Molecular pathogenesis of the obligate intracellular bacterium Coxiella burnetii (original) (raw)
. Author manuscript; available in PMC: 2014 Aug 15.
Published in final edited form as: Nat Rev Microbiol. 2013 Jun 24;11(8):561–573. doi: 10.1038/nrmicro3049
Abstract
The agent of Q fever, Coxiella burnetii, is an obligate intracellular bacterium that causes acute and chronic infections. The study of C. burnetii pathogenesis has benefited from two recent fundamental advances: improved genetic tools and the ability to grow the bacterium in extracellular media. In this Review, we describe how these recent advances have improved our understanding of C. burnetii invasion and host cell modulation, including the formation of replication-permissive _Coxiella_-containing vacuoles. Furthermore, we describe the Dot/Icm (defect in organelle trafficking/intracellular multiplication) system, which is used by C. burnetii to secrete a range of effector proteins into the host cell, and we discuss the role of these effectors in remodelling the host cell.
Coxiella burnetii is the Gram-negative obligate intracellular pathogen that causes Q fever, a condition which presents as either acute or chronic disease (BOX 1). Phylogenetic analyses have shown that isolates causing acute and chronic disease fall into distinct groups, supporting the prediction that there are genetically distinct C. burnetii pathotypes (BOX 2). As an intra-cellular pathogen, this organism has evolved a range of mechanisms to invade and survive within host cells. C. burnetii has a tropism for professional phagocytes and invades such cells using classic phagocytic mechanisms that rely on specific receptor–ligand interactions1. Inside the host cell, invading bacterial pathogens typically subvert phagosomal maturation using a variety of mechanisms2. However, C. burnetii does not follow this paradigm, but instead actively directs the maturation of a phagolysosome-like compartment known as the _Coxiella_-containing vacuole (CCV)3–5.
Box 1. Clinical presentation and epidemiology of Q fever.
Q fever is a zoonosis that is most often transmitted by aerosolized soil or animal products contaminated with Coxiella burnetii. Cattle, sheep and goats are the most common reservoirs for this pathogen, but it also infects many wild animal species and arthropods, and is therefore considered a ubiquitous zoonotic contaminant107. Although infection rarely results in mortality, it does cause significant morbidity. Furthermore, the bacterium is stable for long periods in the environment and has therefore been classified as a category B select agent by the US CDC because of the potential for use as a bioterrorism agent108.
The infectious dose for C. burnetii can be as low as a single organism in the lungs, and acute Q fever usually occurs 2–6 weeks after exposure, although the infection remains asymptomatic in >50% of cases. Symptomatic disease is characterized by high fever, severe headache (retro-orbital), myalgia, malaise, pneumonia and hepatitis, and usually lasts 1–2 weeks109. In symptomatic cases, the acute illness spontaneously resolves within 1–6 weeks and is effectively mitigated by antibiotics such as tetracycline and third-generation cephalosporins110,111. By contrast, some studies have estimated that in 1–2% of cases, acute Q fever develops into a chronic infection that manifests as endocarditis, hepatitis or as an incompletely defined post-Q fever fatigue syndrome112–114. Diagnosis of acute Q fever is difficult because obtaining a culture of the organism is challenging and requires a BL3 containment laboratory. Diagnosis relies on serology using immunofluorescence assays and ELISAs (enzyme-linked immunosorbent assays).
Most outbreaks involve a limited number of people, but the potential impact on a large population was recently highlighted by infections in the Netherlands115. Between 2007 and 2010, more than 4,000 cases of acute Q fever were reported in this country, with an annual summer seasonal peak. The outbreak was mainly restricted to the south of the country, coincident with intensive dairy goat farming in this region, and the source of the infections seemed to be linked to several goat herds in close proximity to population centres. The outbreaks provided a clear demonstration of the threat to public health if adequate diagnostic, therapeutic and epidemiological tools are not developed and available. Following the introduction of veterinary control measures in 2009 (including the vaccination of goat and sheep dairy herds, culling of pregnant animals on infected farms116 and improved hygiene procedures), the number of reported Q fever cases in the Netherlands has returned to the low historical rates (fewer than ten cases per year). In addition, a dramatically improved diagnostics and epidemiology network is now established, and public health officials are anticipating a slow influx of chronic disease cases over the next decade, as a result of the ~15% infection rate estimated for the regional populations in which the outbreaks occurred117.
Box 2. Pathotype-specific virulence.
Pathotype-specific virulence of Coxiella burnetii is a long-standing hypothesis that could account for the difference between the propensity of different isolates to cause either acute or chronic disease118–122. Restriction fragment-length polymorphism (RFLP) and pulse-field gel electrophoresis (PFGE) analysis of DNA isolated from 32 C. burnetii strains led to the differentiation of isolates into six genomic groups (group I to group VI; see the table (group III is not shown)) 119 that showed a distinct pattern of association with acute and chronic disease. Interestingly, isolates from group I, II and III are associated with acute infections, whereas groups IV and V consist of isolates associated with chronic infections (group VI contains the C. burnetii str. Dugway isolates, which have low virulence in guinea pigs). Other genotyping approaches have taken advantage of the fully sequenced C. burnetii genome using either multispacer sequence typing (MST) or multiple-locus variable number of tandem repeats analysis (MLVA). MST analysis suggested that the plasmids QpH1 and QpDV are associated with acute disease isolates, and QpRS is associated with chronic disease isolates, indicating a correlation between genotype and disease manifestation118. Similarly to MST analysis, MLVA analysis demonstrated that there are genetic differences between isolate groups from acute and chronic disease, suggesting that isolates cluster into distinct groups according to disease severity118,121. Consistent with this, there are no polymorphic ORFs between the isolates in restriction group I and the reference isolate C. burnetii str. RSA 493 (also in group I), but 87 polymorphic ORFs were identified between group V isolates and C. burnetii str. RSA 493, and at least half of these were annotated as hypothetical proteins123. These results are intriguing because isolates from group V are associated with chronic infections, suggesting that novel virulence factors contribute to distinct acute and chronic pathotypes119.
In a mouse infection model, isolates from group I induce persistent high-level cytokine secretion throughout the duration of infection, whereas group IV and group V isolates induce only moderate secretion at the peak of clinical disease 124. In the guinea pig model, isolates from group IV fail to induce fever, whereas group I isolates induce high fever and cause death at the highest challenge dose. In the severe combined immunodeficiency (SCID) mouse, which cannot mount an acquired immune response, group I isolates cause severe cachexia and death, whereas group IV and group V isolates induced only transient weight loss. Although this evidence clearly suggests that genetic pathotype differences exist, host factors are also likely to be involved in the development of chronic Q fever, as patients who develop a chronic infection often exhibit elevated interleukin-10 (IL-10) production125, and mice that constitutively express IL-10 have been proposed as a suitable chronic model for Q fever126. It is likely that both pathotype-specific differences and host factors are involved in the development of chronic Q fever.
| Isolate (strain name) | Genome size (bp) | Plasmid present | Source | Restriction group | MST group | Acute disease potential124 |
|---|---|---|---|---|---|---|
| C. burnetii str. Nine Mile IRSA 493 | 1,995,281 | QpH1 | Tick, USA, 1935 | I | 16 | High virulence |
| C. burnetii str. HenzerlingRSA 331 | 2,016,427 | QpH1 | Human blood, Italy, 1945 | II | 18 | Not determined |
| _C. burnetii_str. CbuK_Q154 | 2,102,380 | QpRS | Human heart valve USA, 1976 | IV | 8 | Low virulence |
| _C. burnetii_str. CbuG_Q212 | 2,008,870 | None | Human heart valve, Canada, 1981 | V | 21 | Intermediate virulence |
| C. burnetii str. Dugway5J108-111 | 2,158,758 | QpDG | Rodent, USA, 1958 | VI | 20 | Low virulence |
C. burnetii is readily transmitted between hosts and environmental reservoirs, a characteristic that is partially attributed to the ability of the bacterium to survive in the environment for long periods of time. The environmental stability of the organism seems to be linked to its ability to transition between distinct developmental stages. Small cell variants (SCVs) are metabolically inactive and resistant to numerous harsh environmental conditions. The switch from SCV to large cell variant (LCV) occurs after the invasion of host cells and during acidification of the phagosome, which triggers C. burnetii to become metabolically active6,7. Until recently, the only characterized virulence factor of C. burnetii was lipopolysaccharide (LPS), which was identified through the observation that serial passage in a non-immunologically competent host results in the transition from a smooth to a rough phenotype and the loss of virulence8. Virulent smooth cell variants contain LPS with a complete O antigen and are referred to as phase I organisms, whereas avirulent rough cell variants lack the terminal O antigen sugars and are referred to as phase II organisms. The identification and characterization of other virulence factors has been facilitated by two recent advances: the development of axenic culture media and improvements in the tools available for genetic manipulation (BOX 3).
Box 3. Recent progress in the development of extracellular growth media and genetic tools.
The first successful introduction of exogenous plasmid DNA into Coxiella burnetii was reported more than 10 years ago and involved the transformation of eukaryotic host cell-propagated C. burnetii with a shuttle vector containing a 5.8 kb autonomous replication sequence (ARS)127,128. The generation of new genetic tools has benefited from the development of axenic media to propagate C. burnetii. This advance was made after a breakthrough metabolic study demonstrated that the organism has enhanced metabolic potential in vitro in the presence of acid activation buffer (pH 4.5)129. The second major breakthrough came after the discovery that terminal oxidases are associated with micro-aerophilic respiration130. This led investigators to test the metabolic potential of C. burnetii at low oxygen levels, which revealed that 2.5% oxygen is optimal for growth and metabolism130. In turn, these two observations led to the development of acidified citrate cysteine medium (ACCM), which allows for an ~3 log increase in the number of genome copies after 6 days of culture130.
Shortly after the development of ACCM, genetic transformation and the isolation of single colonies on solid medium were achieved using an improved medium (ACCM-2)75. The ability to isolate single colonies allowed the Himar1_-based transposon system to be used for the generation of mutant libraries131. The original C. burnetii Himar1 mutants were isolated after electroporation and infection of Vero cells, a technical and laborious process131. A second important genetic tool was a stably maintained shuttle vector plasmid for C. burnetii containing the plasmid RSF1010 origin of replication (adapted from use in Legionella pneumophila)127. As an alternative approach to transformation with plasmids, a site-specific Tn_7 transposition system was recently developed132. Finally, an easy-to-use, site-directed mutagenesis strategy has now been developed, which will accelerate the testing of virulence gene candidates in C. burnetii mutants64. This complement of genetic tools and extracellular growth methods will now allow for the fulfilment of Koch’s postulates to verify the functions of a number of genes, including the candidate virulence factor genes.
In this Review, we describe our current understanding of the molecular pathogenesis of C. burnetii, including a discussion of a newly identified virulence factor, the type IV Dot/Icm (defect in organelle trafficking/intra-cellular multiplication) secretion system. We also describe how advances in extracellular cultivation techniques, genetic modification of clonal populations and the use of animal models of acute disease are helping to further our understanding of this intracellular pathogen.
The Coxiella burnetii infection cycle
Adhesion and invasion
After aerosol transmission, C. burnetii targets alveolar macrophages and passively enters these cells by actin-dependent phagocytosis9,10. The identification of a type IV secretion system (T4SS) in C. burnetii suggested that this bacterium uses an active trigger mechanism (mediated by the secretion of T4SS effector proteins) to induce uptake after initial binding to the host cell11. However, recent studies using T4SS mutants have revealed that these bacteria have no defect in their ability to invade either phagocytic cells (monocytes and macrophages) or non-phagocytic cells, supporting the idea that entry into phagocytic cells proceeds via passive actin-dependent phagocytosis and that entry into non-professional phagocytes proceeds via an active zipper mechanism12,13. Non-phagocytic cells do not usually engulf large particles; however, intracellular pathogens can invade these cells after direct contact through the zippering of bacterial ligands to host cell receptors.
Virulent C. burnetii binds to phagocytic cells using as the main receptor and enters the cell αVβ3 integrin via RAC1-dependent phagocytosis, which requires membrane ruffling14,15 (FIG. 1). Interestingly, αVβ3 integrin is typically involved in the removal of apoptotic cells via phagocytosis, and this is generally associated with an inhibition of inflammation16. Thus, the ability to use αVβ3 integrin for invasion might be exploited by C. burnetii as a mechanism to avoid the induction of an inflammatory response. Consistent with this, C. burnetii is characterized as a stealth pathogen that enters cells without alerting the immune system. Although the identity of the bacterial ligand for α Vβ3 integrin has not been determined, a class of likely candidates would be membrane-associated proteins that contain the integrin-binding domain arginine-glycine-aspartic acid (RGD)11. αVβ3 integrin is poorly expressed on resting monocytes, so it is likely that C. burnetii uses other receptors to invade these cells17.
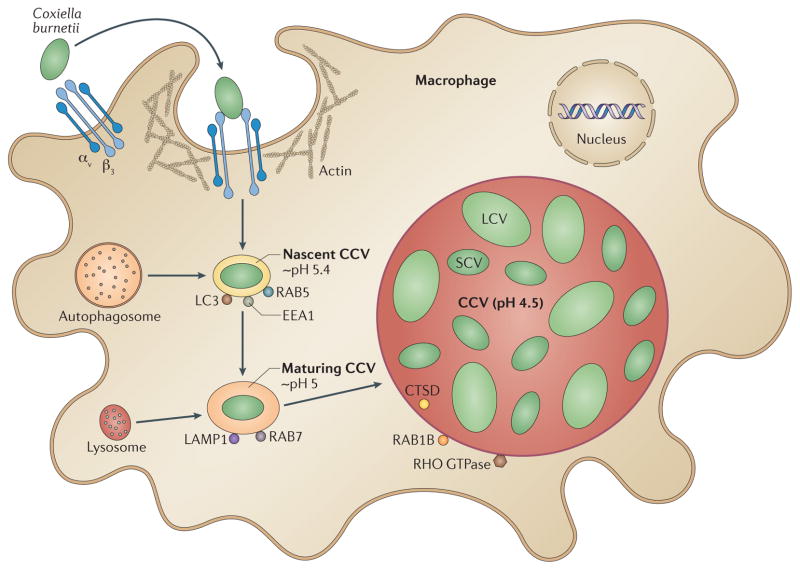
Figure 1. The intracellular trafficking pathway of Coxiella burnetii.
Coxiella burnetii binds to macrophages through αVβ3 integrin, which triggers phagocytosis of the bacterium through an actin-dependent mechanism. The nascent _Coxiella_-containing vacuole (CCV) acquires RAB5 and EEA1 as early as 5 minutes after internalization and acidifies to approximately pH 5.4, which is characteristic of normal phagosomal development. By contrast to phagosomes, the CCV also acquires microtubule-associated protein light-chain 3 (LC3; an autophagosomal marker), a process that is dependent on bacterial protein synthesis. The nascent CCV develops through fusion and fission events with early endosomes and then late endosomes, leading to the disappearance of RAB5 and EEA1 and the acquisition of RAB7 and lysosome-associated membrane glycoprotein 1 (LAMP1) 40 to 60 minutes after internalization, in concurrence with a further acidification to pH 5, which is characteristic of normal phagosomal development. Lysosomal enzymes, including cathepsin D (CTSD), start accumulating in the CCV by 2 hours after internalization, at which point the vacuole is at ~pH 4.5; this is delayed significantly from normal phagolysosomal acquisition of CTSD. This pause in CCV development might allow conversion of the bacteria from small cell variants (SCVs) to the metabolically active large cell variants (LCVs). Between 8 hours and 2 days after internalization, the CCV expands to occupy an increasingly dominant portion of the cytoplasmic space of the host cell. This process is dependent on bacterial protein synthesis and involves the recruitment of both RHO GTPase and RAB1B to the CCV membrane. RHO GTPase is likely to be involved in maintenance of the large vacuole, whereas the recruitment of RAB1B from the ER might facilitate the acquisition of additional membranes to create this spacious CCV.
Although the identity of the receptor on non-professional phagocytes is currently unknown, the mechanism of C. burnetii entry into these cells also appears to be dependent on actin rearrangement9. As the β3-subunit is poorly expressed on bronchial epithelium, it seems unlikely that αVβ3 integrin is the primary receptor for uptake in non-professional phagocytes, assuming that invasion of non-professional phagocytic cells is an important component of respiratory infection18. Histopathological analysis was carried out on the lungs of both animal models of pneumonia and human patients with atypical pneumonia caused by C. burnetii, and this analysis identified monocytes and macrophages as the primary infection sites, but epithelial and endothelial cell infection was also evident19. In a variety of tissue culture models, including mouse fibroblasts (L929), African green monkey kidney cells (Vero), human monocytes (THP-1) and mouse macrophages (J774), C. burnetii cells of different phases exhibit distinct uptake kinetics, whereby avirulent phase II bacteria are more readily internalized than fully virulent phase I bacteria9,10. Although this is an interesting phenomenon, the mechanisms by which these variants enter host cells is not discussed in depth here, as phase II C. burnetii does not cause infections in immunocompetent animal models or humans20. However, both phase I and phase II C. burnetii replicate with similar kinetics and in indistinguishable, proteolytically active CCVs in tissue culture cells21, which is important to note because many studies that examine intracellular trafficking (see below) have used the avirulent phase II variant (as it requires only BL2 containment)22.
Both phase I and phase II variants engage the αVβ3 integrin receptor in monocytes, but dramatic reorganization of the filamentous (F)-actin cytoskeleton is observed with phase I bacteria only14. Cytoskeletal changes during invasion are a common effect of several other invasive pathogens that promote phagocytosis through the secretion of effectors which activate host GTPases23,24. Although the T4SS of C. burnetii is not required for uptake by the host cell, it remains to be determined whether T4SS effectors contribute to the reorganization of the actin cytoskeleton during infection12,13. Interestingly, actin-cytoskeletal ruffling induced via SRC tyrosine kinases is observed following the binding of phase I C. burnetii25. This membrane ruffling requires contact between C. burnetii and host cells and can be induced using purified LPS from phase I bacteria25,26. The ability to induce these ruffles is also dependent on the expression of Toll-like receptor 4 (TLR4) on the host cell surface27. These observations suggest that LPS induces membrane ruffling and the T4SS effectors temporally control downstream signalling cascades after internalization, although this has not yet been demonstrated experimentally.
From phagosome to CCV
Phagocytosis results in the formation of the phagosome, which matures into a phagolysosome following a series of highly ordered and regulated fusion and fission events1. Shortly after internalization at the cell surface, the nascent phagosome develops into an early phagosome and acquires the small GTPase RAB5. This GTPase stimulates fusion with early endosomes, resulting in acidification of the lumen to approximately pH 5.4 (through an incompletely defined process) and acquisition of the early-endosomal marker protein EEA1 (REF. 28). One of the most striking features of the maturing phagosome is that its size remains constant, indicating that membrane components are constantly removed after fusion events1. The late phagosome lacks RAB5 but acquires the GTPase RAB7, lysosome-associated membrane glycoprotein 1 (LAMP1), LAMP2 and the vacuolar ATPase, which pumps protons into the maturing phagosome to further decrease the lumenal pH to about pH 5 (REF. 28). Finally the phagosome fuses with lysosomal compartments to acquire cathepsins and hydrolases, and the vacuolar ATPase further reduces the pH to around pH 4.5 (REFS 1,28).
The maturation of the CCV essentially follows the canonical endosomal pathway described above, but there are a number of important differences that rely on specific bacterial proteins. Thus, these differences are considered pathogenic mechanisms, although how they benefit C. burnetii remains to be determined. After internalization of the bacterium, the nascent CCV is similar in size to a nascent phagosome and sequentially recruits RAB5 and RAB7, indicating that it traffics through the typical endosomal cascade described above22 (FIG. 1). However, maturation of the CCV involves expansion of this compartment to a size that almost completely fills the host cell cytoplasm, an aspect that differs vastly from the unchanging size of the maturing phagolysosome1,29 (FIG. 1). Simultaneous functional inhibition of both RAB5 and RAB7 through dominant-negative mutations impairs expansion of the CCV, whereas inhibition of RAB5 alone (but not RAB7 alone) blocks C. burnetii entry into host cells22. Furthermore, formation of the large CCV is dependent on active C. burnetii protein synthesis, probably because of a requirement for the synthesis and subsequent secretion of effectors across the CCV membrane into the cytoplasm5,12. However, in the absence of protein synthesis, small CCVs still acquire LAMP proteins and acidify, suggesting that the acquisition of LAMPs and acidification are passive processes, similar to the normal maturation process of phagosomes30.
Most intracellular pathogens subvert the endosomal cascade and arrest maturation of the phagosome at an early stage to avoid fusion with lysosomes2. However, in C. burnetii, lysosomal enzymes, including cathepsin D (CTSD) and lysosomal acid phosphatase (ACP2), accumulate in the CCV, although this is delayed until 2 hours after infection (compared with only 15 minutes after infection for phagosomes harbouring inert particles)4. It has been suggested that the delay is caused by interactions of the CCV with the autophagy pathway22. The T4SS of C. burnetii is unlikely to be involved, as translocation of effectors does not occur until at least 8 hours after infection, suggesting that the bacterial proteins which interact with the autophagy pathway remain to be identified13. Autophagy is implicated in host defence through innate and adaptive immunity to intracellular pathogens31, and as early as 5 minutes after internalization, the CCV is decorated with the autophagy marker microtubule-associated protein light-chain 3 (LC3)3,22 (FIG. 1). However, the role of autophagy in C. burnetii pathogenesis is unclear; a recent genome-wide RNAi screen failed to show that autophagy is essential for C. burnetii growth. On the other hand, if autophagy is induced in the host cell before infection with C. burnetii, the intracellular bacterial load and the size of the CCV increase32,33. One potential benefit of the autophagy interaction is that autophagy vesicles are often loaded with nutrients and membranes from their degraded cargo, and these components might serve as fuel for the SCV-to-LCV conversion22,32. The conversion to LCVs coincides with the arrest of CCV endosomal maturation; the percentage of LCVs (as a proportion of the total number of SCVs and LCVs) 1 hour after infection approaches 80%, and by 16 hours the CCV contains only LCVs4,6 (FIG. 1).
Between 8 hours and 2 days after infection, the CCV enlarges drastically and can occupy nearly the entire volume of the host cell34. The large CCV forms as a result of homotypic fusion of multiple smaller CCVs, and it can continue to expand through heterotypic fusion with autophagic, endocytic and lysosome vesicles5,30. Maintenance of the large CCV not only requires protein production by the bacterium30, but also depends on the actin cytoskeleton, as treatment of infected cells with actin-depolymerizing agents results in the formation of small CCVs only35. In addition, C. burnetii infection activates the host cell kinase families protein kinase C, protein kinase A and myosin light chain kinase, which are required for the establishment and maintenance of the CCV36. Furthermore, during this stage of the infection, the CCV interacts with the early secretory pathway (as suggested by the accumulation of RAB1B on the CCV membrane), an interaction which is required for formation of the large CCV37 (FIG. 1). Interaction with the ER through the early secretory pathway might provide a source of lipids for the formation of the large CCV. These interactions are probably orchestrated by C. burnetii effectors, as bacterial protein production is required for the formation of the large CCV; however, the identity and specific functions of these putative effectors remain to be determined37. Interestingly, the CCV membrane contains a comparable amount of cholesterol to that in the plasma membrane, which is twice that of a normal lysosomal compartment38. Furthermore, host genes involved in cholesterol synthesis are upregulated during C. burnetii infection, and inhibition of cholesterol metabolism negatively affects CCV formation, suggesting that C. burnetii directly influences cholesterol metabolism, potentially through interactions with the ER38–40. Although these events are reasonably well characterized, the C. burnetii virulence factors that direct formation of the CCV remain to be identified.
Mature CCV
Approximately 6 days after infection, the CCV is heavily loaded with LCV bacteria, which start differentiating back into SCVs6 (FIG. 1). The mature CCV maintains essentially the same features as the early CCV: the pH remains low (it has a pH of about 4.5–5, the same pH as the phagolysosome in uninfected cells41) and the vacuole contains all of the same markers. In addition, the mature CCV retains fusogenic capacity, which relies on C. burnetii protein synthesis5. Surprisingly, host cell viability is not affected by the drastic expansion of the CCV, which occupies much of the cytoplasm and with a volume much larger than the space required by C. burnetii42. In addition, the generation time and genome stability of the host cell is unchanged43. C. burnetii prolongs host cell viability in two ways: it actively inhibits apoptotic signalling pathways, and it induces pro-survival factors. For example, C. burnetii prevents the induction of apoptosis following exogenous activation with the pro-apototic drug staurosporine, and this inhibitory effect is dependent on protein production by the bacterium42. The anti-apoptotic activity of C. burnetii could be the result of the interplay between beclin 1 (BECN1) and BCL2. BECN1 is an autophagy initiation protein that interacts with the anti-apoptotic protein BCL2, and both are present on the CCV. Their interaction prevents the release of cytochrome c from the mitochondria and thereby leads to apoptosis inhibition, a phenomenon associated with C. burnetii infection and a known function of BCL2 (REFS 44,45). A second anti-apoptotic activity initiated after C. burnetii infection is the sustained activation of the pro-survival signalling proteins ERK1 (also known as MAPK3), ERK2 (also known as MAPK1) and the AKT family46.
Promoting the survival of infected cells seems to be essential for the establishment and maintenance of chronic C. burnetii infections. There is evidence to suggest that during host cell cytokinesis the large CCV is segregated to only one of the daughter cells, leaving the other cell uninfected47. This would provide the bacterium with host cells to infect, thereby promoting a chronic infection. The ability to prevent apoptosis and stimulate pro-survival pathways would also be beneficial for persistent infection, as this activity maintains the host cell to allow continued replication of the bacterium. During acute infection, the infectious dose can be as low as 1–10 C. burnetii cells, which suggests that there are other mechanisms in place that promote the spread of replicating bacteria to infect other susceptible cells48. A recent report demonstrated that C. burnetii is also able to induce apoptosis through the release of cytochrome c in a mechanism that is dependent on bacterial protein synthesis49. Therefore, the prevention of apoptosis seems to be exploited by C. burnetii to cause persistent infections, whereas induction of apoptosis seems to enable spread of the infection to nearby susceptible cells. This strategy is also used by other intracellular pathogens; for example, Mycobacterium tuberculosis inhibits apoptosis early during infection, but induces cell death at a later stage of infection50.
Dot/Icm secretion and host subversion
Sequencing of the C. burnetii genome identified three secretion systems: a type I secretion system (T1SS)51, a T2SS-related pilus biogenesis machinery52 and a conjugation-related T4SS53. Currently, little is known about the role of the T1SS and T2SS in the pathogenesis of C. burnetii. By contrast, the availability of Legionella pneumophila as a surrogate host for the expression of putative C. burnetii T4SS effectors has provided insight into the role of this secretion system, and this is the focus of the discussion below.
The T4BSS (the Dot/Icm secretion system)
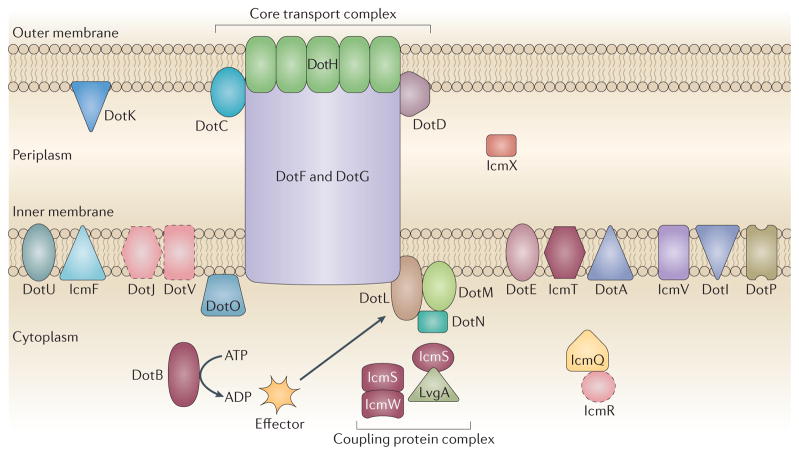
T4SSs translocate effector substrates from the bacterial cytosol directly into the cytosol of eukaryotic host cells and are intimately involved in the pathogenesis of many bacteria. These multicomponent protein machines can be subdivided into several substructures, including a pilus, a core transport complex and a type IV coupling protein complex54 (FIG. 2). Furthermore, T4SSs are separated into two subgroups, T4ASS and T4BSS, on the basis of homology to the Agrobacterium tumefaciens and L. pneumophila systems, respectively55. The T4BSS, which is also known as the Dot/Icm system, resembles the conjugation machinery encoded on IncI plasmids. Studies with L. pneumophila have identified several core complexes that define the topology of the Dot/Icm apparatus, such as the subcomplex consisting of DotC, DotD, DotF (also known as IcmG), DotG (also known as IcmE) and DotH (also known as IcmK), which bridges the inner and outer bacterial membranes and is functionally the core transport complex56 (FIG. 2). A second subcomplex comprises the coupling protein DotL (also known as IcmO) (which recruits IcmSW-dependent substrates to the secretion apparatus), DotM (also known as IcmP), DotN (also known as IcmJ), IcmS and IcmW57 (FIG. 2). The C. burnetii str. Nine Mile I isolate encodes homologues for 24 of the 27 Dot/Icm proteins found in L. pneumophila, and this high degree of similarity between the two systems suggests that they are structurally similar11. The C. burnetii genome lacks dotV (also known as icmC), icmR and dotJ (also known as icmM), but has a gene duplication of dotI (also known as icmL; resulting in the genes icmL.1 and icmL.2); the similarity of DotJ and DotI suggests that this DotI duplication can substitute for the absence of DotJ, a functional homologue of IcmR11,58 (FIG. 2). The conservation between these two T4BSSs became apparent when it was shown that dotB, icmS, icmW and icmT from C. burnetii can complement the intracellular growth defect associated with an L. pneumophila strain in which these genes are mutated59. This was the first indication that C. burnetii encodes a functional Dot/Icm system and led to the use of L. pneumophila as a surrogate host to identify and characterize C. burnetii effector proteins60,61 (TABLE 1). Using this system, transcriptional analysis demonstrated that several C. burnetii dot/icm genes are actively transcribed at early time points after infection62, and similarly to in L. pneumophila, the Dot/Icm system localizes to the poles of C. burnetii cells during infection63. Using transposon and site-specific mutants, several studies have now confirmed the dependence of C. burnetii on the Dot/Icm system for intracellular survival64,12,13. Current research is now focused on identifying the roles of the T4SS effectors to provide insight into the pathogenic mechanisms of this intracellular bacterium.
Figure 2. The type IVB secretion systems of Coxiella burnetii and Legionella pneumophila.
Coxiella burnetii encodes 24 of the 27 Legionella pneumophila type IVB secretion system (T4BSS) components. The C. burnetii system lacks three homologues, DotJ (defect in organelle trafficking J; also known as IcmM), DotV and IcmR (intracellular multiplication R), which are depicted with a dashed outline. Studies using L. pneumophila identified a subcomplex termed the core transport complex, which links the inner and outer membranes and is composed of DotC, DotD, DotF, DotG and DotH. As this complex is also conserved in the T4ASS, it is hypothesized that C. burnetii assembles a similar complex. A second subcomplex consists of the coupling protein DotL (providing a link between the substrates and the transport complex), DotM, DotN, IcmS and IcmW. The ATPase activity of DotB is required for secretion, but the function of this activity remains to be determined. C. burnetii proteins that can substitute for the corresponding proteins in the L. pneumophila system are shown in maroon.
Table 1.
Coxiella burnetii Dot/Icm substrates with predicted effector functions
| C. burnetii str. Nine Mile I locus | Encoded protein | Genetic differences between homologues* | Encoded protein domains | Protein localization‡ | Protein function | Refs | ||
|---|---|---|---|---|---|---|---|---|
| C. burnetii str. CbuK_Q154 | C. burnetii str. CbuG_Q212 | C. burnetii str. Dugway | ||||||
| CBU0072 | AnkA | – | A | AS | Ankyrin | Cytoplasm | ND | 60 |
| CBU0144/0145 | AnkB | AS | – | – | Ankyrin | Cytoplasm | ND | 60 |
| CBU0447§ | AnkF | – | – | – | Ankyrin | Cytoplasm | ND | 60 |
| CBU0781 | AnkG | NT | – | – | Ankyrin | Microtubules | Anti-apoptotic | 60,78 |
| CBUA0006§ | CpeA | – | – | – | ND | Puncta | ND | 67 |
| CBUA0013§ | CpeB | – | – | – | ND | Autophagosome | ND | 67 |
| CBUA0014 | CpeC | A | A | A | F-box | Ubiquitin-rich structures | ND | 67 |
| CBUA0015 | CpeD | A | A | A | Kinase, coil | ER (partial) | ND | 67 |
| CBUA0016 | CpeE | A | A | A | ND | No distinct localization | ND | 67 |
| CBUA0023§ | CpeF | – | – | – | ND | No distinct localization | ND | 67 |
| CBU1532 | CaeB | AS, CT | AS | AS | ND | Mitochondria | Anti-apoptotic | 13,77 |
| CBU1524 | CaeA | – | – | AS | ND | Nucleus | Anti-apoptotic | 13,77 |
| CBU1314§ | Not yet named | – | – | – | Coiled-coil | Nucleus | ND | 61 |
| CBU1543§ | Not yet named | – | – | – | Coiled-coil | Vesicles | ND | 61 |
| CBU1556§ | Not yet named | – | – | – | Coiled-coil | Cytoskeleton-like filaments | ND | 61 |
| CBU0801§ | Not yet named | – | – | – | ND | Vesicles | ND | 61 |
| CBU1825§ | Not yet named | – | – | – | ND | Mitochondria | ND | 13 |
| CBU1379 | Not yet named | – | – | FS | Protein kinase | No distinct localization | ND | 61 |
| CBU1457§ | Not yet named | – | – | – | Tetratricopeptide | ND | ND | 61 |
| CBU2078§ | Not yet named | – | – | – | Fic | ND | ND | 61 |
| CBU0077§ | Not yet named | – | – | – | ND | Lysosome | ND | 13 |
| CBU0635§ | Not yet named | NT | – | – | ND | Golgi, vesicles | Inhibits the secretory pathway | 13 |
Identification and characterization of Dot/Icm substrates
Initially, all C. burnetii effector screens used L. pneumophila as a surrogate host65,66. However, the development of C. burnetii shuttle vectors (BOX 3) enabled the adaptation of fluorescence-based β-lactamase (TEM1) translocation assays for C. burnetii, which were used to confirm the results observed using L. pneumophila61,67. Recent genetic and bioinformatic screens have identified 60 C. burnetii Dot/Icm substrates (the best characterized of which are shown in TABLE 1), and ~60 further substrates have recently been added to this list13,60,61,65,67,68 (Robert A. Heinzen, personal communication). To date, the target and function of the majority of the ~120 Dot/Icm substrates remain undefined. A large number of the encoding genes (23 of the 60 substrate genes initially characterized) have a GC content that is significantly different from the average GC content of the C. burnetii genome (42.7%)61. The obligate intracellular nature of C. burnetii and the presence of eukaryotic-like domains in many substrates suggest that the genes encoding these substrates were acquired by interdomain horizontal gene transfer from a eukaryotic source51,69. Furthermore, similarly to L. pneumophila effectors, many of the C. burnetii T4SS substrates contain a carboxy-terminal motif or recognition sequence that is required for translocation61,66,67.
Bacterial pathogens use the T4SS as part of their virulence programme to control the activity of various effector proteins in order to ensure the proper progression of infection. For example, the L. pneumophila effector protein LubX acts as an E3 ubiquitin ligase that targets the effector SidH for degradation several hours after infection, representing a mechanism of temporal control70. C. burnetii effector functions are likely to be controlled by similar temporal mechanisms, including transcriptional control, translocation efficiency and protein stability in host cells. This hypothesis was supported by the recent observation that C. burnetii Dot/Icm effectors are not translocated until 8 hours after infection and also require acidification of the CCV for translocation13,71. Although this expression pattern is in agreement with the ability of dot/icm mutants to passively traffic through the endocytic pathway to the phagolysosome, it does not explain how C. burnetii is able to manipulate the autophagy pathway immediately after uptake into host cells, nor how the bacterium delays lysosomal fusion12,13. Thus, it is possible that T4SS-independent effectors are involved in the manipulation of the autophagy pathway and the delay in lysosomal fusion.
Another mechanism of control is executed at the transcriptional level by the two-component regulatory system PmrAB, which has been shown to directly regulate the Dot/Icm secretion system in both L. pneumophila and C. burnetii72. The PmrAB system regulates virulence genes in other pathogenic bacteria, such as Pseudomonas aeruginosa, in response to low Mg2+ levels, high Fe3+ levels and low pH73. Bioinformatic analysis revealed that genes encoding five of the structural Dot/Icm proteins (dotP (also known as icmD), icmQ, icmV, icmW and dotD) contain a PmrA-binding sequence located upstream of the −10 region of their promoters. Furthermore, 20 putative PmrA-regulated C. burnetii genes have been confirmed as Dot/Icm substrates, indicating that this system might represent a global virulence regulator in C. burnetii61,65,72. Bioinformatic analysis also suggests that other unknown mechanisms exist for regulating non-PmrA-regulated effectors, as many newly identified effectors do not contain PmrA-binding sequences but are still secreted. Although L. pneumophila and C. burnetii are phylogenetically related, only ten of the identified Dot/Icm substrates from C. burnetii display significant similarity to any of the >250 L. pneumophila T4SS substrates. This is not surprising considering that L. pneumophila and C. burnetii replicate in distinct vacuolar compartments during co-infection74, and suggests that L. pneumophila and C. burnetii Dot/Icm effectors have specific functions that target diverse vesicle trafficking pathways to create a distinct permissive replicative environment for each bacterium.
Plasticity and redundancy in Dot/Icm substrates from different pathotypes
The Dot/Icm effectors exhibit striking heterogeneity among the sequenced C. burnetii isolates belonging to different pathotypes13. Only 19 of the currently identified effectors, including three plasmid-encoded effectors, are fully conserved among the acute and chronic isolates (TABLE 1). The fact that these effectors have been maintained in in all C. burnetii isolates suggests that they have essential roles for intracellular survival. The plasticity of the remaining effectors might be mediated, in part, by extensive recombination between abundant insertion sequences (ISs) found within the C. burnetii genome51. In isolates causing chronic and avirulent disease, the majority of homologues for the C. burnetii Nine Mile I effectors are either truncated or found as pseudogenes. This suggests that the truncated homologues are nonfunctional, a hypothesis supported by the observation that ectopic expression of CBU1532 (encoded by C. burnetii str. Nine Mile I) leads to rounding of the host cell, whereas the alternative-start homologue CBUD0454 (encoded by C. burnetii str. Dugway) does not induce this phenotype13. Thus, we speculate that several group-specific Dot/Icm effectors participate in pathotype-specific virulence, a hypothesis that is now testable with site-specific mutagenesis methods64,75.
Compared with other secretion systems, the Dot/Icm system from L. pneumophila has an astounding number of substrates. Dot/Icm is capable of translocating at least 8.5% of the L. pneumophila proteome (which corresponds to approximately 250 out of 2,943 ORFs). Why would a pathogen need so many effectors? An intriguing recent study suggests that at least 30% of L. pneumophila Dot/Icm effectors are not involved in the establishment of mammalian infection, but have been acquired instead to enable adaptation to multiple hosts76. Although a survey of the C. burnetii proteome for secretion substrates is incomplete, it seems that a substantial proportion of the proteome — to date, 5.8% — does serve as T4SS substrates13,60,61,65,67,68 (Robert A. Heinzen, personal communication). C. burnetii is capable of infecting and colonizing a wide range of mammalian and arthropod hosts, and a diverse pool of T4SS substrates might facilitate this wide host range. In addition, L. pneumophila encodes several examples of multiple effectors that target the same host pathway to orchestrate intracellular bacterial survival. Evidence suggests that C. burnetii uses a similar strategy, as three effectors have been identified that inhibit apoptosis (AnkG, CaeA and CaeB)77,78 (TABLE 1).
Diverse cytosolic functions for Dot/Icm effectors
The majority of the identified Dot/Icm substrates are annotated as hypothetical proteins. However, most encode one or more eukaryotic-like domains implicated in protein–protein interactions, and these domains provide clues to protein functionality. These domains include ankyrin repeats, coiled-coil domains and tetratricopeptide repeats13,61,66 (TABLE 1). Other substrates have eukaryotic-like domains that are implicated in post-translational modifications, such as putative serine/threonine kinase domains, F-box domains and Fic domains13,61,67 (TABLE 1). Eukaryotic proteins carrying these motifs are involved in a variety of host processes, including apoptosis, ubiquitylation, lipid metabolism and membrane trafficking, suggesting that C. burnetii effectors are involved in modulating many of these cellular processes65,79–83. For example, the T4SS effectors AnkG, CaeA and CaeB all prevent intrinsic apoptosis77,78 (FIG. 3; TABLE 1). Modulation of apoptosis by AnkG is dependent on the ability of AnkG to bind the host mitochondrial matrix protein p32 (also known as C1QBP); the mechanism of action of p32 during apoptosis has not yet been established, but the protein is thought to regulate the opening of the permeability transition pore78. CaeB localizes to the mitochondria and inhibits mitochondrial membrane permeabilization, thus preventing the release of pro-apoptotic proteins77 (FIG. 3). These two effectors target the same cellular pathway, albeit through different mechanisms. Although CaeA localizes to the nucleus, the exact mechanism of apoptosis inhibition by this protein remains to be determined77. As discussed above, for C. burnetii, the benefits of inhibiting apoptosis include the maintenance of host cell viability, despite the mature CCV occupying almost the entire volume of the cell.
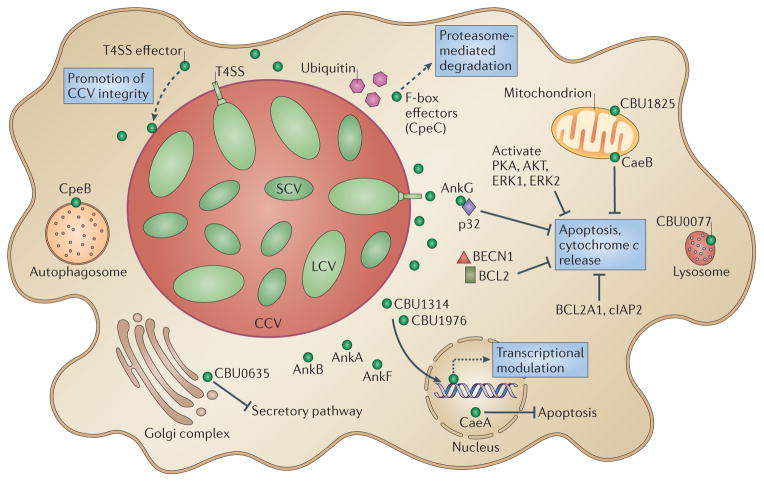
Figure 3. Roles of type IV secretion system effectors during Coxiella burnetii intracellular infection.
A new family of type IV secretion system (T4SS) effectors has been identified that localizes to the _Coxiella_-containing vacuole (CCV) and might have diverse functions, such as CCV stability and fusogenicity. Located next to the Golgi apparatus, CBU0635 interferes with the host secretory pathway on ectopic expression. At least three T4SS effectors localize to the nucleus (CaeA, CBU1314 and CBU1976) when ectopically expressed in host cells, and these proteins might be involved in the transcriptional modulation that occurs during Coxiella burnetii infection. Exogenously stimulated apoptosis is modulated by C. burnetii, and an ankyrin repeat-containing protein (AnkG) has anti-apoptotic activity through an interaction with p32, a host protein that is involved in pathogen-induced apoptosis. A second T4SS effector, CaeB, alters mitochondrial membrane permeability during exogenously stimulated apoptosis. C. burnetii infection also increases the synthesis of anti-apoptotic proteins BCL2-related protein A1 (BCL2A1; also known as BFL1) and cIAP2 (also known as BIRC3) and causes activation of the pro-survival kinases AKT, ERK1 and ERK2. Other effectors might also be involved in the modulation of apoptosis. Several T4SS effectors have F-box domains and are potentially involved in promoting proteasome-mediated degradation of proteins; these effectors include CpeC, which localizes to ubiquitin-rich compartments in host cells. The effector CpeB has been localized to the autophagosome, and AnkA, AnkB and AnkF localize to the cytoplasm. Dashed arrows represent predicted effector functions that have not yet been experimentally demonstrated. All T4SS effectors are shown as green circles. BECN1, beclin 1; LCV, large cell variant; PKA, protein kinase A; SCV, small cell variant.
Functional assays serve as a starting point for characterizing the many effector functions of C. burnetii, and ectopic expression of effector proteins in eukaryotic cells has demonstrated that many effectors target specific host organelles13,61,66,67. For example, at least three effectors (CaeA, CBU1314 and CBU1976) are targeted to the nucleus13,61, and this probably depends on the presence of nuclear localization motifs (TABLE 1). As C. burnetii infection has been shown to alter host gene transcription84, it is reasonable to suggest that these putative nuclear effectors are involved in this process (FIG. 3). In addition, several vesicle-associated effectors target the CCV and autophagosome compartments13,66. Recently, researchers have identified a family of CCV-binding proteins, several of which are essential for intracellular replication (Robert A. Heinzen, personal communication). These effectors might serve as CCV stabilizers or as docking points for signalling molecules, and/or they might promote the fusogenic activities of the CCV (FIG. 3). Because C. burnetii alters the fusogenic capacity of the CCV, identifying effectors that disturb the secretory pathway in eukaryotic cells is crucial. C. burnetii actively modulates vesicle trafficking pathways by recruiting membranes to the CCV, and there is evidence that the early secretory pathway contributes to CCV biogenesis37. Dot/Icm effectors that interfere with the host secretory pathway have been identified for L. pneumophila using a SEAP (secreted embryonic alkaline phosphatase) assay65,85–87. In C. burnetii, an effector (CBU0635) that disrupts mammalian secretory pathways has been identified using the same assay and localizes next to the Golgi apparatus13 (FIG. 3; TABLE 1). Further analysis will be required to determine the mechanism of action of CBU0635 and its contribution to C. burnetii virulence.
Ubiquitin-related F-box-containing proteins represent an interesting group of Dot/Icm effectors. Similarly to L. pneumophila, C. burnetii encodes several paralogues of F-box-containing proteins; three of them have been confirmed to be Dot/Icm effectors, and one (CpeC) is associated with ubiquitin61,67. On the basis of their function in eukaryotic cells, this group of proteins might promote proteasome-mediated protein degradation70,88. For example, AnkB (an F-box protein) of L. pneumophila targets non-essential host proteins (through ubiquitylation) for degradation by the 26S proteasome, providing a source of amino acids for bacterial growth88. A second possible mechanism involves the degradation of host proteins that impede bacterial replication. Whether C. burnetii F-box proteins have similar roles remains to be determined.
Our knowledge of the functions of the Dot/Icm effectors from C. burnetii is still in its infancy. However, the recent description of a site-directed mutagenesis system for C. burnetii64 should aid in the characterization of all ~120 identified effectors and help to establish how they contribute to pathogenesis.
Immune evasion strategies
C. burnetii has several immune evasion strategies that are associated with the structure of its LPS. Virulent phase I C. burnetii produces LPS with a complete O antigen and exhibits serum resistance (the bacterium only moderately activates complement and prevents surface deposition of complement factor C3b)89. Furthermore, the structure of this so-called smooth LPS masks bacterial recognition by the pattern recognition receptor TLR2 (REF. 90), which recognizes ligands within LPS. By contrast, avirulent phase II C. burnetii produces a rough LPS, which lacks the terminal O antigen sugars and is readily detected by TLR2; this detection induces the production of interleukin-12 (IL-12) and tumour necrosis factor (TNF), and activates macrophages to mediate bacterial clearance91. Conversely, phase I C. burnetii does not induce maturation of primary dendritic cells (DCs) and induces the production of relatively low levels of IL-12 and TNF90. This suggests that the LPS of phase I bacteria, containing the full-length O antigen, shields TLR2 ligands on the bacterial cell surface90. Other studies have shown that LPS from both phase I and phase II C. burnetii induces the production of TNF in macrophages, but this induction is likely to be due to contaminating TLR2 ligands, as purified LPS (which could contain contaminating TLR ligands) but not chromatography-purified lipid A (the LPS component recognized by TLR4) induces TLR2 signalling15,91,92.
Early studies showed that C. burnetii LPS is ~1,000-fold less endotoxic than the LPS from enteric species such as Escherichia coli93. It was subsequently discovered that C. burnetii uses a second LPS-dependent intrinsic immune evasion strategy: rather than serving as an agonist of TLR4 signalling, C. burnetii lipid A acts as a TLR4 antagonist91. The lipid A portion of C. burnetii LPS (both phase I and phase II variants) is composed of a tetra-acylated structure, a form of lipid A that is associated with antagonism of TLR4 signalling in several other bacterial pathogens, including Yersinia pestis94–96. Elucidating the role of TLR4 in C. burnetii pathogenesis was complicated by the observation that this cell surface receptor seems to be involved in actin rearrangement and phagocytosis of virulent phase I C. burnetii27. However, studies suggest that the ability of TLR4 to distinguish between the LPS of phase I and phase II variants is not dependent on the structure of lipid A, but depends on novel signalling mechanisms involving O antigen-mediated phagocytosis (the O antigen is not present in phase II LPS)27,91. Further studies will be required to resolve these findings, especially because the transcriptional profiles of host cells infected with either phase I or phase II C. burnetii do not show expression of genes required for TLR4 signalling97.
The ability of C. burnetii to evade detection by pathogen recognition receptors prevents the activation of infected macrophages and provides a replication-permissive intracellular niche. A number of studies have shown that the activation of infected cells via interferon- (IFN ) inhibits C. burnetii replication98–101 owing primarily to the production of reactive oxygen species (ROS) and reactive nitrogen species (RNS)102. C. burnetii prevents the production of ROS via the secretion of an acid phosphatase (CBU0335); the mechanism of action has yet to be confirmed, but it is likely that this protein prevents assembly of the NADPH oxidase complex, although this mechanism would be insufficient to protect against the high level of ROS that is induced by IFNγ103,104. RNS production requires de novo synthesis of inducible nitric oxide synthase (iNOS), which occurs after the induction of pro-inflammatory cytokines such as those produced after TLR stimulation105. Indeed, in contrast to E. coli LPS, which stimulates macrophages to secrete nitrates, exposure of macrophages to C. burnetii does not lead to nitrate secretion, which is not surprising considering that C. burnetii invades cells without stimulating TLRs97.
Host cell death is often induced after infection as an immune defence mechanism50. This serves two purposes: it eliminates the infected cell, and it allows the infected apoptotic blebs to be engulfed by DCs and the subsequent presentation of antigens through major histocompatibility complex (MHC) class I molecules, which induces protective immunity to intracellular pathogens50. Thus, the ability of C. burnetii to inhibit apoptosis represents an immune evasion strategy. Autophagy is another mechanism used by the innate immune system to remove intracellular pathogens. However, as mentioned above, C. burnetii actively recruits autophagy components to the CCV, and the induction of autophagy actually promotes C. burnetii intracellular replication3,32,106.
Conclusions
C. burnetii is ubiquitous in the environment, has a wide host range and has the potential to cause epidemic outbreaks, owing in part to its aerosol transmission and extreme stability. Thus, understanding the pathogenic mechanisms of this organism is a high research priority. From our understanding of the intracellular trafficking pathway and the host processes that are activated or suppressed during infection, it is clear that C. burnetii has adapted to a low-pH CCV and inhibits activation of its primary host cells, macrophages, by evading recognition by the innate immune system.
Research on C. burnetii is currently undergoing a renaissance. The long-standing idea that LPS is the sole virulence determinant is now considered invalid, and there is a growing appreciation for the essential role of a functional T4SS to promote intracellular growth. The C. burnetii T4SS transports more than 100 established substrates, but those that are essential for intracellular replication remain to be determined. Recent reports suggest that, compared with L. pneumophila, C. burnetii has several essential effectors that are secreted via the T4SS, demonstrating that there is less redundancy in C. burnetii than in L. pneumophila. High-throughput screening of random and defined mutant libraries is ongoing and will surely identify the next series of essential genes for intracellular growth in vitro and in vivo. Finally, the development of site-specific mutagenesis methods now allows the roles of predicted pathways and factors to be validated in order to gain an increasingly refined understanding of the molecular pathogenesis of Q fever.
Acknowledgments
This work was supported by US National Institutes of Health grants AI037744, U54 AI057156, AI078213, AI088430, AI090142, and AI092153 (to J.E.S.). The authors thank L. R. Hendrix, K. Russell-Lodrigue and C. Farris for critical review and helpful discussions.
Glossary
Axenic culture media
Media used for the growth of intracellular bacteria in the absence of host cells
Category B select agent
An infectious disease agent of the second highest priority, as defined by the US CDC guidelines for potential misuse. These agents are usually readily transmitted by aerosol, are stable in the environment and cause moderate morbidity and low mortality
BL3 containment
A level of biocontainment that includes a separation from general traffic areas by double doors, airlocks and negative air flow. Access is limited to trained personnel and requires users to wear personal protective equipment. All research activities with potential exposure of the agent to the atmosphere are conducted within biosafety cabinets
Integrin
A highly conserved family of heterodimeric surface glycoproteins involved in binding to extracellular matrix components such as fibronectin and vitronectin through arginine-glycine-aspartic acid (RGD) domains
BL2 containment
A level of biocontainment that involves the use of standard laboratory space in which work using an infectious agent is carried out within a biosafety cabinet
SRC tyrosine kinases
A family of kinases that was originally identified through homology to Rous sarcoma virus oncogene v-src and is involved in signal transduction from cellular receptors
Cachexia
Loss of weight, fatigue and weakness that are associated with severe inflammatory disease and cannot be reversed by nutritional supplementation
_Himar1_-based transposon system
A eukaryotic horn fly element that is extensively used to create mutations in bacteria and relies on only an AT dinucleotide for insertion
C. burnetii str. Nine Mile I
The original Coxiella burnetii strain isolated from ticks in 1935. This strain was later serially passaged in embryonated hen eggs and guinea pigs to obtain the avirulent isolate C. burnetii str. Nine Mile II
Two-component regulatory system
A bacterial signal transduction system involving a sensor kinase that responds to an environmental stimulus by phosphorylating a response regulator, which controls the transcription of downstream genes
Insertion sequences
Mobile genetic elements consisting of short inverted repeats flanking one or more ORFs
Ankyrin repeats
Eukaryotic protein domains consisting of repeating segments of 33 amino acids that form a helix–turn–helix motif and mediate protein–protein interactions. These domains are some of the most commonly found domains in eukaryotic proteins
Coiled-coil domains
Structural motifs that are found in proteins and consist of 2–5 α-helices wrapped around each other in a left-handed manner to form a superhelix
Tetratricopeptide repeats
Structural motifs that mediate protein–protein interactions and are composed of a degenerate ~34 amino acid sequence that is often arranged in a tandem array
F-box domains
Structural motifs composed of approximately 50 amino acids and that contain tryptophan-aspartic acid repeats. These domains function as protein–protein interaction domains. F-box proteins were first characterized as components of ubiquitin ligase complexes
Fic domains
(Filamentation induced by cyclic AMP domains). Protein domains that mediate ampylation of proteins and regulate protein function
Footnotes
Competing interests statement
The authors declare no competing financial interests.
References
- 1.Flannagan RS, Jaumouillé V, Grinstein S. The cell biology of phagocytosis. Annu Rev Pathol. 2012;7:61–98. doi: 10.1146/annurev-pathol-011811-132445. [DOI] [PubMed] [Google Scholar]
- 2.Flannagan RS, Cosio G, Grinstein S. Antimicrobial mechanisms of phagocytes and bacterial evasion strategies. Nature Rev Microbiol. 2009;7:355–366. doi: 10.1038/nrmicro2128. [DOI] [PubMed] [Google Scholar]
- 3.Beron W, Gutierrez MG, Rabinovitch M, Colombo MI. Coxiella burnetii localizes in a Rab7- labeled compartment with autophagic characteristics. Infect Immun. 2002;70:5816–5821. doi: 10.1128/IAI.70.10.5816-5821.2002. The first report to show that CCV formation is disrupted by autophagy inhibitors and that LC3 localizes to the CCV. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Howe D, Mallavia LP. Coxiella burnetii exhibits morphological change and delays phagolysosomal fusion after internalization by J774A.1 cells. Infect Immun. 2000;68:3815–3821. doi: 10.1128/iai.68.7.3815-3821.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Howe D, Melnicâakova J, Barâak I, Heinzen RA. Fusogenicity of the Coxiella burnetii parasitophorous vacuole. Ann NY Acad Sci. 2003;990:556–562. doi: 10.1111/j.1749-6632.2003.tb07426.x. [DOI] [PubMed] [Google Scholar]
- 6.Coleman SA, Fischer ER, Howe D, Mead DJ, Heinzen RA. Temporal analysis of Coxiella burnetii morphological differentiation. J Bacteriol. 2004;186:7344–7352. doi: 10.1128/JB.186.21.7344-7352.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Coleman SA, et al. Proteome and antigen profiling of Coxiella burnetii developmental forms. Infect Immun. 2007;75:290–298. doi: 10.1128/IAI.00883-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Stoker MBP, Fiset P. Phase variation of the Nine Mile and other strains of Rickettsia burnetii. Can J Microbiol. 1956;2:310–321. doi: 10.1139/m56-036. [DOI] [PubMed] [Google Scholar]
- 9.Baca OG, Klassen DA, Aragon AS. Entry of Coxiella burnetii into host cells. Acta Virol. 1993;37:143–155. [PubMed] [Google Scholar]
- 10.Tujulin E, Macellaro A, Lilliehook B, Norlander L. Effect of endocytosis inhibitors on Coxiella burnetii interaction with host cells. Acta Virol. 1998;42:125–131. [PubMed] [Google Scholar]
- 11.Seshadri R, et al. Complete genome sequence of the Q-fever pathogen Coxiella burnetii. Proc Natl Acad Sci USA. 2003;100:5455–5460. doi: 10.1073/pnas.0931379100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Beare PA, et al. Dot/Icm type IVB secretion system requirements for Coxiella burnetii growth in human macrophages. mBio. 2011;2:e00175–11. doi: 10.1128/mBio.00175-11. Work that determines the essential role of the T4SS and the temporal control of the C. burnetii T4SS. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Carey KL, Newton HJ, Luhrmann A, Roy CR. The Coxiella burnetii Dot/Icm system delivers a unique repertoire of type IV effectors into host cells and is required for intracellular replication. PLoS Pathog. 2011;7:e1002056. doi: 10.1371/journal.ppat.1002056. A paper which demonstrates that CCV biogenesis and C. burnetii intracellular replication require a functional T4SS. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Capo C, et al. Subversion of monocyte functions by Coxiella burnetii: impairment of the cross-talk between αvβ3 integrin and CR3. J Immunol. 1999;163:6078–6085. [PubMed] [Google Scholar]
- 15.Dellacasagrande J, et al. αvβ3 integrin and bacterial lipopolysaccharide are involved in Coxiella burnetii-stimulated production of tumor necrosis factor by human monocytes. Infect Immun. 2000;68:5673–5678. doi: 10.1128/iai.68.10.5673-5678.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Dupuy AG, Caron E. Integrin-dependent phagocytosis – spreading from microadhesion to new concepts. J Cell Sci. 2008;121:1773–1783. doi: 10.1242/jcs.018036. [DOI] [PubMed] [Google Scholar]
- 17.De Fougerolles AR, Koteliansky VE. Regulation of monocyte gene expression by the extracellular matrix and its functional implications. Immunol Rev. 2002;186:208–220. doi: 10.1034/j.1600-065x.2002.18617.x. [DOI] [PubMed] [Google Scholar]
- 18.Damjanovich L, Albelda SM, Mette SA, Buck CA. Distribution of integrin cell adhesion receptors in normal and malignant lung tissue. Am J Respir Cell Mol Biol. 1992;6:197–206. doi: 10.1165/ajrcmb/6.2.197. [DOI] [PubMed] [Google Scholar]
- 19.Russell-Lodrigue KE, Zhang GQ, McMurray DN, Samuel JE. Clinical and pathologic changes in a guinea pig aerosol challenge model of acute Q fever. Infect Immun. 2006;74:6085–6091. doi: 10.1128/IAI.00763-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Moos A, Hackstadt T. Comparative virulence of intra-and interstrain lipopolysaccharide variants of Coxiella burnetii in the guinea pig model. Infect Immun. 1987;55:1144–1150. doi: 10.1128/iai.55.5.1144-1150.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Howe D, Shannon JG, Winfree S, Dorward DW, Heinzen RA. Coxiella burnetii phase I and II variants replicate with similar kinetics in degradative phagolysosome-like compartments of human macrophages. Infect Immun. 2010;78:3465–3474. doi: 10.1128/IAI.00406-10. An article showing that phase I and phase II C. burnetii cells replicate within similar vacuoles in human-derived macrophages and THP-1 cells, indicating that differences in virulence are not determined by the terminal compartment that these bacteria reside in. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Romano PS, Gutierrez MG, Beron W, Rabinovitch M, Colombo MI. The autophagic pathway is actively modulated by phase II Coxiella burnetii to efficiently replicate in the host cell. Cell Microbiol. 2007;9:891–909. doi: 10.1111/j.1462-5822.2006.00838.x. [DOI] [PubMed] [Google Scholar]
- 23.Fu Y, Galan JE. A Salmonella protein antagonizes Rac-1 and Cdc42 to mediate host-cell recovery after bacterial invasion. Nature. 1999;401:293–297. doi: 10.1038/45829. [DOI] [PubMed] [Google Scholar]
- 24.Subtil A, Wyplosz B, Balana ME, Dautry-Varsat A. Analysis of Chlamydia caviae entry sites and involvement of Cdc42 and Rac activity. J Cell Sci. 2004;117:3923–3933. doi: 10.1242/jcs.01247. [DOI] [PubMed] [Google Scholar]
- 25.Meconi S, et al. Activation of protein tyrosine kinases by Coxiella burnetii: role in actin cytoskeleton reorganization and bacterial phagocytosis. Infect Immun. 2001;69:2520–2526. doi: 10.1128/IAI.69.4.2520-2526.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Meconi S, et al. Coxiella burnetii induces reorganization of the actin cytoskeleton in human monocytes. Infect Immun. 1998;66:5527–5533. doi: 10.1128/iai.66.11.5527-5533.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Honstettre A, et al. Lipopolysaccharide from Coxiella burnetii is involved in bacterial phagocytosis, filamentous actin reorganization, and inflammatory responses through Toll-like receptor 4. J Immunol. 2004;172:3695–3703. doi: 10.4049/jimmunol.172.6.3695. [DOI] [PubMed] [Google Scholar]
- 28.Kinchen JM, Ravichandran KS. Phagosome maturation: going through the acid test. Nature Rev Mol Cell Biol. 2008;9:781–795. doi: 10.1038/nrm2515. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Voth DE, Heinzen RA. Lounging in a lysosome: the intracellular lifestyle of Coxiella burnetii. Cell Microbiol. 2007;9:829–840. doi: 10.1111/j.1462-5822.2007.00901.x. [DOI] [PubMed] [Google Scholar]
- 30.Howe D, Melnicâakovâa J, Barâak I, Heinzen RA. Maturation of the Coxiella burnetii parasitophorous vacuole requires bacterial protein synthesis but not replication. Cell Microbiol. 2003;5:469–480. doi: 10.1046/j.1462-5822.2003.00293.x. The first report indicating that maturation of the CCV is a bacterially driven process. [DOI] [PubMed] [Google Scholar]
- 31.Levine B, Mizushima N, Virgin HW. Autophagy in immunity and inflammation. Nature. 2011;469:323–335. doi: 10.1038/nature09782. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Gutierrez MG, et al. Autophagy induction favours the generation and maturation of the Coxiella-replicative vacuoles. Cell Microbiol. 2005;7:981–993. doi: 10.1111/j.1462-5822.2005.00527.x. [DOI] [PubMed] [Google Scholar]
- 33.McDonough JA, et al. Host pathways important for Coxiella burnetii infection revealed by genome-wide RNA interference screening. mBio. 2013;4:e00606–12. doi: 10.1128/mBio.00606-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Roman MJ, Crissman HA, Samsonoff WA, Hechemy KE, Baca OG. Analysis of Coxiella burnetii isolates in cell culture and the expression of parasite-specific antigens on the host membrane surface. Acta Virol. 1991;35:503–510. [PubMed] [Google Scholar]
- 35.Aguilera M, et al. Actin dynamics and Rho GTPases regulate the size and formation of parasitophorous vacuoles containing Coxiella burnetii. Infect Immun. 2009;77:4609–4620. doi: 10.1128/IAI.00301-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Hussain SK, Broederdorf LJ, Sharma UM, Voth DE. Host kinase activity is required for Coxiella burnetii parasitophorous vacuole formation. Front Microbiol. 2010;1:137. doi: 10.3389/fmicb.2010.00137. The first evidence that host cell kinases are involved in CCV maturation, further defining the host–pathogen interface. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Campoy EM, Zoppino FC, Colombo MI. The early secretory pathway contributes to the growth of the Coxiella-replicative niche. Infect Immun. 2011;79:402–413. doi: 10.1128/IAI.00688-10. An article demonstrating that the CCV interacts with the ER at late time points during infection. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Howe D, Heinzen RA. Coxiella burnetii inhabits a cholesterol-rich vacuole and influences cellular cholesterol metabolism. Cell Microbiol. 2006;8:496–507. doi: 10.1111/j.1462-5822.2005.00641.x. [DOI] [PubMed] [Google Scholar]
- 39.Howe D, Heinzen RA. Replication of Coxiella burnetii is inhibited in CHO K-1 cells treated with inhibitors of cholesterol metabolism. Ann NY Acad Sci. 2005;1063:123–129. doi: 10.1196/annals.1355.020. [DOI] [PubMed] [Google Scholar]
- 40.Espenshade PJ, Hughes AL. Regulation of sterol synthesis in eukaryotes. Annu Rev Genet. 2007;41:401–427. doi: 10.1146/annurev.genet.41.110306.130315. [DOI] [PubMed] [Google Scholar]
- 41.Akporiaye ET, Rowatt JD, Aragon AA, Baca OG. Lysosomal response of a murine macrophage-like cell line persistently infected with Coxiella burnetii. Infect Immun. 1983;40:1155–1162. doi: 10.1128/iai.40.3.1155-1162.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Voth DE, Howe D, Heinzen RA. Coxiella burnetii inhibits apoptosis in human THP-1 cells and monkey primary alveolar macrophages. Infect Immun. 2007;75:4263–4271. doi: 10.1128/IAI.00594-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Baca OG, Scott TO, Akporiaye ET, DeBlassie R, Crissman HA. Cell cycle distribution patterns and generation times of L929 fibroblast cells persistently infected with Coxiella burnetii. Infect Immun. 1985;47:366–369. doi: 10.1128/iai.47.2.366-369.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Luhrmann A, Roy CR. Coxiella burnetii inhibits activation of host cell apoptosis through a mechanism that involves preventing cytochrome c release from mitochondria. Infect Immun. 2007;75:5282–5289. doi: 10.1128/IAI.00863-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Yang J, et al. Prevention of apoptosis by Bcl-2: release of cytochrome c from mitochondria blocked. Science. 1997;275:1129–1132. doi: 10.1126/science.275.5303.1129. [DOI] [PubMed] [Google Scholar]
- 46.Voth DE, Heinzen RA. Sustained activation of Akt and Erk1/2 is required for Coxiella burnetii antiapoptotic activity. Infect Immun. 2009;77:205–213. doi: 10.1128/IAI.01124-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Roman MJ, Coriz PD, Baca OG. A proposed model to explain persistent infection of host cells with Coxiella burnetii. J Gen Microbiol. 1986;132:1415–1422. doi: 10.1099/00221287-132-5-1415. [DOI] [PubMed] [Google Scholar]
- 48.Tigertt WD, Benenson AS, Gochenour WS. Airborne Q fever. Bacteriol Rev. 1961;25:285–293. doi: 10.1128/br.25.3.285-293.1961. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Zhang Y, Zhang G, Hendrix LR, Tesh VL, Samuel JE. Coxiella burnetii induces apoptosis during early stage infection via a caspase-independent pathway in human monocytic THP-1 cells. PLoS ONE. 2012;7:e30841. doi: 10.1371/journal.pone.0030841. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Ashida H, et al. Cell death and infection: a double-edged sword for host and pathogen survival. J Cell Biol. 2011;195:931–942. doi: 10.1083/jcb.201108081. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Beare PA, et al. Comparative genomics reveal extensive transposon-mediated genomic plasticity and diversity among potential effector proteins within the genus Coxiella. Infect Immun. 2009;77:642–656. doi: 10.1128/IAI.01141-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Peabody CR, et al. Type II protein secretion and its relationship to bacterial type IV pili and archaeal flagella. Microbiology. 2003;149:3051–3072. doi: 10.1099/mic.0.26364-0. [DOI] [PubMed] [Google Scholar]
- 53.Sexton JA, Vogel JP. Type IVB secretion by intracellular pathogens. Traffic. 2002;3:178–185. doi: 10.1034/j.1600-0854.2002.030303.x. [DOI] [PubMed] [Google Scholar]
- 54.Zechner EL, Lang S, Schildbach JF. Assembly and mechanisms of bacterial type IV secretion machines. Phil Trans R Soc B. 2012;367:1073–1087. doi: 10.1098/rstb.2011.0207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Alvarez-Martinez CE, Christie PJ. Biological diversity of prokaryotic type IV secretion systems. Microbiol Mol Biol Rev. 2009;73:775–808. doi: 10.1128/MMBR.00023-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Vincent CD, et al. Identification of the core transmembrane complex of the Legionella Dot/Icm type IV secretion system. Mol Microbiol. 2006;62:1278–1291. doi: 10.1111/j.1365-2958.2006.05446.x. [DOI] [PubMed] [Google Scholar]
- 57.Vincent CD, Friedman JR, Jeong KC, Sutherland MC, Vogel JP. Identification of the DotL coupling protein subcomplex of the Legionella Dot/Icm type IV secretion system. Mol Microbiol. 2012;85:378–391. doi: 10.1111/j.1365-2958.2012.08118.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Hiroki N, Tomoko K. Type IVB secretion systems of Legionella and other Gram-negative bacteria. Front Microbiol. 2011;2:136. doi: 10.3389/fmicb.2011.00136. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Zamboni DS, McGrath S, Rabinovitch M, Roy CR. Coxiella burnetii express type IV secretion system proteins that function similarly to components of the Legionella pneumophila Dot/Icm system. Mol Microbiol. 2003;49:965–976. doi: 10.1046/j.1365-2958.2003.03626.x. [DOI] [PubMed] [Google Scholar]
- 60.Voth DE, et al. The Coxiella burnetii ankyrin repeat domain-containing protein family is heterogeneous with C-terminal truncations that influence Dot/Icm-mediated secretion. J Bacteriol. 2009;191:4232–4242. doi: 10.1128/JB.01656-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Chen C, et al. Large-scale identification and translocation of type IV secretion substrates by Coxiella burnetii. Proc Natl Acad Sci USA. 2010;107:21755–21760. doi: 10.1073/pnas.1010485107. Research establishing a shuttle vector and translocation assays for C. burnetii, leading to the identification of a large number of T4SS effectors and confirming the presence of a functional secretion system in C. burnetii during infection. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Morgan JK, Luedtke BE, Thompson HA, Shaw EI. Coxiella burnetii type IVB secretion system region I genes are expressed early during the infection of host cells. FEMS Microbiol Lett. 2010;311:61–69. doi: 10.1111/j.1574-6968.2010.02072.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Morgan JK, Luedtke BE, Shaw EI. Polar localization of the Coxiella burnetii type IVB secretion system. FEMS Microbiol Lett. 2010;305:177–183. doi: 10.1111/j.1574-6968.2010.01926.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Beare PA, Larson CL, Gilk SD, Heinzen RA. Two systems for targeted gene deletion in Coxiella burnetii. Appl Environ Microbiol. 2012;78:4580–4589. doi: 10.1128/AEM.00881-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Pan X, Luhrmann A, Satoh A, Laskowski-Arce MA, Roy CR. Ankyrin repeat proteins comprise a diverse family of bacterial type IV effectors. Science. 2008;320:1651–1654. doi: 10.1126/science.1158160. A paper that identifies ankyrin repeat proteins as T4SS effectors in C. burnetii and L. pneumophila, indicating that these proteins are conserved T4SS effectors. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Voth DE, Heinzen RA. Coxiella type IV secretion and cellular microbiology. Curr Opin Microbiol. 2009;12:74–80. doi: 10.1016/j.mib.2008.11.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Voth DE, et al. The Coxiella burnetii cryptic plasmid is enriched in genes encoding type IV secretion system substrates. J Bacteriol. 2011;193:1493–1503. doi: 10.1128/JB.01359-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Lifshitz Z, et al. Computational modeling and experimental validation of the Legionella and Coxiella virulence-related type-IVB secretion signal. Proc Natl Acad Sci USA. 2013;110:e707–e715. doi: 10.1073/pnas.1215278110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.de Felipe KS, et al. Evidence for acquisition of Legionella type IV secretion substrates via interdomain horizontal gene transfer. J Bacteriol. 2005;187:7716–7726. doi: 10.1128/JB.187.22.7716-7726.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Kubori T, Shinzawa N, Kanuka H, Nagai H. Legionella metaeffector exploits host proteasome to temporally regulate cognate effector. PLoS Pathog. 2011;6:e1001216. doi: 10.1371/journal.ppat.1001216. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Newton HJ, McDonough JA, Roy CR. Effector protein translocation by the Coxiella burnetii Dot/Icm type IV secretion system requires endocytic maturation of the pathogen-occupied vacuole. PLoS ONE. 2013;8:e54566. doi: 10.1371/journal.pone.0054566. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Zusman T, et al. The response regulator PmrA is a major regulator of the icm/dot type IV secretion system in Legionella pneumophila and Coxiella burnetii. Mol Microbiol. 2007;63:1508–1523. doi: 10.1111/j.1365-2958.2007.05604.x. A report showing that components of the T4SS and some putative T4SS effectors are co-regulated, implying that there is a link between, and temporal regulation of, the secretion apparatus and its effectors. [DOI] [PubMed] [Google Scholar]
- 73.McPhee JB, Lewenza S, Hancock RE. Cationic antimicrobial peptides activate a two-component regulatory system, PmrA-PmrB, that regulates resistance to polymyxin B and cationic antimicrobial peptides in Pseudomonas aeruginosa. Mol Microbiol. 2003;50:205–217. doi: 10.1046/j.1365-2958.2003.03673.x. [DOI] [PubMed] [Google Scholar]
- 74.Sauer JD, et al. Specificity of Legionella pneumophila and Coxiella burnetii vacuoles and versatility of Legionella pneumophila revealed by coinfection. Infect Immun. 2005;73:4494–4504. doi: 10.1128/IAI.73.8.4494-4504.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Omsland A, et al. Isolation from animal tissue and genetic transformation of Coxiella burnetii are facilitated by an improved axenic growth medium. Appl Environ Microbiol. 2011;77:3720–3725. doi: 10.1128/AEM.02826-10. The first description of the clonal isolation of transformed C. burnetii using axenic media. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Huang L, et al. The E Block motif is associated with Legionella pneumophila translocated substrates. Cell Microbiol. 2011;13:227–245. doi: 10.1111/j.1462-5822.2010.01531.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Klingenbeck L, Eckart RA, Berens C, Luhrmann A. The Coxiella burnetii type IV secretion system substrate CaeB inhibits intrinsic apoptosis at the mitochondrial level. Cell Microbiol. 2013;15:675–687. doi: 10.1111/cmi.12066. [DOI] [PubMed] [Google Scholar]
- 78.Luhrmann A, Nogueira CV, Carey KL, Roy CR. Inhibition of pathogen-induced apoptosis by a Coxiella burnetii type IV effector protein. Proc Natl Acad Sci USA. 2010;107:18997–19001. doi: 10.1073/pnas.1004380107. An article which describes the only characterized C. burnetii T4SS effector that prevents host cell apoptosis by targeting pro-apoptotic protein p32. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Ge J, et al. A Legionella type IV effector activates the NF-κB pathway by phosphorylating the IκB family of inhibitors. Proc Natl Acad Sci USA. 2009;106:13725–13730. doi: 10.1073/pnas.0907200106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.de Felipe KS, et al. Legionella eukaryotic-like type IV substrates interfere with organelle trafficking. PLoS Pathog. 2008;4:e1000117. doi: 10.1371/journal.ppat.1000117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Shen X, et al. Targeting eEF1A by a Legionella pneumophila effector leads to inhibition of protein synthesis and induction of host stress response. Cell Microbiol. 2009;11:911–926. doi: 10.1111/j.1462-5822.2009.01301.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Banga S, et al. Legionella pneumophila inhibits macrophage apoptosis by targeting pro-death members of the Bcl2 protein family. Proc Natl Acad Sci USA. 2007;104:5121–5126. doi: 10.1073/pnas.0611030104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Campodonico EM, Chesnel L, Roy CR. A yeast genetic system for the identification and characterization of substrate proteins transferred into host cells by the Legionella pneumophila Dot/Icm system. Mol Microbiol. 2005;56:918–933. doi: 10.1111/j.1365-2958.2005.04595.x. [DOI] [PubMed] [Google Scholar]
- 84.Ren Q, Robertson SJ, Howe D, Barrows LF, Heinzen RA. Comparative DNA microarray analysis of host cell transcriptional responses to infection by Coxiella burnetii or Chlamydia trachomatis. Ann NY Acad Sci. 2003;990:701–713. doi: 10.1111/j.1749-6632.2003.tb07447.x. [DOI] [PubMed] [Google Scholar]
- 85.Murata T, et al. The Legionella pneumophila effector protein DrrA is a Rab1 guanine nucleotide-exchange factor. Nature Cell Biol. 2006;8:971–977. doi: 10.1038/ncb1463. [DOI] [PubMed] [Google Scholar]
- 86.Machner MP, Isberg RR. Targeting of host Rab GTPase function by the intravacuolar pathogen Legionella pneumophila. Dev Cell. 2006;11:47–56. doi: 10.1016/j.devcel.2006.05.013. [DOI] [PubMed] [Google Scholar]
- 87.Nagai H, Kagan JC, Zhu X, Kahn RA, Roy CR. A bacterial guanine nucleotide exchange factor activates ARF on Legionella phagosomes. Science. 2002;295:679–682. doi: 10.1126/science.1067025. [DOI] [PubMed] [Google Scholar]
- 88.Price CT, Al-Quadan T, Santic M, Rosenshine I, Abu Kwaik Y. Host proteasomal degradation generates amino acids essential for intracellular bacterial growth. Science. 2011;334:1553–1557. doi: 10.1126/science.1212868. [DOI] [PubMed] [Google Scholar]
- 89.Vishwanath S, Hackstadt T. Lipopolysaccharide phase variation determines the complement-mediated serum susceptibility of Coxiella burnetii. Infect Immun. 1988;56:40–44. doi: 10.1128/iai.56.1.40-44.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Shannon JG, Howe D, Heinzen RA. Virulent Coxiella burnetii does not activate human dendritic cells: role of lipopolysaccharide as a shielding molecule. Proc Natl Acad Sci USA. 2005;102:8722–8727. doi: 10.1073/pnas.0501863102. A study showing that lipopolysaccharide is used by virulent C. burnetii to evade immune surveillance. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Zamboni DS, et al. Stimulation of toll-like receptor 2 by Coxiella burnetii is required for macrophage production of pro-inflammatory cytokines and resistance to infection. J Biol Chem. 2004;279:54405–54415. doi: 10.1074/jbc.M410340200. The first report to indicate that lipid A from either phase I or phase II C. burnetii cannot stimulate TLR4. [DOI] [PubMed] [Google Scholar]
- 92.Hirschfeld M, Ma Y, Weis JH, Vogel SN, Weis JJ. Cutting edge: repurification of lipopolysaccharide eliminates signaling through both human and murine Toll-like receptor 2. J Immunol. 2000;165:618–622. doi: 10.4049/jimmunol.165.2.618. [DOI] [PubMed] [Google Scholar]
- 93.Amano K, Williams JC, Missler SR, Reinhold VN. Structure and biological relationships of Coxiella burnetii lipopolysaccharide. J Biol Chem. 1987;262:4740–4747. [PubMed] [Google Scholar]
- 94.Stoenner HG, Lackman DB. The biologic properties of Coxiella burnetii isolated from rodents collected in Utah. Am J Hyg. 1960;71:45–51. doi: 10.1093/oxfordjournals.aje.a120088. [DOI] [PubMed] [Google Scholar]
- 95.Toman R, et al. Physicochemical characterization of the endotoxins from Coxiella burnetii strain Priscilla in relation to their bioactivities. BMC Biochem. 2004;5:1. doi: 10.1186/1471-2091-5-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Telepnev MV, et al. Tetraacylated lipopolysaccharide of Yersinia pestis can inhibit multiple Toll-like receptor-mediated signaling pathways in human dendritic cells. J Infect Dis. 2009;200:1694–1702. doi: 10.1086/647986. [DOI] [PubMed] [Google Scholar]
- 97.Benoit M, Barbarat B, Bernard A, Olive D, Mege JL. Coxiella burnetii, the agent of Q fever, stimulates an atypical M2 activation program in human macrophages. Eur J Immunol. 2008;38:1065–1070. doi: 10.1002/eji.200738067. [DOI] [PubMed] [Google Scholar]
- 98.Dellacasagrande J, Capo C, Raoult D, Mege JL. IFN-γ-mediated control of Coxiella burnetii survival in monocytes: the role of cell apoptosis and TNF. J Immunol. 1999;162:2259–2265. [PubMed] [Google Scholar]
- 99.Howe D, Barrows LF, Lindstrom NM, Heinzen RA. Nitric oxide inhibits Coxiella burnetii replication and parasitophorous vacuole maturation. Infect Immun. 2002;70:5140–5147. doi: 10.1128/IAI.70.9.5140-5147.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Turco J, Thompson HA, Winkler H. Interferon-γ inhibits growth of Coxiella burnetii in mouse fibroblasts. Infect Immun. 1984;45:781–783. doi: 10.1128/iai.45.3.781-783.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Zamboni DS, Rabinovitch M. Nitric oxide partially controls Coxiella burnetii phase II infection in mouse primary macrophages. Infect Immun. 2003;71:1225–1233. doi: 10.1128/IAI.71.3.1225-1233.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Brennan RE, Russell K, Zhang G, Samuel JE. Both inducible nitric oxide synthase and NADPH oxidase contribute to the control of virulent phase I Coxiella burnetii infections. Infect Immun. 2004;72:6666–6675. doi: 10.1128/IAI.72.11.6666-6675.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Hill J, Samuel JE. Coxiella burnetii acid phosphatase inhibits the release of reactive oxygen intermediates in polymorphonuclear leukocytes. Infect Immun. 2011;79:414–420. doi: 10.1128/IAI.01011-10. Work identifying a specific protein that mediates the inhibition of ROS release in infected polymorphonuclear neutrophils. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Siemsen DW, Kirpotina LN, Jutila MA, Quinn MT. Inhibition of the human neutrophil NADPH oxidase by Coxiella burnetii. Microbes Infect. 2009;11:671–679. doi: 10.1016/j.micinf.2009.04.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Vila-del Sol V, Diaz-Munoz MD, Fresno M. Requirement of tumor necrosis factor α and nuclear factor-κB in the induction by IFN-γ of inducible nitric oxide synthase in macrophages. J Leukoc Biol. 2007;81:272–283. doi: 10.1189/jlb.0905529. [DOI] [PubMed] [Google Scholar]
- 106.Vazquez CL, Colombo MI. Coxiella burnetii modulates Beclin 1 and Bcl-2, preventing host cell apoptosis to generate a persistent bacterial infection. Cell Death Differ. 2010;17:421–438. doi: 10.1038/cdd.2009.129. [DOI] [PubMed] [Google Scholar]
- 107.Babudieri C. Q fever: a zoonosis. Adv Vet Sci. 1959;5:81–84. [Google Scholar]
- 108.CDC & Department of Health and Human Services. Possession, use, and transfer of select agents and toxins. Final rule. Fed Regist. 2008;73:61363–61366. [PubMed] [Google Scholar]
- 109.Fournier PE, Marrie TJ, Raoult D. Diagnosis of Q fever. J Clin Microbiol. 1998;36:1823–1834. doi: 10.1128/jcm.36.7.1823-1834.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Million M, Thuny F, Richet H, Raoult D. Long-term outcome of Q fever endocarditis: a 26-year personal survey. Lancet Infect Dis. 2010;10:527–535. doi: 10.1016/S1473-3099(10)70135-3. [DOI] [PubMed] [Google Scholar]
- 111.Raoult D, et al. Treatment of Q fever endocarditis: comparison of 2 regimens containing doxycycline and ofloxacin or hydroxychloroquine. Arch Intern Med. 1999;159:167–173. doi: 10.1001/archinte.159.2.167. [DOI] [PubMed] [Google Scholar]
- 112.Marmion BP, et al. Long-term persistence of Coxiella burnetii after acute primary Q fever. QJM. 2005;98:7–20. doi: 10.1093/qjmed/hci009. [DOI] [PubMed] [Google Scholar]
- 113.Penttila IA, et al. Cytokine dysregulation in the post-Q-fever fatigue syndrome. QJM. 1998;91:549–560. doi: 10.1093/qjmed/91.8.549. [DOI] [PubMed] [Google Scholar]
- 114.Marmion BP, Shannon M, Maddocks I, Storm P, Penttila I. Protracted debility and fatigue after acute Q fever. Lancet. 1996;347:977–978. doi: 10.1016/s0140-6736(96)91469-5. [DOI] [PubMed] [Google Scholar]
- 115.Enserink M. Infectious diseases. Questions abound in Q-fever explosion in the Netherlands. Science. 2010;327:266–267. doi: 10.1126/science.327.5963.266-a. [DOI] [PubMed] [Google Scholar]
- 116.Whelan J, et al. Q fever among culling workers, the Netherlands, 2009–2010. Emerg Infect Dis. 2011;17:1719–1723. doi: 10.3201/eid1709.110051. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 117.Roest HI, et al. The Q fever epidemic in The Netherlands: history, onset, response and reflection. Epidemiol Infect. 2011;139:1–12. doi: 10.1017/S0950268810002268. [DOI] [PubMed] [Google Scholar]
- 118.Glazunova O, et al. Coxiella burnetii genotyping. Emerg Infect Dis. 2005;11:1211–1217. doi: 10.3201/eid1108.041354. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Hendrix LR, Samuel JE, Mallavia LP. Differentiation of Coxiella burnetii isolates by analysis of restriction-endonuclease-digested DNA separated by SDS-PAGE. J Gen Microbiol. 1991;137:269–276. doi: 10.1099/00221287-137-2-269. [DOI] [PubMed] [Google Scholar]
- 120.Samuel JE, Frazier ME, Mallavia LP. Correlation of plasmid type and disease caused by Coxiella burnetii. Infect Immun. 1985;49:775–779. doi: 10.1128/iai.49.3.775-779.1985. The proposal that pathotype-specific differences exist between clinical isolates of C. burnetii. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Svraka S, Toman R, Skultety L, Slaba K, Homan WL. Establishment of a genotyping scheme for Coxiella burnetii. FEMS Microbiol Lett. 2006;254:268–274. doi: 10.1111/j.1574-6968.2005.00036.x. [DOI] [PubMed] [Google Scholar]
- 122.Thiele D, Willems H. Is plasmid based differentiation of Coxiella burnetii in ‘acute’ and ‘chronic’ isolates still valid? Eur J Epidemiol. 1994;10:427–434. doi: 10.1007/BF01719667. [DOI] [PubMed] [Google Scholar]
- 123.Beare PA, et al. Genetic diversity of the Q fever agent, Coxiella burnetii, assessed by microarray-based whole-genome comparisons. J Bacteriol. 2006;188:2309–2324. doi: 10.1128/JB.188.7.2309-2324.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124.Russell-Lodrigue KE, et al. Coxiella burnetii isolates cause genogroup-specific virulence in mouse and guinea pig models of acute Q fever. Infect Immun. 2009;77:5640–5650. doi: 10.1128/IAI.00851-09. An investigation that clearly establishes pathotype differences in animal models of acute Q fever. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 125.Capo C, et al. Production of interleukin-10 and transforming growth factor β by peripheral blood mononuclear cells in Q fever endocarditis. Infect Immun. 1996;64:4143–4147. doi: 10.1128/iai.64.10.4143-4147.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 126.Meghari S, et al. Persistent Coxiella burnetii infection in mice overexpressing IL-10: an efficient model for chronic Q fever pathogenesis. PLoS Pathog. 2008;4:e23. doi: 10.1371/journal.ppat.0040023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 127.Chen SY, Hoover TA, Thompson HA, Williams JC. Characterization of the origin of DNA replication of the Coxiella burnetii chromosome. Ann NY Acad Sci. 1990;590:491–503. doi: 10.1111/j.1749-6632.1990.tb42259.x. [DOI] [PubMed] [Google Scholar]
- 128.Suhan M, et al. Cloning and characterization of an autonomous replication sequence from Coxiella burnetii. J Bacteriol. 1994;176:5233–5243. doi: 10.1128/jb.176.17.5233-5243.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 129.Hackstadt T, Williams JC. Biochemical stratagem for obligate parasitism of eukaryotic cells by Coxiella burnetii. Proc Natl Acad Sci USA. 1981;78:3240–3244. doi: 10.1073/pnas.78.5.3240. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 130.Omsland A, et al. Host cell-free growth of the Q fever bacterium Coxiella burnetii. Proc Natl Acad Sci USA. 2009;106:4430–4434. doi: 10.1073/pnas.0812074106. Work which characterizes the cell-free growth medium of C. burnetii and determines the anaerobic and acidophilic tropism of this bacterium. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 131.Beare PA, et al. Characterization of a Coxiella burnetii ftsZ mutant generated by Himar1 transposon mutagenesis. J Bacteriol. 2009;191:1369–1381. doi: 10.1128/JB.01580-08. An article detailing the Himar1-based transposon system for C. burnetii, building the foundation for forward genetics screens. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 132.Beare P, Sandoz K, Omsland A, Rockey D, Heinzen R. Advances in genetic manipulation of obligate intracellular bacterial pathogens. Front Microbiol. 2011;2:97. doi: 10.3389/fmicb.2011.00097. [DOI] [PMC free article] [PubMed] [Google Scholar]