi: 10.1038/modpathol.3800603. [DOI] [PubMed] [Google Scholar]
- 71.Yang Y, Macleod V, Bendre M, et al. Heparanase promotes the spontaneous metastasis of myeloma cells to bone. Blood. 2005;105:1303–1309. doi: 10.1182/blood-2004-06-2141. [DOI] [PubMed] [Google Scholar]
- 72.Zetser A, Bashenko Y, Miao H-Q, Vlodavsky I, Ilan N. Heparanase affects adhesive and tumorigenic potential of human glioma cells. Cancer Res. 2003;63:7733–7741. [PubMed] [Google Scholar]
- 73.Vlodavsky I, Korner G, Ishai-Michaeli R, Bashkin P, Bar-Shavit R, Fuks Z. Extracellular matrix-resident growth factors and enzymes: possible involvement in tumor metastasis and angiogenesis. Cancer Metastasis Rev. 1990;9:203–226. doi: 10.1007/BF00046361. [DOI] [PubMed] [Google Scholar]
- 74.Witz IP. Tumor-microenvironment interactions: dangerous liaisons. Adv Cancer Res. 2008;100:203–229. doi: 10.1016/S0065-230X(08)00007-9. [DOI] [PubMed] [Google Scholar]
- 75.Ruoslahti E. How cancer spreads. Sci Am. 1996;275:72–77. doi: 10.1038/scientificamerican0996-72. [DOI] [PubMed] [Google Scholar]
- 76.Liotta LA, Kohn EC. The microenvironment of the tumour-host interface. Nature. 2001;411:375–379. doi: 10.1038/35077241. [DOI] [PubMed] [Google Scholar]
- 77.Kessenbrock K, Plaks V, Werb Z. Matrix metalloproteinases: regulators of the tumor microenvironment. Cell. 2010;141:52–67. doi: 10.1016/j.cell.2010.03.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Mason SD, Joyce JA. Proteolytic networks in cancer. Trends Cell Biol. 2011;21:228–237. doi: 10.1016/j.tcb.2010.12.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Stupack DG, Cheresh DA. Integrins and angiogenesis. Curr Top Dev Biol. 2004;64:207–238. doi: 10.1016/S0070-2153(04)64009-9. [DOI] [PubMed] [Google Scholar]
- 80.Noel A, Jost M, Maquoi E. Matrix metalloproteinases at cancer tumor-host interface. Sem Cell Dev Biol. 2008;19:52–60. doi: 10.1016/j.semcdb.2007.05.011. [DOI] [PubMed] [Google Scholar]
- 81.Damme J, Struyf S, Opdenakker G. Chemokine-protease interactions in cancer. Semin Cancer Biol. 2004;14:201–208. doi: 10.1016/j.semcancer.2003.10.007. [DOI] [PubMed] [Google Scholar]
- 82.Gocheva V, Joyce JA. Cysteine cathepsins and the cutting edge of cancer invasion. Cell Cycle. 2007;6:60–64. doi: 10.4161/cc.6.1.3669. [DOI] [PubMed] [Google Scholar]
- 83.Lankelma JM, Voorend DM, Barwari T, et al. Cathepsin L, target in cancer treatment? Life Sci. 2010;86:225–233. doi: 10.1016/j.lfs.2009.11.016. [DOI] [PubMed] [Google Scholar]
- 84.Turk B, Turk D, Turk V. Lysosomal cysteine proteases: more than scavengers. Biochim Biophys Acta. 2000;1477:98–111. doi: 10.1016/s0167-4838(99)00263-0. [DOI] [PubMed] [Google Scholar]
- 85.Bix G, Iozzo RV. Matrix revolutions: “tails” of basement-membrane components with angiostatic functions. Trends Cell Biol. 2005;15:52–60. doi: 10.1016/j.tcb.2004.11.008. [DOI] [PubMed] [Google Scholar]
- 86.Purushothaman A, Chen L, Yang Y, Sanderson RD. Heparanase stimulation of protease expression implicates it as a master regulator of the aggressive tumor phenotype in myeloma. J Biol Chem. 2008;283:32628–32636. doi: 10.1074/jbc.M806266200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Zcharia E, Jia J, Zhang X, et al. Newly generated heparanase knock-out mice unravel co-regulation of heparanase and matrix metalloproteinases. PLoS ONE. 2009;4:e5181. doi: 10.1371/journal.pone.0005181. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Chen L, Sanderson RD. Heparanase regulates levels of syndecan-1 in the nucleus. PLoS ONE. 2009;4(3):e4947. doi: 10.1371/journal.pone.0004947. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Paget S. The distribution of a secondary growths in cancer of the breast. Lancet. 1889;133:571–583. [PubMed] [Google Scholar]
- 90.Talmadge JE, Fidler IJ. AACR centennial series: the biology of cancer metastasis: historical perspective. Cancer Res. 2010;70:5649–5669. doi: 10.1158/0008-5472.CAN-10-1040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Nicolson GL. Organ specificity of tumor metastasis: role of preferential adhesion, invasion and growth of malignant cells at specific secondary sites. Cancer Metastasis Rev. 1988;7:143–188. doi: 10.1007/BF00046483. [DOI] [PubMed] [Google Scholar]
- 92.Kaplan RN, Psaila B, Lyden D. Bone marrow cells in the ‘pre-metastatic niche’: within bone and beyond. Cancer Metastasis Rev. 2006;25:521–529. doi: 10.1007/s10555-006-9036-9. [DOI] [PubMed] [Google Scholar]
- 93.Kaplan RN, Rafii S, Lyden D. Preparing the “soil”: the premetastatic niche. Cancer Res. 2006;66:11089–11093. doi: 10.1158/0008-5472.CAN-06-2407. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Kaplan RN, Riba RD, Zacharoulis S, et al. VEGFR1-positive haematopoietic bone marrow progenitors initiate the pre-metastatic niche. Nature. 2005;438:820–827. doi: 10.1038/nature04186. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Ferrara N, Hillan KJ, Gerber HP, Novotny W. Discovery and development of bevacizumab, an anti-VEGF antibody for treating cancer. Nat Rev Drug Discov. 2004;3:391–400. doi: 10.1038/nrd1381. [DOI] [PubMed] [Google Scholar]
- 96.Franco OE, Shaw AK, Strand DW, Hayward SW. Cancer associated fibroblasts in cancer pathogenesis. Sem Cell Dev Biol. 2010;21:33–39. doi: 10.1016/j.semcdb.2009.10.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Shimoda M, Mellody KT, Orimo A. Carcinoma-associated fibroblasts are a rate-limiting determinant for tumour progression. Sem cell Dev Biol. 2010;221:19–25. doi: 10.1016/j.semcdb.2009.10.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Bhowmick NA, Chytil A, Plieth D, et al. TGF-beta signaling in fibroblasts modulates the oncogenic potential of adjacent epithelia. Science. 2004;303:848–851. doi: 10.1126/science.1090922. [DOI] [PubMed] [Google Scholar]
- 99.Bhowmick NA, Neilson EG, Moses HL. Stromal fibroblasts in cancer initiation and progression. Nature. 2004;432:332–337. doi: 10.1038/nature03096. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Kalluri R, Zeisberg M. Fibroblasts in cancer. Nat Rev Cancer. 2006;6:392–401. doi: 10.1038/nrc1877. [DOI] [PubMed] [Google Scholar]
- 101.Erez N, Truitt M, Olson P, Arron ST, Hanahan D. Cancer-associated fibroblasts are activated in incipient neoplasia to orchestrate tumor-promoting inflammation in an NF-kappaB-dependent manner. Cancer Cell. 2010;17:135–147. doi: 10.1016/j.ccr.2009.12.041. [DOI] [PubMed] [Google Scholar]
- 102.Grivennikov SI, Greten FR, Karin M. Immunity, inflammation, and cancer. Cell. 2010;140:883–899. doi: 10.1016/j.cell.2010.01.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Mantovani A, Sica A. Macrophages, innate immunity and cancer: balance, tolerance, and diversity. Cur Opin Immunol. 2010;22:231–237. doi: 10.1016/j.coi.2010.01.009. [DOI] [PubMed] [Google Scholar]
- 104.Condeelis J, Pollard JW. Macrophages: obligate partners for tumor cell migration, invasion, and metastasis. Cell. 2006;124:263–266. doi: 10.1016/j.cell.2006.01.007. [DOI] [PubMed] [Google Scholar]
- 105.Dirkx AE, Oude Egbrink MG, Wagstaff J, Griffioen AW. Monocyte/macrophage infiltration in tumors: modulators of angiogenesis. J Leukoc Biol. 2006;80:1183–1196. doi: 10.1189/jlb.0905495. [DOI] [PubMed] [Google Scholar]
- 106.Pollard JW. Tumour-educated macrophages promote tumour progression and metastasis. Nat Rev Cancer. 2004;4:71–78. doi: 10.1038/nrc1256. [DOI] [PubMed] [Google Scholar]
- 107.Edovitsky E, Lerner I, Zcharia E, Peretz T, Vlodavsky I, Elkin M. Role of endothelial heparanase in delayed-type hypersensitivity. Blood. 2006;107:3609–3616. doi: 10.1182/blood-2005-08-3301. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.Waterman M, Ben-Izhak O, Eliakim R, Groisman G, Vlodavsky I, Ilan N. Heparanase upregulation by colonic epithelium in inflammatory bowel disease. Mod Pathol. 2007;20:8–14. doi: 10.1038/modpathol.3800710. [DOI] [PubMed] [Google Scholar]
- 109.Clevers H. Colon cancer–understanding how NSAIDs work. N Engl J Med. 2006;354:761–763. doi: 10.1056/NEJMcibr055457. [DOI] [PubMed] [Google Scholar]
- 110.Karin M, Greten FR. NF-kappaB: linking inflammation and immunity to cancer development and progression. Nat Rev Immunol. 2005;5:749–759. doi: 10.1038/nri1703. [DOI] [PubMed] [Google Scholar]
- 111.Greten FR, Eckmann L, Greten TF, et al. IKKbeta links inflammation and tumorigenesis in a mouse model of colitis-associated cancer. Cell. 2004;118:285–296. doi: 10.1016/j.cell.2004.07.013. [DOI] [PubMed] [Google Scholar]
- 112.Popivanova BK, Kitamura K, Wu Y, et al. Blocking TNF-alpha in mice reduces colorectal carcinogenesis associated with chronic colitis. J Clin Invest. 2008;118:560–570. doi: 10.1172/JCI32453. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Okayasu I, Ohkusa T, Kajiura K, Kanno J, Sakamoto S. Promotion of colorectal neoplasia in experimental murine ulcerative colitis. Gut. 1996;39:87–92. doi: 10.1136/gut.39.1.87. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Karin M. Nuclear factor-kappaB in cancer development and progression. Nature. 2006;441:431–436. doi: 10.1038/nature04870. [DOI] [PubMed] [Google Scholar]
- 115.Yu H, Pardoll D, Jove R. STATs in cancer inflammation and immunity: a leading role for STAT3. Nat Rev Cancer. 2009;9:798–809. doi: 10.1038/nrc2734. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 116.Okayasu I, Hatakeyama S, Yamada M, Ohkusa T, Inagaki Y, Nakaya R. A novel method in the induction of reliable experimental acute and chronic ulcerative colitis in mice. Gastroenterology. 1990;98:694–702. doi: 10.1016/0016-5085(90)90290-h. [DOI] [PubMed] [Google Scholar]
- 117.Zcharia E, Metzger S, Chajek-ShaulL T, et al. Transgenic expression of mammalian heparanase uncovers physiological functions of heparan sulfate in tissue morphogenesis, vascularization, and feeding behavior. FASEB J. 2004;18:252–263. doi: 10.1096/fj.03-0572com. [DOI] [PubMed] [Google Scholar]
- 118.Mahida YR. The key role of macrophages in the immunopathogenesis of inflammatory bowel disease. Inflamm Bowel Dis. 2000;6:21–33. doi: 10.1097/00054725-200002000-00004. [DOI] [PubMed] [Google Scholar]
- 119.Sanchez-Munoz F, Dominguez-Lopez A, Yamamoto-Furusho JK. Role of cytokines in inflammatory bowel disease. World J Gastroenterol. 2008;14:4280–4288. doi: 10.3748/wjg.14.4280. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120.Elson CO, Sartor RB, Tennyson GS, Riddell RH. Experimental models of inflammatory bowel disease. Gastroenterology. 1995;109:1344–1367. doi: 10.1016/0016-5085(95)90599-5. [DOI] [PubMed] [Google Scholar]
- 121.Krieglstein CF, Cerwinka WH, Sprague AG, et al. Collagen-binding integrin alpha1beta1 regulates intestinal inflammation in experimental colitis. J Clin Invest. 2002;110:1773–1782. doi: 10.1172/JCI200215256. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 122.Coussens LM, Fingleton B, Matrisian LM. Matrix metalloproteinase inhibitors and cancer: trials and tribulations. Science. 2002;295:2387–2392. doi: 10.1126/science.1067100. [DOI] [PubMed] [Google Scholar]
- 123.Brunn GJ, Bungum MK, Johnson GB, Platt JL. Conditional signaling by Toll-like receptor 4. FASEB J. 2005;19:872–874. doi: 10.1096/fj.04-3211fje. [DOI] [PubMed] [Google Scholar]
- 124.Fukata M, Chen A, Vamadevan AS, et al. Toll-like receptor-4 promotes the development of colitis-associated colorectal tumors. Gastroenterology. 2007;133:1869–1881. doi: 10.1053/j.gastro.2007.09.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 125.Fukata M, Hernandez Y, Conduah D, et al. Innate immune signaling by Toll-like receptor-4 (TLR4) shapes the inflammatory microenvironment in colitis-associated tumors. Inflamm Bowel Dis. 2009;15:997–1006. doi: 10.1002/ibd.20880. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 126.Kumar V, Abbas A, Fausto N, (eds) (2005) Pathologic basis of disease: Elsevier Saunders
- 127.Mestre AM, Rao S, Hornby JR, Soe-Htwe T, Khachigian LM, Hulett MD. Early Growth Response gene 1 (EGR1) regulates heparanase gene transcription in tumor cells. J Biol Chem. 2005;280:35136–35147. doi: 10.1074/jbc.M503414200. [DOI] [PubMed] [Google Scholar]
- 128.Subbaramaiah K, Yoshimatsu K, Scherl E, et al. Microsomal prostaglandin E synthase-1 is overexpressed in inflammatory bowel disease. Evidence for involvement of the transcription factor Egr-1. J Biol Chem. 2004;279:12647–12658. doi: 10.1074/jbc.M312972200. [DOI] [PubMed] [Google Scholar]
- 129.Fiebiger E, Maehr R, Villadangos J, et al. Invariant chain controls the activity of extracellular cathepsin L. J Exp Med. 2002;196:1263–1269. doi: 10.1084/jem.20020762. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 130.Bouma G, Strober W. The immunological and genetic basis of inflammatory bowel disease. Nat Rev Immunol. 2003;3:521–533. doi: 10.1038/nri1132. [DOI] [PubMed] [Google Scholar]
- 131.Borja-Cacho D, Jensen EH, Saluja AK, Buchsbaum DJ, Vickers SM. Molecular targeted therapies for pancreatic cancer. Am J Surg. 2008;196:430–441. doi: 10.1016/j.amjsurg.2008.04.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 132.Raimondi S, Maisonneuve P, Lowenfels AB. Epidemiology of pancreatic cancer: an overview. Nat Rev Gastroenterol Hepatol. 2009;6:699–708. doi: 10.1038/nrgastro.2009.177. [DOI] [PubMed] [Google Scholar]
- 133.Shaib YH, Davila JA, El-Serag HB. The epidemiology of pancreatic cancer in the United States: changes below the surface. Aliment Pharmacol Ther. 2006;24:87–94. doi: 10.1111/j.1365-2036.2006.02961.x. [DOI] [PubMed] [Google Scholar]
- 134.Muller MW, Friess H, Koninger J, et al. Factors influencing survival after bypass procedures in patients with advanced pancreatic adenocarcinomas. Am J Surg. 2008;195:221–228. doi: 10.1016/j.amjsurg.2007.02.026. [DOI] [PubMed] [Google Scholar]
- 135.Network NCC (2008) NCCN clinical practice guidelines in oncology: pancreatic adenocarcinoma. v.1
- 136.Cohen SJ, Dobelbower R, Jr, Lipsitz S, et al. A randomized phase III study of radiotherapy alone or with 5-fluorouracil and mitomycin-C in patients with locally advanced adenocarcinoma of the pancreas: Eastern Cooperative Oncology Group study E8282. Int J Radiat Oncol Biol Phys. 2005;62:1345–1350. doi: 10.1016/j.ijrobp.2004.12.074. [DOI] [PubMed] [Google Scholar]
- 137.Neoptolemos JP, Stocken DD, Friess H, et al. A randomized trial of chemoradiotherapy and chemotherapy after resection of pancreatic cancer. N Engl J Med. 2004;350:1200–1210. doi: 10.1056/NEJMoa032295. [DOI] [PubMed] [Google Scholar]
- 138.Camphausen K, Moses MA, Beecken W-D, Khan MK, Folkman J, O’Reilly MS. Radiation therapy to a primary tumor accelerates metastatic growth in mice. Cancer Res. 2001;61:2207–2211. [PubMed] [Google Scholar]
- 139.Kaliski A, Maggiorella L, Cengel KA, et al. Angiogenesis and tumor growth inhibition by a matrix metalloproteinase inhibitor targeting radiation-induced invasion. Mol Cancer Ther. 2005;4:1717–1728. doi: 10.1158/1535-7163.MCT-05-0179. [DOI] [PubMed] [Google Scholar]
- 140.Madani I, Neve W, Mareel M. Does ionizing radiation stimulate cancer invasion and metastasis? Bull Cancer. 2008;95:292–300. doi: 10.1684/bdc.2008.0598. [DOI] [PubMed] [Google Scholar]
- 141.Ohuchida K, Mizumoto K, Murakami M, et al. Radiation to stromal fibroblasts increases invasiveness of pancreatic cancer cells through tumor-stromal interactions. Cancer Res. 2004;64:3215–3222. doi: 10.1158/0008-5472.can-03-2464. [DOI] [PubMed] [Google Scholar]
- 142.Park CM, Park MJ, Kwak HJ, et al. Ionizing radiation enhances matrix metalloproteinase-2 secretion and invasion of glioma cells through Src/epidermal growth factor receptor-mediated p38/Akt and phosphatidylinositol 3-kinase/Akt signaling pathways. Cancer Res. 2006;66:8511–8519. doi: 10.1158/0008-5472.CAN-05-4340. [DOI] [PubMed] [Google Scholar]
- 143.Qian L-W, Mizumoto K, Urashima T, et al. Radiation-induced increase in invasive potential of human pancreatic cancer cells and its blockade by a matrix metalloproteinase inhibitor, CGS27023. Clin Cancer Res. 2002;8:1223–1227. [PubMed] [Google Scholar]
- 144.Wild-Bode C, Weller M, Rimner A, Dichgans J, Wick W. Sublethal irradiation promotes migration and invasiveness of glioma cells: implications for radiotherapy of human glioblastoma. Cancer Res. 2001;61:2744–2750. [PubMed] [Google Scholar]
- 145.Meirovitz A, Hermano E, Lerner I, et al. Role of heparanase in radiation-enhanced invasiveness of pancreatic carcinoma. Cancer Res. 2011;71:2772–2780. doi: 10.1158/0008-5472.CAN-10-3402. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 146.Koliopanos A, Friess H, Kleeff J, Shi X, Liao Q, Pecker I, Vlodavsky I, Zimmermann A, Buchler MW. Cancer Res. 2001;61:4655–4659. [PubMed] [Google Scholar]
- 147.Quiros RM, Rao G, Plate J, et al. Elevated serum heparanase-1 levels in patients with pancreatic carcinoma are associated with poor survival. Cancer. 2006;106:532–540. doi: 10.1002/cncr.21648. [DOI] [PubMed] [Google Scholar]
- 148.Rohloff J, Zinke J, Schoppmeyer K, Tannapfel A, Witzigmann H, Mossner J, Wittekind C, Caca K. Br J Cancer. 2002;86:1270–1275. doi: 10.1038/sj.bjc.6600232. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 149.Hoffmann AC, Mori R, Vallbohmer D, et al. High expression of heparanase is significantly associated with dedifferentiation and lymph node metastasis in patients with pancreatic ductal adenocarcinomas and correlated to PDGFA and via HIF1a to HB-EGF and bFGF. J Gastrointest Surg. 2008;12:1674–1681. doi: 10.1007/s11605-008-0628-2. [DOI] [PubMed] [Google Scholar]
- 150.Mestre AM, Khachigian LM, Santiago FS, Staykova MA, Hulett MD. Regulation of inducible heparanase gene transcription in activated T cells by early growth response 1. J Biol Chem. 2003;278:50377–50385. doi: 10.1074/jbc.M310154200. [DOI] [PubMed] [Google Scholar]
- 151.Mestre AM, Staykova MA, Hornby JR, Willenborg DO, Hulett MD. Expression of the heparan sulfate-degrading enzyme heparanase is induced in infiltrating CD4+ T cells in experimental autoimmune encephalomyelitis and regulated at the level of transcription by early growth response gene 1. J Leukoc Biol. 2007;82:1289–1300. doi: 10.1189/jlb.0507315. [DOI] [PubMed] [Google Scholar]
- 152.Srinivasan R, Mager GM, Ward RM, Mayer J, Svaren J. NAB2 represses transcription by interacting with the CHD4 subunit of the nucleosome remodeling and deacetylase (NuRD) complex. J Biol Chem. 2006;281:15129–15137. doi: 10.1074/jbc.M600775200. [DOI] [PubMed] [Google Scholar]
- 153.Casu B, Guerrini M, Torri G. Structural and conformational aspects of the anticoagulant and anti-thrombotic activity of heparin and dermatan sulfate. Curr Pharm Des. 2004;10:939–949. doi: 10.2174/1381612043452794. [DOI] [PubMed] [Google Scholar]
- 154.Casu B, Vlodavsky I, Sanderson RD. Non-anticoagulant heparins and inhibition of cancer. Pathophysiol Haemost Thromb. 2008;36:195–203. doi: 10.1159/000175157. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 155.Naggi A, Casu B, Perez M, et al. Modulation of the heparanase-inhibiting activity of heparin through selective desulfation, graded N-acetylation, and glycol splitting. J Biol Chem. 2005;280:12103–12113. doi: 10.1074/jbc.M414217200. [DOI] [PubMed] [Google Scholar]
- 156.Chung SI, Seong J, Park YN, Kim WW, Oh HJ, Han KH. Identification of proteins indicating radiation-induced hepatic toxicity in cirrhotic rats. J Rsd Res 51:643–650 [DOI] [PubMed]
- 157.Iriyama S, Matsunaga Y, Takahashi K, Matsuzaki K, Kumagai N, Amano S. Activation of heparanase by ultraviolet B irradiation leads to functional loss of basement membrane at the dermal-epidermal junction in human skin. Arch Dermatol Res. 2011;303:253–261. doi: 10.1007/s00403-010-1117-5. [DOI] [PubMed] [Google Scholar]
- 158.Chen T, Tang XD, Wan Y, et al. HLA-A2-restricted cytotoxic T lymphocyte epitopes from human heparanase as novel targets for broad-spectrum tumor immunotherapy. Neoplasia. 2008;10:977–986. doi: 10.1593/neo.08576. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 159.Tang W, Nakamura Y, Tsujimoto M, et al. Heparanase: a key enzyme in invasion and metastasis of gastric carcinoma. Mod Pathol. 2002;15:593–598. doi: 10.1038/modpathol.3880571. [DOI] [PubMed] [Google Scholar]
- 160.Tang XD, Liang GP, Li C, et al. Cytotoxic T lymphocyte epitopes from human heparanase can elicit a potent anti-tumor immune response in mice. Cancer Immunol Immunother. 2010;59:1041–1047. doi: 10.1007/s00262-010-0829-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 161.Wang GZ, Tang XD, Lu MH_,_ et al (2011) Multiple antigenic peptides of human heparanase elicit a much more potent immune response against tumors. Canc Prev Res (PMID: 21505182) [DOI] [PubMed]
- 162.Beckhove P, Feuerer M, Dolenc M, et al. Specifically activated memory T cell subsets from cancer patients recognize and reject xenotransplanted autologous tumors. J Clin Invest. 2004;114:67–76. doi: 10.1172/JCI20278. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 163.Feuerer M, Beckhove P, Garbi N, et al. Bone marrow as a priming site for T-cell responses to blood-borne antigen. Nat Med. 2003;9:1151–1157. doi: 10.1038/nm914. [DOI] [PubMed] [Google Scholar]
- 164.Khazaie K, Prifti S, Beckhove P, et al. Persistence of dormant tumor cells in the bone marrow of tumor cell-vaccinated mice correlates with long-term immunological protection. Proc Natl Acad Sci USA. 1994;91:7430–7434. doi: 10.1073/pnas.91.16.7430. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 165.Schirrmacher V, Feuerer M, Fournier P, Ahlert T, Umansky V, Beckhove P. T-cell priming in bone marrow: the potential for long-lasting protective anti-tumor immunity. Trends Mol Med. 2003;9:526–534. doi: 10.1016/j.molmed.2003.10.001. [DOI] [PubMed] [Google Scholar]
- 166.Bai L, Beckhove P, Feuerer M, et al. Cognate interactions between memory T cells and tumor antigen-presenting dendritic cells from bone marrow of breast cancer patients: bidirectional cell stimulation, survival and antitumor activity in vivo. Int J Cancer. 2003;103:73–83. doi: 10.1002/ijc.10781. [DOI] [PubMed] [Google Scholar]
- 167.Schmitz-Winnenthal FH, Volk C, Z’Graggen K, et al. High frequencies of functional tumor-reactive T cells in bone marrow and blood of pancreatic cancer patients. Cancer Res. 2005;65:10079–10087. doi: 10.1158/0008-5472.CAN-05-1098. [DOI] [PubMed] [Google Scholar]
- 168.Wagner P, Koch M, Nummer D, et al. Detection and functional analysis of tumor infiltrating T-lymphocytes (TIL) in liver metastases from colorectal cancer. Ann Surg Oncol. 2008;15:2310–2317. doi: 10.1245/s10434-008-9971-5. [DOI] [PubMed] [Google Scholar]
- 169.Schuetz F, Ehlert K, Ge Y, et al. Treatment of advanced metastasized breast cancer with bone marrow-derived tumour-reactive memory T cells: a pilot clinical study. Cancer Immunol Immunother. 2008;58:887–900. doi: 10.1007/s00262-008-0605-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 170.Sommerfeldt N, Beckhove P, Ge Y, et al. Heparanase: a new metastasis-associated antigen recognized in breast cancer patients by spontaneously induced memory T lymphocytes. Cancer Res. 2006;66:7716–77123. doi: 10.1158/0008-5472.CAN-05-2363. [DOI] [PubMed] [Google Scholar]
- 171.Bonertz A, Weitz J, Pietsch DH, et al. Antigen-specific Tregs control T cell responses against a limited repertoire of tumor antigens in patients with colorectal carcinoma. J Clin Invest. 2009;119:3311–3321. doi: 10.1172/JCI39608. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 172.Schuetz F, Ehlert K, Ge Y, et al. Treatment of advanced metastasized breast cancer with bone marrow-derived tumour-reactive memory T cells: a pilot clinical study. Cancer Immunol Immunother. 2009;58:887–900. doi: 10.1007/s00262-008-0605-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 173.Levy-Adam F, Ilan N, Vlodavsky I. Tumorigenic and adhesive properties of heparanase. Semin Cancer Biol. 2010;20:153–160. doi: 10.1016/j.semcancer.2010.06.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 174.Levy-Adam F, Feld S, Suss-Toby E, Vlodavsky I, Ilan N. Heparanase facilitates cell adhesion and spreading by clustering of cell surface heparan sulfate proteoglycans. PLoS ONE. 2008;3:e2319. doi: 10.1371/journal.pone.0002319. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 175.Kim MY, Oskarsson T, Acharyya S, et al. Tumor self-seeding by circulating cancer cells. Cell. 2009;139:1315–1326. doi: 10.1016/j.cell.2009.11.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 176.Leung CT, Brugge JS. Tumor self-seeding: bidirectional flow of tumor cells. Cell. 2009;139:1226–1228. doi: 10.1016/j.cell.2009.12.013. [DOI] [PubMed] [Google Scholar]
- 177.Joyce JA, Freeman C, Meyer-Morse N, Parish CR, Hanahan D. A functional heparan sulfate mimetic implicates both heparanase and heparan sulfate in tumor angiogenesis and invasion in a mouse model of multistage cancer. Oncogene. 2005;24:4037–4051. doi: 10.1038/sj.onc.1208602. [DOI] [PubMed] [Google Scholar]
- 178.Friedmann Y, Vlodavsky I, Aingorn H, Aviv A, Peretz T, Pecker I, Pappo O. Am J Pathol. 2000;157(4):1167–1175. doi: 10.1016/S0002-9440(10)64632-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 179.Brun R, Naroditsky I, Waterman M, et al. Heparanase expression by Barrett’s epithelium and during esophageal carcinoma progression. Mod Pathol. 2009;22:1548–1554. doi: 10.1038/modpathol.2009.115. [DOI] [PubMed] [Google Scholar]
- 180.Cohen-Kaplan V, Naroditsky I, Zetser A, Ilan N, Vlodavsky I, Doweck I. Heparanase induces VEGF C and facilitates tumor lymphangiogenesis. Int J Cancer. 2008;123:2566–2573. doi: 10.1002/ijc.23898. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 181.Buczek-Thomas JA, Hsia E, Rich CB, Foster JA, Nugent MA. Inhibition of histone acetyltransferase by glycosaminoglycans. J Cell Biochem. 2008;105:108–120. doi: 10.1002/jcb.21803. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 182.Kovalszky I, Dudas J, Olah-Nagy J, et al. Inhibition of DNA topoisomerase I activity by heparan sulfate and modulation by basic fibroblast growth factor. Mol Cell Biochem. 1998;183:11–23. doi: 10.1023/a:1006898920637. [DOI] [PubMed] [Google Scholar]
- 183.Ohkawa T, Naomoto Y, Takaoka M, et al. Localization of heparanase in esophageal cancer cells: respective roles in prognosis and differentiation. Lab Invest. 2004;84:1289–1304. doi: 10.1038/labinvest.3700159. [DOI] [PubMed] [Google Scholar]
- 184.Schubert SY, Ilan N, Shushy M, Ben-Izhak O, Vlodavsky I, Goldshmidt O. Human heparanase nuclear localization and enzymatic activity. Lab Invest. 2004;84:535–544. doi: 10.1038/labinvest.3700084. [DOI] [PubMed] [Google Scholar]
- 185.Kobayashi M, Naomoto Y, Nobuhisa T, et al. Heparanase regulates esophageal keratinocyte differentiation through nuclear translocation and heparan sulfate cleavage. Differentiation. 2006;74:235–243. doi: 10.1111/j.1432-0436.2006.00072.x. [DOI] [PubMed] [Google Scholar]
- 186.Purushothaman A, Hurst DR, Pisano C, Mizumoto S, Sugahara K, Sanderson RD (2011) Heparanase-mediated loss of nuclear syndecan-1 enhances histone acetyltransferase (HAT) activity to promote expression of genes that drive an aggressive tumor phenotype. J Biol Chem PMID: 21757697 [DOI] [PMC free article] [PubMed]
- 187.Szymczak M, Kuzniar J, Klinger M. The role of heparanase in diseases of the glomeruli. Arch Immunol Ther Exp. 2010;58:45–56. doi: 10.1007/s00005-009-0061-6. [DOI] [PubMed] [Google Scholar]
- 188.Hoven MJ, Rops AL, Vlodavsky I, Levidiotis V, Berden JH, Vlag J. Heparanase in glomerular diseases. Kidney Int. 2007;72:543–548. doi: 10.1038/sj.ki.5002337. [DOI] [PubMed] [Google Scholar]
- 189.Levidiotis V, Freeman C, Tikellis C, Cooper ME, Power DA. Heparanase is involved in the pathogenesis of proteinuria as a result of glomerulonephritis. J Am Soc Nephrol. 2004;15:68–78. doi: 10.1097/01.asn.0000103229.25389.40. [DOI] [PubMed] [Google Scholar]
- 190.Levidiotis V, Kanellis J, Ierino FL, Power DA. Increased expression of heparanase in puromycin aminonucleoside nephrosis. Kidney Int. 2001;60:1287–1296. doi: 10.1046/j.1523-1755.2001.00934.x. [DOI] [PubMed] [Google Scholar]
- 191.Levidiotis V, Freeman C, Tikellis C, Cooper ME, Power DA. Heparanase inhibition reduces proteinuria in a model of accelerated anti-glomerular basement membrane antibody disease. Nephrology. 2005;10:167–173. doi: 10.1111/j.1440-1797.2005.00388.x. [DOI] [PubMed] [Google Scholar]
- 192.Ritchie JP, Ramani VC, Ren Y, et al. SST0001, a chemically modified heparin, inhibits myeloma growth and angiogenesis via disruption of the heparanase/syndecan-1 axis. Clin Cancer Res. 2011;17:1382–1393. doi: 10.1158/1078-0432.CCR-10-2476. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 193.Higgins WJ, Fox DM, Kowalski PS, Nielsen JE, Worrall DM. Heparin enhances serpin inhibition of the cysteine protease cathepsin L. J Biol Chem. 2010;285:3722–3729. doi: 10.1074/jbc.M109.037358. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 194.Arvatz G, Shafat I, Levy-Adam F, Ilan N, Vlodavsky I. The heparanase system and tumor metastasis: is heparanase the seed and soil? Cancer Metastasis Rev. 2011;30:253–268. doi: 10.1007/s10555-011-9288-x. [DOI] [PubMed] [Google Scholar]
- 195.Dredge K, Hammond E, Handley P, et al. PG545, a dual heparanase and angiogenesis inhibitor, induces potent anti-tumour and anti-metastatic efficacy in preclinical models. Br J Cancer. 2011;104:635–642. doi: 10.1038/bjc.2011.11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 196.Zhang L, Sullivan PS, Goodman JC, Gunaratne PH, Marchetti D. MicroRNA-1258 suppresses breast cancer brain metastasis by targeting heparanase. Cancer Res. 2011;71:645–654. doi: 10.1158/0008-5472.CAN-10-1910. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 197.Gohji K, Hirano H, Okamoto M, Kitazawa S, Toyoshima M, Dong J, Katsuoka Y, Nakajima M. Int J Cancer. 2001;95:295–301. doi: 10.1002/1097-0215(20010920)95:5<295::aid-ijc1051>3.0.co;2-a. [DOI] [PubMed] [Google Scholar]
- 198.Gohji K, Okamoto M, Kitazawa S, Toyoshima M, Dong J, Katsuoka Y, Nakajima M. J Urol. 2001;166:1286–1290. [PubMed] [Google Scholar]
- 199.Shinyo Y, Kodama J, Hongo A, Yoshinouchi M, Hiramatsu Y. Ann Oncol. 2003;14:1505–1510. doi: 10.1093/annonc/mdg407. [DOI] [PubMed] [Google Scholar]
- 200.Nobuhisa T, Naomoto Y, Ohkawa T, Takaoka M, Ono R, Murata T, Gunduz M, Shirakawa Y, Yamatsuji T, Haisa M, Matsuoka J, Tsujigiwa H, Nagatsuka H, Nakajima M, Tanaka N. J Cancer Res Clin Oncol. 2005;131:229–237. doi: 10.1007/s00432-004-0644-x. [DOI] [PubMed] [Google Scholar]
- 201.Sato T, Yamaguchi A, Goi T, Hirono Y, Takeuchi K, Katayama K, Matsukawa S. J Surg Oncol. 2004;87:174–181. doi: 10.1002/jso.20097. [DOI] [PubMed] [Google Scholar]
- 202.Watanabe M, Aoki Y, Kase H, Tanaka K. Gynecol Obstet Invest. 2003;56:77–82. doi: 10.1159/000072821. [DOI] [PubMed] [Google Scholar]
- 203.Endo K, Maejara U, Baba H, Tokunaga E, Koga T, Ikeda Y, Toh Y, Kohnoe S, Okamura T, Nakajima M, Sugimachi K. Anticancer Res. 2001;21:3365–3369. [PubMed] [Google Scholar]
- 204.Takaoka M, Naomoto Y, Ohkawa T, Uetsuka H, Shirakawa Y, Uno F, Fujiwara T, Gunduz M, Nagatsuka H, Nakajima M, Tanaka N, Haisa M. Lab Invest. 2003;83:613–622. doi: 10.1097/01.lab.0000067482.84946.bd. [DOI] [PubMed] [Google Scholar]
- 205.Takahashi H, Ebihara S, Okazaki T, Suzuki S, Asada M, Kubo H, Sasaki H (2004) Lung Canc (Amsterdam, Netherlands) 45: 207–214 [DOI] [PubMed]
- 206.Cohen E, Doweck I, Naroditsky I, Ben-Izhak O, Kremer R, Best LA, Vlodavsky I, Ilan N. Cancer. 2008;113:1004–1011. doi: 10.1002/cncr.23680. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 207.Kelly T, Miao H-Q, Yang Y, Navarro E, Kussie P, Huang Y, MacLeod V, Casciano J, Joseph L, Zhan F, Zangari M, Barlogie B, Shaughnessy J, Sanderson RD. Cancer Res. 2003;63:8749–8756. [PubMed] [Google Scholar]
- 208.Bar-Sela G, Kaplan-Cohen V, Ilan N, Vlodavsky I, Ben-Izhak O. Histopathology. 2006;49:188–193. doi: 10.1111/j.1365-2559.2006.02469.x. [DOI] [PubMed] [Google Scholar]
- 209.Zheng LD, Tong QS, Tang ST, Du ZY, Liu Y, Jiang GS, Cai JB. World J Pediatr. 2009;5:206–210. doi: 10.1007/s12519-009-0039-9. [DOI] [PubMed] [Google Scholar]
- 210.Kim AW, Xu X, Hollinger EF, Gattuso P, Godellas CV, Prinz RA. J Gastrointest Surg. 2002;6:167–172. doi: 10.1016/s1091-255x(01)00087-7. [DOI] [PubMed] [Google Scholar]
- 211.Mikami S, Oya M, Shimoda M, Mizuno R, Ishida M, Kosaka T, Mukai M, Nakajima M, Okada Y. Clin Cancer Res. 2008;14:6055–6061. doi: 10.1158/1078-0432.CCR-08-0750. [DOI] [PubMed] [Google Scholar]
- 212.Ben-Izhak O, Kaplan-Cohen V, Ilan N, Gan S, Vlodavsky I, Nagler R. Neoplasia. 2006;8:879–884. doi: 10.1593/neo.06382. [DOI] [PMC free article] [PubMed] [Google Scholar]