Identification of Zika virus epitopes reveals immunodominant and protective roles for dengue virus cross-reactive CD8+ T cells (original) (raw)
. Author manuscript; available in PMC: 2018 Apr 27.
Abstract
CD8+ T cells play an important role in controlling Flavivirus infection, including Zika virus (ZIKV). Here, we have identified 25 HLA-B*0702-restricted epitopes and 1 HLA-A*0101-restricted epitope using interferon (IFN)-γ enzyme-linked immunospot (ELISPOT) and intracellular cytokine staining (ICS) in ZIKV-infected IFN-α/β receptor-deficient HLA transgenic mice. The cross-reactivity of ZIKV epitopes to dengue virus (DENV) was tested using IFN-γ-ELISPOT and IFN-γ-ICS on CD8+ T cells from DENV-infected mice, and five cross-reactive HLA-B*0702-binding peptides were identified by both assays. ZIKV/DENV cross-reactive CD8+ T cells in DENV-immune mice expanded post ZIKV challenge and dominated in the subsequent CD8+ T cell response. ZIKV challenge following immunization of mice with ZIKV-specific and ZIKV/DENV cross-reactive epitopes elicited CD8+ T cell responses that reduced infectious ZIKV levels, and CD8+ T cell depletions confirmed that CD8+ T cells mediated this protection. These results identify ZIKV-specific and ZIKV/DENV cross-reactive epitopes and demonstrate both an altered immunodominance pattern in the DENV-immune setting relative to naive, as well as a protective role for epitope-specific CD8+ T cells against ZIKV. These results have important implications for ZIKV vaccine development and provide a mouse model for evaluating anti-ZIKV CD8+ T cell responses of human relevance.
Zika virus (ZIKV) is a positive-sense, single-stranded, enveloped RNA flavivirus that shares vector and host space with other flaviviruses, including dengue virus (DENV), yellow fever virus and Japanese encephalitis virus1,2. ZIKV was first isolated in 1947 in Uganda from a sentinel rhesus macaque and, until recently, was known to cause mild, self-limiting and sporadic disease in Africa and Southeast Asia3. A causal relationship between ZIKV and a congenital syndrome including microcephaly was confirmed in the 2015 Brazilian outbreak, and signs of microcephaly have been seen in ZIKV-infected mice4–6. ZIKV has also been linked to Guillain–Barre syndrome7 and case reports of sexual transmission are mounting8,9.
With the new disease syndromes caused by and associated with ZIKV infection, there is an urgent need to address fundamental gaps in the understanding of ZIKV immunology and pathogenesis. Signs of clinical Zika disease have historically been similar to signs of dengue fever, and ZIKV’s immunological similarity to DENV has also been documented. BLAST search results show that ZIKV and DENV have about 52–57% amino acid sequence homology. Indeed, the serologic cross-reactivity of these two viruses has probably contributed to the misdiagnosis and underdiagnosis of ZIKV, and cases of concurrent infection with ZIKV and DENV have also been documented10. Cellular immunity to flaviviruses is also cross-reactive, and cross-reactive T cells may play a dual role in protection and pathogenesis11–13. However, so far, ZIKV epitopes recognized by human CD4+ or CD8+ T cells have not been identified, and their identification would accelerate the investigation of immunity and pathogenesis, as well as the development of vaccines and potentially diagnostics.
Epidemiological and laboratory studies from the relatively large body of knowledge on the four serotypes of DENV indicate that the severe and potentially fatal form of dengue disease occurs most commonly when patients are infected with a second DENV serotype after infection by, and recovery from, a first heterologous DENV serotype14,15. One hypothesis, termed ‘original T cell antigenic sin’, suggests that disease severity increases in secondary infection because T cells primed during the first DENV infection predominate in the subsequent infection with a different DENV serotype, and these serotype-cross-reactive T cells fail to mount an appropriate immune response to the second DENV serotype11–13. Similar T cell cross-reactivity may exist between ZIKV and DENV, as ZIKV and DENV share high amino acid identity. Consistent with this homology, several recent studies have revealed cross-reactivity between ZIKV and DENV at the antibody response level. In particular, both plasma and monoclonal antibodies isolated from DENV-exposed donors can have potent neutralizing activity against ZIKV, and can mediate antibody-dependent enhancement (ADE) of ZIKV infection16–18. In fact, monoclonal antibodies isolated from ZIKV-immune donors can induce ADE of DENV infection in vitro and in vivo in mice19. Although our recent work demonstrated that CD8+ T cells are protective against ZIKV infection in H-2b mice20, the understanding of T cell-mediated responses to ZIKV remains minimal. As ZIKV and DENV will continue to co-circulate in many regions of the world due to their common vectors and geographical distributions, it is critical to continue exploring the protective versus potentially pathogenic influence of T cells induced by prior DENV exposure on ZIKV infection. However, knowledge about the human T cell epitopes that are unique to ZIKV or shared with DENV is lacking. As a consequence, few tools are available for investigating ZIKV-specific T cell immunity and vaccine development.
In the present study, we computationally predicted 107 and 90 ZIKV epitopes to bind two common human leucocyte antigen (HLA) class I molecules, HLA-B*0702 and HLA-A*0101, respectively, and screened these epitopes by IFN-γ-enzyme-linked immunospot (ELISPOT) analysis of epitope-stimulated CD8+ T cells from IFN-α/β receptor (Ifnar)−/− HLA-B*0702 and _Ifnar_−/− HLA-A*0101 transgenic mice that were infected with either ZIKV strain FSS13025 (Asian lineage) or MR766 (African lineage). Twenty-five HLA-B*0702-restricted epitopes and 1 HLA-A*0101-restricted epitope were confirmed to induce IFN-γ production by intracellular cytokine staining (ICS). We performed a similar analysis with DENV serotype 2 (DENV2) strain S221 and identified five ZIKV/DENV cross-reactive HLA-B*0702-restricted epitopes. Prior infection with DENV2 shaped the subsequent anti-ZIKV CD8+ T cell response. Specifically, the memory ZIKV/DENV cross-reactive CD8+ T cells expanded during subsequent ZIKV infection and thus the CD8+ T cell response to ZIKV in DENV-immune mice was narrow (focused on the cross-reactive epitopes). In contrast, the anti-ZIKV CD8+ T cell response in naive animals was broad (included both ZIKV-specific and ZIKV/DENV cross-reactive epitopes). Immunization of mice with a cocktail of ZIKV-specific or ZIKV/DENV cross-reactive peptides afforded protection against ZIKV challenge. CD8+ T cell depletion of peptide-immunized mice confirmed that this protection was mediated by CD8+ T cells. Overall, we identified human-relevant ZIKV-derived epitopes recognized by CD8+ T cells in naive and DENV-immune settings, revealed an altered immunodominance pattern of the anti-ZIKV CD8+ T cell response in naive versus DENV-immune conditions, and demonstrated that ZIKV-specific and ZIKV/DENV cross-reactive CD8+ T cells can contribute to protection against ZIKV. These findings imply that ZIKV vaccine development efforts should elicit and evaluate CD8+ T cell responses.
Results
Identification of HLA-B*0702- and HLA-A*0101-restricted ZIKV-derived epitopes
We previously generated in vivo models of DENV infection in HLA transgenic _Ifnar_−/− mice21,22. _Ifnar_−/− mice were used instead of wild-type mice, because DENV cannot block type I IFN signalling and replicate in murine cells23–25. Our HLA transgenic _Ifnar_−/− mouse models of DENV infection have been validated by several observations. (1) The epitopes identified in mice are also recognized by peripheral blood mononuclear cells (PBMCs) from DENV-exposed humans21,22. (2) A dominance of the HLA-B*0702-restricted response is observed in both mice and humans21,26. (3) The CD8+ T cell response targets both structural and nonstructural (NS) proteins in DENV3, but predominantly NS proteins in the other three DENV serotypes in both mice and humans22,27,28. (4) CD8+ T cell responses are broad (targeting both structural and NS proteins) following primary and homotypic secondary DENV infection in both mice and humans, whereas CD8+ T cell responses following heterotypic secondary infection in mice22 and natural reinfections in humans26,29,30 focus on the conserved NS proteins. Similar to DENV, ZIKV cannot evade type I IFN-mediated immunity31–33. We therefore used _Ifnar_−/− HLA-B*0702 and HLA-A*0101 transgenic mice to identify ZIKV-derived HLA-restricted epitopes.
A total of 107 HLA-B*0702-binding epitope candidates (8-, 9-, 10- and 11-mers), representing the top 2% of candidates predicted by the Immune Epitope Database and Analysis Resource (IEDAR), were chosen for synthesis. The numbers of peptides in C, prM, M, E, NS1, NS2A, NS2B, NS3, NS4A, NS4B and NS5 were 3, 2, 6, 13, 12, 12, 3, 23, 4, 16 and 13, respectively. Seven days after infection of _Ifnar_−/− HLA-B*0702 transgenic mice with ZIKV strain FSS13025 or MR766, CD8+ T cells were isolated from splenocytes and screened by IFN-γ ELISPOT assay. ZIKV MR766 infection induced a stronger and broader CD8+ T cell response than ZIKV FSS13025 in _Ifnar_−/− HLA-B*0702 transgenic mice (Supplementary Fig. 1a–c). The frequencies of peptide-specific IFN-γ-producing CD8+ T cells in mice infected with ZIKV FSS13025 and ZIKV MR766 ranged from 140 to 1,223 spot-forming cells (SFCs) per 106 CD8+ T cells and 98 to 1,362 SFCs per 106 CD8+ T cells, respectively. Table 1 shows the key characteristics of the positive peptides. A total of 19 and 36 epitopes were derived from ZIKV FSS13025 and ZIKV MR766, respectively. The two ZIKV strains shared 18 epitopes, including the following 9 immunodominant peptides: FSS/MR-NS2A75–84, FSS/MR-NS2A89–99, FSS-NS2A133–141, MR-NS2A133–141, FSS/MR-NS2A148–155, FSS/MR-NS2B68–75, FSS/MR-NS3206–215, FSS/MR-NS3574–582 and FSS/MR-NS4B426–435 (Table 1). The FSS-C25–35 epitope is present in ZIKV FSS13025 but not ZIKV MR766. The NS3 protein contained the largest number of epitopes and NS2A induced the highest magnitude of CD8+ T cell responses. The majority of immunodominant epitopes were located in NS2A, but the most immunodominant epitope, FSS/MR-NS4B426–435, was contained in NS4B (Supplementary Fig. 1a–c).
Table 1.
Characteristics of identified Zika virus epitopes.
| Peptides* | Sequences | HLA | IEDB prediction Percentile rank | Conservation† | SFCs per106 CD8+ T cells (% IFN-γ+ CD8+ T cells)‡ | ||||
|---|---|---|---|---|---|---|---|---|---|
| FSS13025 | MR766 | SPH2015 | FSS13025 | MR766 | S221 | ||||
| FSS-C25–35 | SPFGGLKRLPA | B*0702 | 0.75 | Y | Y | 802 (0.79%) | |||
| FSS/MR-M4–12 | LPSHSTRKL | B*0702 | 0.4 | Y | Y | Y | 235 | 398 (0.28%) | |
| FSS/MR-E38–45 | KPTVDIEL | B*0702 | 1.45 | Y | Y | Y | 103 | 51 | |
| FSS/MR-E170–178 | TPNSPRAEA | B*0702 | 0.8 | Y | Y | Y | 155 | ||
| FSS/MR-E173–180 | SPRAEATL | B*0702 | 0.25 | Y | Y | Y | 167 | 108 | |
| FSS/MR-E233–242 | TPHWNNKEAL | B*0702 | 0.35 | Y | Y | Y | 158 | 218 (0.3%) | |
| FSS-E337–347 | GPCKVPAQMAV | B*0702 | 1.7 | Y | Y | 271 | |||
| FSS/MR-NS199–107 | GPQRLPVPV | B*0702 | 1.4 | Y | Y | Y | 103 | ||
| MR-NS2A31–40 | VVMILGGFSM | B*0702 | 1.6 | Y | 106 (0.37%) | 51 | |||
| FSS/MR-NS2A75–84 | RPALLVSFIF | B*0702 | 1.6 | Y | Y | Y | 713 (0.53%) | 582 (2.04%) | 88 (0.58%) |
| FSS/MR-NS2A89–99 | TPRESMLLAL | B*0702 | 0.15 | Y | Y | Y | 807 (0.56%) | 823 (0.95%) | 55 |
| FSS-NS2A133–141 | VPRTDNITL | B*0702 | 0.3 | Y | Y | 786 (0.9%) | 1,182 (1.07%) | ||
| MR-NS2A133–141 | VPRTDNIAL | B*0702 | 0.2 | Y | 648 (0.54%) | 776 (1.39%) | |||
| MR-NS2A141–150 | LPILAALTPL | B*0702 | 0.2 | Y | 540 (0.28%) | ||||
| FSS/MR-NS2A148–155 | TPLARGTL | B*0702 | 0.45 | Y | Y | Y | 706 (0.28%) | 1,055 (0.87%) | |
| FSS/MR-NS2B68–75 | SPRLDVAL | B*0702 | 0.6 | Y | Y | Y | 963 (0.56%) | 993 (0.77%) | |
| FSS/MR-NS3188–196 | LPEIVREAI | B*0702 | 1.2 | Y | Y | Y | 179 (0.23%) | 63 | |
| FSS/MR-NS3206–215 | APTRVVAAEM | B*0702 | 0.2 | Y | Y | Y | 749 (0.3%) | 724 (0.53%) | 606 (0.33%) |
| FSS/MR-NS3309–317 | FPDSNSPIM | B*0702 | 1.2 | Y | Y | Y | 335 | 565 (0.49%) | |
| FSS/MR-NS3405–413 | RVIDSRRCL | B*0702 | 1.1 | Y | Y | Y | 236 (0.23%) | 718 (0.4%) | |
| FSS/MR-NS3427–436 | GPMPVTHASA | B*0702 | 0.25 | Y | Y | Y | 140 | 204 | 136 |
| FSS/MR-NS3492–501 | RPEADKVAAI | B*0702 | 0.55 | Y | Y | Y | 157 | ||
| FSS/MR-NS3574–582 | KPRWMDARV | B*0702 | 0.3 | Y | Y | Y | 969 (0.79%) | 1,095 (1.9%) | 350 (0.63%) |
| FSS/MR-NS3581–589 | RVCSDHAAL | B*0702 | 1.3 | Y | Y | Y | 411 (0.31%) | 44 | |
| MR-NS3596-NS4A1 | AAGKRGAAL | B*0702 | 0.6 | Y | 148 | ||||
| FSS/MR-NS4A36–45 | RPYKAAAAQL | B*0702 | 0.25 | Y | Y | Y | 120 | ||
| FSS/MR-NS4A125–133 | SPQDNQMAI | B*0702 | 0.5 | Y | Y | Y | 159 | ||
| FSS/MR-NS4B35–44 | RPASAWAIYA | B*0702 | 0.35 | Y | Y | Y | 180 (0.49%) | ||
| FSS/MR-NS4B105–115 | TPLTLIVAIIL | B*0702 | 1.55 | Y | Y | Y | 163 (0.4%) | ||
| FSS/MR-NS4B210–220 | SPNKYWNSSTA | B*0702 | 0.45 | Y | Y | Y | 195 | 49 | |
| FSS/MR-NS4B426–435 | RPGAFCIKVL | B*0702 | 0.3 | Y | Y | Y | 1,223 (2.28%) | 1,362 (2.36%) | 90 (0.3%) |
| FSS/MR-NS561–70 | APTQGSASSL | B*0702 | 0.3 | Y | Y | Y | 98 | 47 | |
| FSS/MR-NS5140–149 | RPRVCTKEEF | B*0702 | 0.1 | Y | Y | Y | 162 | 311 (0.24%) | 0.28% |
| MR-NS5332–341 | RPAEGGKTVM | B*0702 | 0.1 | Y | 277 (0.35%) | ||||
| FSS/MR-NS5539–546 | VPTGRTTW | B*0702 | 0.4 | Y | Y | Y | 121 (0.25%) | 54 (0.18%) | |
| FSS/MR-NS5587–596 | IPYLGKREDL | B*0702 | 0.95 | Y | Y | Y | 178 | 438 (0.26%) | |
| FSS/MR-NS5605–614 | RPRTTWAENI | B*0702 | 0.25 | Y | Y | Y | 383 (0.22%) | 641 (0.55%) | |
| FSS-C125-prM8 | VTRRGNAYY | A*0101 | 0.7 | Y | 266 | ||||
| FSS/MR-prM40–48 | HMCDATMSY | A*0101 | 1 | Y | Y | Y | 288 | ||
| FSS/MR-E81–90 | YLDKQSDTQY | A*0101 | 0.25 | Y | Y | Y | 495 | 258 | |
| FSS/MR-E159–167 | ETDENRAKV | A*0101 | 1.3 | Y | Y | Y | 512 | 516 | |
| FSS/MR-E195–203 | GLDFSDLYY | A*0101 | 0.2 | Y | Y | Y | 993 | 647 | |
| FSS/MR-E198–206 | FSDLYYLTM | A*0101 | 0.25 | Y | Y | Y | 398 | 388 | |
| FSS/MR-E377–386 | ELDPPFGDSY | A*0101 | 0.25 | Y | Y | Y | 286 | ||
| FSS/MR-NS123–31 | DVEAWRDRY | A*0101 | 0.95 | Y | Y | Y | 583 | 260 | |
| FSS/MR-NS3272–280 | FTDPSSIAA | A*0101 | 0.25 | Y | Y | Y | 286 | ||
| FSS/MR-NS4A14–22 | MTERFQEAI | A*0101 | 1.15 | Y | Y | Y | 266 | ||
| FSS/MR-NS4B231–239 | YLAGASLIY | A*0101 | 0.55 | Y | Y | Y | 1,646 (0.60%) | 1,574 (1.38%) | |
| FSS/MR-NS4B270–278 | MSALEFYSY | A*0101 | 0.3 | Y | Y | Y | 378 | ||
| FSS/MR-NS5509–517 | YAQMWQLLY | A*0101 | 0.2 | Y | Y | Y | 313 | 225 |
Ninety HLA-A*0101-binding epitope candidates were also chosen for synthesis. The numbers of peptides in C, prM, M, E, NS1, NS2A, NS2B, NS3, NS4A, NS4B and NS5 were 2, 5, 1, 10, 11, 3, 3, 18, 1, 20 and 16, respectively. In contrast to HLA-B*0702 mice, FSS13026 induced a stronger CD8+ T cell response than MR766 in _Ifnar_−/− HLA-A*0101 transgenic mice (Supplementary Fig. 1d–f). As shown in Table 1, the frequencies of peptide-specific IFN-γ-producing CD8+ T cells in mice infected with ZIKV FSS13025 and ZIKV MR766 ranged from 286 to 1,646 SFCs per 106 CD8+ T cells and 225 to 1,574 SFCs per 106 CD8+ T cells, respectively. Thirteen peptides were identified as positive in total. The two ZIKV strains shared 7 positive peptides, including 4 immunodominant peptides: FSS/MR-E159–167, FSS/MR-E195–203, FSS/MR-NS123–31 and FSS/MR-NS4B231–239 (Table 1). E protein contained the largest number of epitopes and induced the highest magnitude of CD8+ T cell response. The most immunodominant epitope, FSS/MR-NS4B231–239, was contained in NS4B (Supplementary Fig. 1d–f).
Epitope confirmation and characterization of cytokine secretion
To further characterize the epitopes identified via IFN-γ ELISPOT analysis, splenocytes were isolated from ZIKV-infected _Ifnar_−/− HLA-B*0702 transgenic mice, stimulated with each of 37 positive peptides, and the frequency of IFN-γ- and/or tumour necrosis factor-α (TNF-α)-producing CD3+ CD8+ T cells was determined by ICS (Supplementary Fig. 2). The percentages of IFN-γ-producing CD8+ T cells ranged from 0.22 to 2.28% and 0.23 to 2.36% of total CD3+ CD8+ T cells in ZIKV FSS13025- and ZIKV MR766-infected mice, respectively (Table 1). Twenty-five of the 37 IFN-γ ELISPOT-positive peptides were confirmed by IFN-γ ICS. Some of the FSS/MR-NS2A89–99 and FSS/MR-NS4B426–435 peptide-stimulated CD8+ T cells simultaneously secreted both IFN-γ and TNF-α in mice infected with either ZIKV FSS13025 or ZIKV MR766. In ZIKV FSS13025-infected mice, FSS-C25–35 also stimulated production of both IFN-γ and TNF-α (Fig. 1a). In ZIKV MR766-infected mice, four additional peptides (FSS/MR-NS2A75–84, FSS/MR-NS2A148–155, FSS/MR-NS2B68–75 and FSS/MR-NS3574–582) stimulated simultaneous production of IFN-γ and TNF-α (Fig. 1b). Collectively, these results define the specificity of the anti-ZIKV CD8+ T cell response restricted by HLA-B*0702 in this mouse model. The epitope map shows that NS proteins NS2A, NS3, NS4B and NS5 are the major targets of the HLA-B*0702-restricted CD8+ T cell response to both African- and Asian-lineage ZIKV, and that the African-lineage ZIKV MR766 contained more HLA-B*0702-restricted CD8+ T cell epitopes than the Asian-lineage ZIKV FSS13025. However, among 13 IFN-γ-ELISPOT-positive HLA-A*0101-binding peptides, only FSS/MR-NS4B231–239 was confirmed by IFN-γ ICS in mice infected with either FSS13025 or MR766 (Fig. 1c,d). The percentages of IFN-γ-producing CD8+ T cells in ZIKV FSS13025- and ZIKV MR766-infected _Ifnar_−/− HLA-A*0101 transgenic mice were 0.6 and 1.38%, respectively (Table 1).
Figure 1. Cytokine secretion pattern of CD8+ T cells directed to HLA-B*0702- and HLA-A*0101-binding ZIKV epitopes identified via IFN-γ ELISPOT.
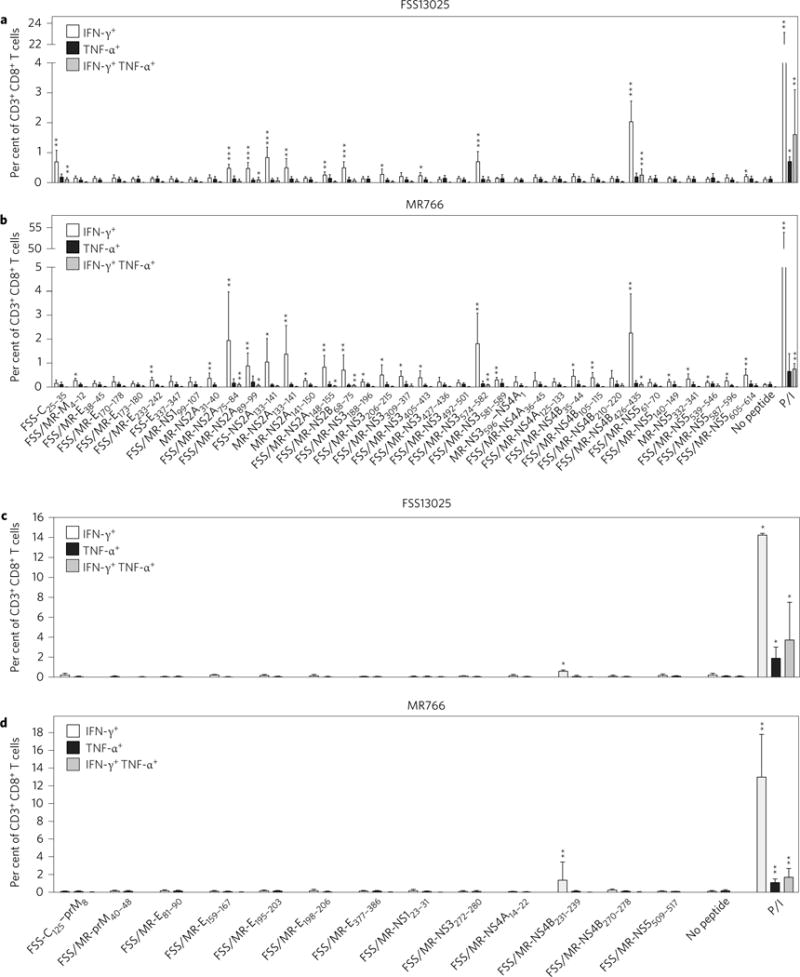
a–d, Splenocytes isolated from _Ifnar_−/− HLA-B*0702 transgenic mice (a,b) and _Ifnar_−/− HLA-A*0101 transgenic mice (c,d) 7 days after r.o. infection with 1 × 102 FFU of ZIKV strain FSS13025 or MR766 were stimulated with each of the 37 positive HLA-B*0702-binding peptides or 13 positive HLA-A*0101-binding peptides identified via IFN-γ ELISPOT and an ICS assay was then performed. Data represent the average of two independent experiments and are expressed as mean ± s.e.m. *P < 0.05, **P < 0.01, ***P < 0.001. Two-tailed Mann–Whitney test. P/I denotes phorbol myristate acetate/ionomycin. Supplementary Tables 2 and 3 give exact values of n and P.
Cross-reactivity of ZIKV epitopes with DENV
To evaluate the potential cross-reactivity of the HLA-B*0702-restricted ZIKV-derived epitopes with DENV, CD8+ T cells from spleens of _Ifnar_−/− HLA-B*0702 transgenic mice infected with DENV2 strain S221 were stimulated by each of 37 ZIKV-derived epitopes identified by IFN-γ ELISPOT analysis. Cross-reactivity was determined using both IFN-γ ELISPOT and IFN-γ ICS assays (Supplementary Fig. 3). Thirteen peptides were positive, as determined by IFN-γ ELISPOT (Supplementary Fig. 3a), and the frequency of epitope-specific IFN-γ+ CD8+ T cells ranged from 44 to 606 SFCs per 106 CD8+ T cells (Table 1). Six peptides were positive based on IFN-γ ICS assay, and the percentages of IFN-γ-producing CD8+ T cells ranged from 0.18 to 0.63% (Supplementary Fig. 3b and Table 1). Of these six IFN-γ ICS-positive peptides, five epitopes (FSS/MR-NS2A75–84, FSS/MR-NS3206–215, FSS/MR-NS3574–582, FSS/MR-NS4B426–435 and FSS/MR-NS5539–546) were positive in both IFN-γ ELISPOT and IFN-γ ICS assays (Table 1). These results thus identified at least five HLA-B*0702-restricted ZIKV/DENV cross-reactive epitopes with as many as 13 additional HLA-B*0702-restricted epitopes possibly being cross-reactive, as determined by the IFN-γ ELISPOT assay alone. Thirteen positive HLA-A*0101-binding peptides were tested in DENV2 S221-infected _Ifnar_−/− HLA-A*0101 transgenic mice, but no positive peptide was found by IFN-γ ICS (Supplementary Fig. 4).
Immunodominance of cross-reactive memory CD8+ T cells during ZIKV infection of DENV-immune mice
The majority of people in the Latin American countries with recent ZIKV outbreaks have previously been exposed to DENV34. To investigate how prior exposure to DENV impacts the ZIKV-specific CD8+ T cell response, _Ifnar_−/− HLA-B*0702 transgenic mice were infected with DENV2 strain S221 for 4 weeks, followed by challenge of these DENV-immune mice with ZIKV FSS13025. On day 3 post-ZIKV infection (a time point that is too early for the development of a DENV-specific naive CD8+ T cell response in mouse models35,36), splenocytes from mock-infected and DENV2-immune mice were stimulated by each of 23 ZIKV epitopes that were identified by IFN-γ ICS. In naive mice, no significant epitope-specific CD8+ T cells were induced (Fig. 2a). In DENV2-immune mice, IFN-γ-expressing CD8+ T cells directed to the three ZIKV/DENV cross-reactive peptides (MR-NS2A31–40, FSS/MR-NS2A75–84 and FSS/MR-NS3574–582) were detected (Fig. 2b). The percentages of these three epitope-specific IFN-γ-producing CD8+ T cells were 0.42 ± 0.12%, 0.44 ± 0.24% and 2.61 ± 1.19%, respectively. FSS/MR-NS2A75–84- and FSS/MR-NS3574–582-specific CD8+ T cells were double-positive with expression of both IFN-γ and TNF-α. Similar analysis on day 7 post-ZIKV infection revealed that eight ZIKV-specific and four ZIKV/DENV cross-reactive CD8+ T cell responses were induced in naive mice (Fig. 2c); in comparison, two ZIKV-specific and five ZIKV/DENV cross-reactive CD8+ T cell responses were elicited in DENV2-immune mice (Fig. 2d). Figure 2e summarizes the frequency and magnitude of ZIKV-specific, ZIKV/DENV cross-reactive and total epitope-specific (both ZIKV-specific and ZIKV/DENV cross-reactive) CD8+ T cell responses in naive versus DENV-immune mice at 3 and 7 days following ZIKV infection. The results demonstrate that, upon ZIKV challenge of DENV-immune mice, memory ZIKV/DENV cross-reactive CD8+ T cells are activated, and the immunodominance pattern of the CD8+ T cell response to ZIKV infection is altered relative to naive mice. The anti-ZIKV CD8+ T cell response in naive mice is broad and includes recognition of both ZIKV-specific and cross-reactive epitopes, whereas the CD8+ T cell response to ZIKV infection in DENV-immune mice is directed to cross-reactive epitopes.
Figure 2. Impact of prior DENV2 infection on the ZIKV-specific CD8+ T cell response.
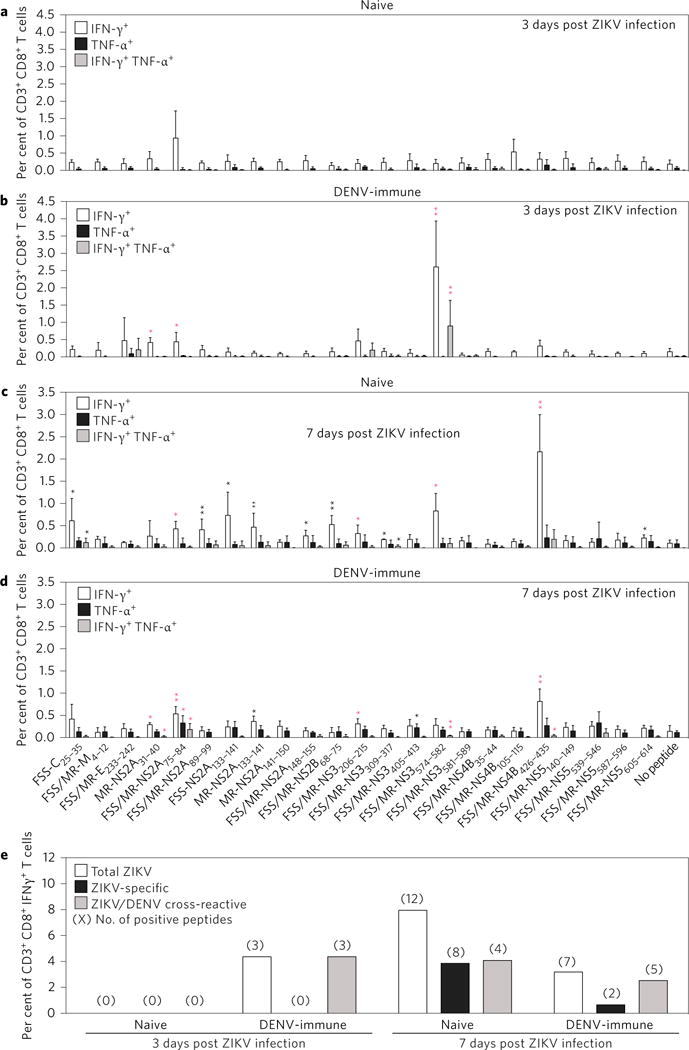
a–d, _Ifnar_−/− HLA-B*0702 transgenic mice were inoculated i.p. with 2 × 103 FFU of DENV2 strain S221 for 4 weeks. Naive mice and DENV2 strain S221-immune mice were challenged r.o. with 1 × 104 FFU of ZIKV FSS13025 for 3 days and the percentages of peptide-specific IFN-γ+ and/or TNF-α+ CD8+ T cells were detected by ICS assay (a,b). In addition, separate groups of naive mice and DENV2 strain S221-immune mice were challenged r.o. with 1 × 104 FFU of ZIKV FSS13025 for 7 days and the percentages of peptide-specific IFN-γ+ and/or TNF-α+ CD8+ T cells were detected by ICS assay (c,d). Data are expressed as mean ± s.e.m. *P < 0.05, **P < 0.01. Two-tailed Mann–Whitney test. Black asterisk indicates ZIKV-specific response; red asterisk represents the ZIKV/DENV cross-reactive response. e, All positive peptides were grouped according to the ZIKV specificity of the immune response. Numbers in parentheses indicate the number of positive peptides in this group. Supplementary Tables 2 and 3 give exact n and P values.
Protective immunity conferred by immunization of mice with ZIKV-specific and ZIKV/DENV cross-reactive peptides
Based on increasing numbers of recent studies using mouse models and human donor samples that implicate a protective role for serotype-cross-reactive CD8+ T cells against DENV13,26,37,38, we next hypothesized that ZIKV/DENV cross-reactive CD8+ T cells play a protective role against ZIKV infection. To directly address the role of ZIKV epitope-specific and ZIKV/DENV cross-reactive CD8+ T cells in protection against ZIKV infection, _Ifnar_−/− HLA-B*0702 transgenic mice were immunized with a cocktail of either 3 ZIKV immunodominant peptides (FSS-NS2A133–141, FSS/MR766-NS2B68–75 and FSS/MR766-NS4B426–435) or four ZIKV/DENV cross-reactive peptides (FSS/MR766-NS4B426–435, FSS/MR766-NS2A75–84, FSS/MR766-NS3206–215 and FSS/MR766-NS3574–582), followed by challenge with ZIKV FSS13025. On day 3 after viral challenge, epitope-specific CD8+ T cells in the spleen were examined via ICS, and viral titres in the serum and brain were assessed via focus-forming assay. We focused on measuring infectious virus in these two tissues because viraemia is a defining feature of human ZIKV infection and the brain appears to be a major target of ZIKV in both fetal and adult infection settings39.
As expected, no significant epitope-specific CD8+ T cell responses were detected in mock-immunized mice (Supplementary Fig. 5a and Fig. 3a,d). In comparison, a significant frequency for each individual epitope-specific CD8+ T cell was observed in mice immunized with the three ZIKV immunodominant epitopes; the percentages of IFN-γ-producing cells among antigen-experienced (CD44+ CD62L−) CD8+ T cells directed to the FSS-NS2A133–141, FSS/MR766-NS2B68–75 and FSS/MR766-NS4B426–435 epitopes were 0.86 ± 0.71%, 0.61 ± 0.47% and 1.63 ± 0.76%, respectively (Fig. 3a). In addition, the levels of infectious virus in the serum and brain of mice immunized with these three ZIKV immunodominant epitopes were significantly lower than those of mock-immunized mice (Fig. 3b,c). Similarly, in mice that were immunized with the four ZIKV/DENV cross-reactive peptides, antigen-experienced (CD44+ CD62L−) CD8+ T cells recognizing four epitopes were detectable: the percentages of FSS/MR766-NS4B426–435-, FSS/MR766-NS2A75–84-, FSS/MR766-NS3206–215- and FSS/MR766-NS3574–582-specific cells were 1.52 ± 0.42%, 3.27 ± 1.91%, 0.66 ± 0.24% and 0.84 ± 0.39%, respectively (Fig. 3d). Infectious ZIKV in both the serum and brain of these mice immunized with the four ZIKV/DENV cross-reactive peptides were significantly lower relative to mock-immunized mice (Fig. 3e,f). Taken together, these results demonstrate that both peptide immunization protocols (that is, the three ZIKV immunodominant and the four ZIKV/DENV cross-reactive epitope cocktails) elicit antigen-experienced, epitope-specific CD8+ T cells upon ZIKV challenge, and reduce infectious ZIKV titres in tissues. Importantly, they indicate that CD8+ T cells recognizing not only ZIKV immunodominant epitopes but also ZIKV/DENV cross-reactive epitopes can contribute to a reduction in infectious ZIKV titres in vivo.
Figure 3. ZIKV-specific and ZIKV/DENV cross-reactive peptide immunization elicited a CD8+ T cell response and mediated protection against ZIKV.
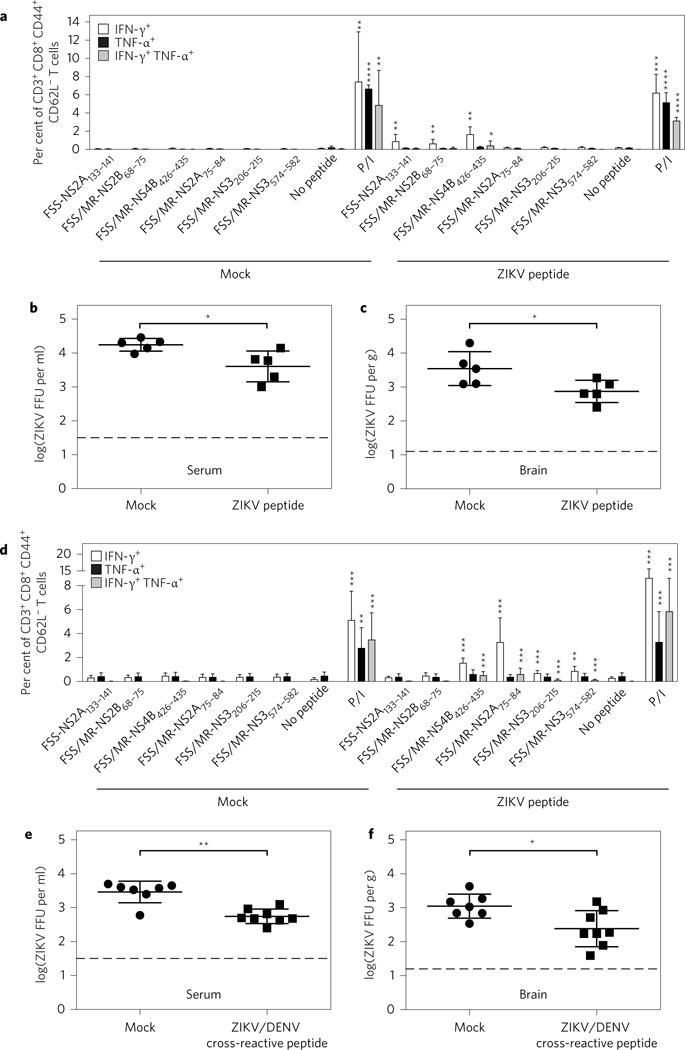
a–f, Five-week-old _Ifnar_−/− HLA-B*0702 transgenic mice were divided into four groups: mock versus ZIKV peptide (a–c) and mock versus ZIKV/DENV cross-reactive peptide (d–f). Peptide groups received corresponding peptide immunizations as described in the Methods. All groups were challenged r.o. with 1 × 104 FFU of ZIKV FSS13025 for 3 days. CD3+ CD8+ CD44+ CD62L− T cells were gated and the percentages of IFN-γ+ and/or TNF-α+ cells were determined by ICS. The levels of infectious ZIKV in sera and brains were measured via FFA. Data are expressed as mean ± s.e.m. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. Two-tailed Mann–Whitney test. Dashed lines represent limit of detection. Supplementary Tables 2 and 3 give exact n and P values.
CD8+ T cell depletion confirms epitope-specific CD8+ T cell-mediated protection
To confirm the protective role of epitope-specific CD8+ T cells in ZIKV infection, we firstly immunized _Ifnar_−/− HLA-B*0702 and HLA-A*0101 transgenic mice with 6 HLA-B*0702-restricted epitopes and 5 HLA-A*0101-restricted epitopes, respectively, then treated the peptide-immunized mice with anti-CD8 antibody to deplete CD8+ T cells. As expected, antigen-experienced CD8+ T cell responses were absent in mock-immunized mice (Figs 4a,b and 5a,b), but were present in peptide-immunized mice (Figs 4c and 5c). Infectious ZIKV levels in the serum, liver and brain of peptide-immunized mice were significantly lower than those in mock-immunized mice (isotype peptide-immunized versus isotype mock in Figs 4e–g and 5e). When CD8+ T cell-depleting anti-CD8 antibody was administered to mice, the majority of peptide-specific CD8+ T cells were absent (Figs 4d and 5d) and infectious ZIKV levels in the serum, liver and brain of CD8+ T cell-depleted peptide-immunized mice were the same as in anti-CD8 antibody-treated mock-immunized mice, but significantly higher than levels in isotype control antibody-treated peptide-immunized mice that were CD8+ T cell-sufficient (anti-CD8 peptide-immunized versus isotype peptide-immunized in Figs 4e–g and 5e). Collectively, these results confirm that protective immunity induced by ZIKV peptide immunization is mediated by CD8+ T cells.
Figure 4. HLA-B*0702-restricted ZIKV peptide immunization-mediated protection is mediated by CD8+ T cells.
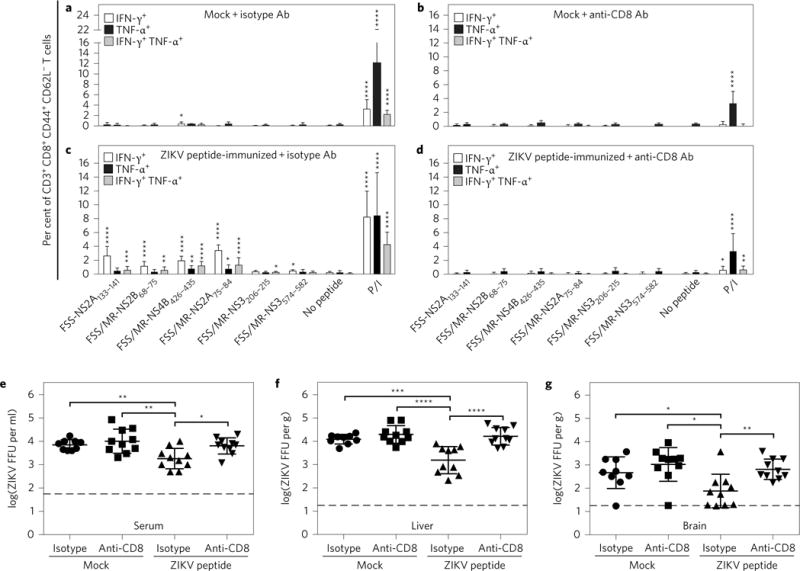
a–g, Five-week-old _Ifnar_−/− HLA-B*0702 transgenic mice were divided into four groups: mock + isotype antibody (Ab) versus mock + anti-CD8 Ab versus peptide-immunized + isotype Ab versus peptide-immunized + anti-CD8 Ab. Peptide-immunized _Ifnar_−/− HLA-B*0702 transgenic mice were immunized with a cocktail of six peptides, as described in the Methods. Mock-immunized mice and peptide-immunized mice were injected i.p. with isotype control Ab and anti-mouse CD8 Ab 3 days and 1 day before ZIKV challenge. All groups were challenged r.o. with 1 × 104 FFU of ZIKV FSS13025 for 3 days. CD3+ CD8+ CD44+ CD62L− T cells were gated and the percentages of IFN-γ+ and/or TNF-α+ cells in mock and peptide groups were determined by ICS. Levels of infectious ZIKV in tissues were measured via FFA. Data are expressed as mean ± s.e.m. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. Two-tailed Mann–Whitney test. Supplementary Tables 2 and 3 give exact n and P values.
Figure 5. HLA-A*0101-restricted ZIKV peptide immunization-mediated protection is mediated by CD8+ T cells.
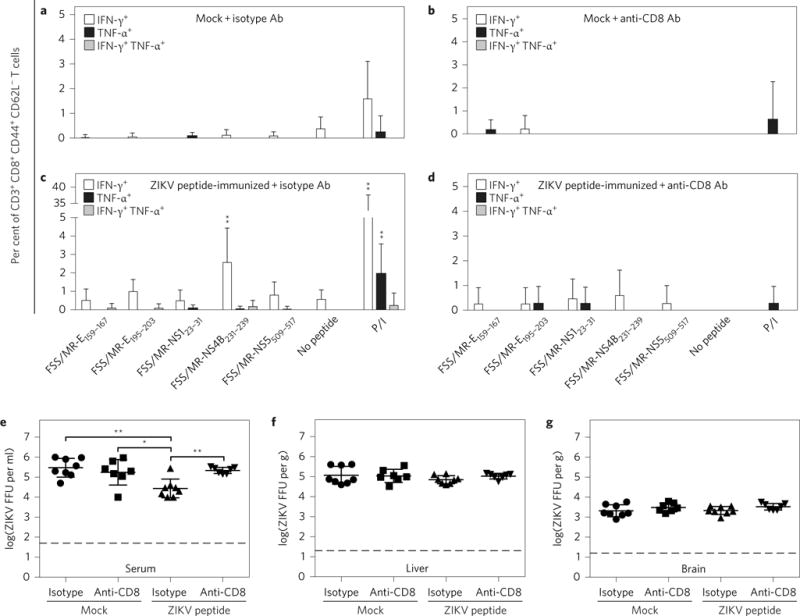
a–g, Five-week-old _Ifnar_−/− HLA-A*0101 transgenic mice were divided into four groups: mock + isotype Ab versus mock + anti-CD8 Ab versus peptide-immunized + isotype Ab versus peptide-immunized + anti-CD8 Ab. Peptide-immunized _Ifnar_−/− HLA-A*0101 transgenic mice were immunized with a cocktail of five peptides as described in the Methods. Mock-immunized mice and peptide-immunized mice were injected i.p. with isotype control Ab and anti-mouse CD8 Ab 3 days and 1 day before ZIKV challenge. All groups were challenged r.o. with 1 × 104 FFU of ZIKV FSS13025 for 3 days. CD3+ CD8+ CD44+ CD62L− T cells were gated and the percentages of IFN-γ+ and/or TNF-α+ cells in mock and peptide groups were determined by ICS. The levels of infectious ZIKV in tissues were measured via FFA. Data are expressed as mean ± s.e.m. *P < 0.05, **P < 0.01. Two-tailed Mann–Whitney test. Supplementary Tables 2 and 3 give exact n and P values.
Discussion
The goals of the present study were to define the specificity and role of ZIKV-specific and ZIKV/DENV cross-reactive CD8+ T cell epitopes that are restricted by common HLA molecules using the _Ifnar_−/− HLA-B*0702 and HLA-A*0101 transgenic mouse models. Prior studies have shown the value of using these mouse models to investigate DENV epitopes of relevance to human T cell responses21,22, and our recent study in LysMCre+Ifnar f l/f l C57BL/6 mice found that H-2b CD8+ T cells protect against ZIKV infection20. Therefore, we first identified CD8+ T cell epitopes that were recognized in _Ifnar_−/− HLA-B*0702 and HLA-A*0101 transgenic mice infected with either the African- or Asian-lineage ZIKV. Most identified epitopes are conserved, not only in ZIKV FSS13025 and ZIKV MR766, but also in the Brazilian outbreak strain ZIKV SPH2015. The majority of 37 HLA-B*0702-restricted CD8+ T cell epitopes identified in ZIKV are located in NS proteins. In contrast, only 13 HLA-A*0101-restricted epitopes were identified and the majority of HLA-A*0101-restricted ZIKV epitopes resided in structural protein E. In comparison, about half of mouse H-2b-restricted ZIKV epitopes were located in structural proteins and the majority of CD8+ T cell responses were to E protein20.
To assess the magnitude and functional quality of cross-reactive T cell responses between ZIKV and DENV, we next identified ZIKV/DENV cross-reactive epitopes using DENV2-infected mice. We tested 37 IFN-γ ELISPOT-confirmed HLA-B*0702-restricted ZIKV epitopes in DENV2-infected mice, and identified 14 peptides reactive with DENV2 in ELISPOT and/or ICS assays, whereas none of the 13 HLA-A*0101-restricted ZIKV epitopes were cross-reactive with DENV2. These 14 HLA-B*0702-restricted ZIKV/DENV cross-reactive epitopes and their DENV2 variants have 0–8 amino-acid substitutions (Supplementary Table 1). It is noteworthy that five DENV2 variants (RPTFAAGLLL, APTRVVAAEM, KPRWLDARI, TPRMCTREEF and LPAIVREAI) had been identified as HLA-B*0702-restricted epitopes in both mouse models and humans21. Three ZIKV peptides and the corresponding DENV2 variants have the same C-terminal amino-acid residue, suggesting that these ZIKV peptides are probably human epitopes as well. Of these five peptides, FSS/MR766-NS3206–215 (APTRVVAAEM) is conserved among many flaviviruses, including ZIKV, four DENV serotypes, West Nile virus, Japanese encephalitis virus, Usutu virus, Murray Valley encephalitis virus and Kunjin virus.
Our investigation of the effect of heterologous DENV/ZIKV infection on HLA-B*0702-restricted T cell response revealed that the ZIKV/DENV cross-reactive CD8+ T cells elicited by prior DENV infection expanded in the early phase of ZIKV challenge and then dominated in the later CD8+ T cell response to ZIKV. Moreover, both ZIKV-specific and ZIKV/DENV cross-reactive CD8+ T cell responses in DENV2-immune mice were weaker in terms of both magnitude and breadth than responses in primary ZIKV infection. These results indicate that prior DENV immunity can affect both the specificity and magnitude of CD8+ T cell response to ZIKV. This phenomenon was also observed during heterotypic DENV infection in mice22 and natural reinfections in humans26,29,30, implying that ZIKV infection in DENV-immune people may behave in a similar manner to heterotypic DENV infection.
In humans, congenital microcephaly and additional birth defects result in part from infection of the fetal neuronal stem cells40,41. Our recent research showed that ZIKV can also infect adult mouse neural progenitor cells, resulting in reduced cell proliferation and cell death39. Therefore, in addition to minimizing viraemia, ZIKV vaccine candidates should protect from brain infection (ZIKV encephalitis). In the present study, we selected six immunodominant HLA-B*0702-restricted epitopes for peptide immunization because (1) these peptides were positive in both IFN-γ ELISPOT and ICS assays for both ZIKV FSS13025 and ZIKV MR766 infection; (2) all six peptides were conserved in both ZIKV FSS13025 and ZIKV SPH2015, while five peptides were also shared by ZIKV MR766; and (3) four peptides were cross-reactive with DENV2, as confirmed by IFN-γ ELISPOT and ICS assays. We then divided these peptides into two groups for immunization: the ZIKV peptide group (two ZIKV-specific peptides and one ZIKV/DENV cross-reactive peptide) and the ZIKV/DENV cross-reactive peptide group (four ZIKV/DENV cross-reactive peptides). As expected based on our recent studies demonstrating a protective role for DENV serotype-specific and cross-reactive HLA-B7-restricted T cell response in mice38, both ZIKV peptide and ZIKV/DENV cross-reactive peptide immunization elicited significant CD8+ T cell responses and reduced infectious ZIKV levels in mouse sera and brains, revealing the potential of these epitopes for preventing ZIKV encephalitis. CD8+ T cell depletion assays in mice immunized with 6 HLA-B*0702- or 5 HLA-A*0101-restricted epitopes further confirmed epitope-specific CD8+ T cell-mediated protection. The finding of cross-reactive peptides, combined with protection against ZIKV seen in our study, raises the possibility of developing a single vaccine that can confer protection against multiple strains of ZIKV and DENV. Although recent studies have demonstrated that vaccination with subunit and inactivated ZIKV strains provides protection42,43, a waning vaccine-induced antibody response, at least in theory, may cause ADE. In domestic mammals, attenuated virus vaccine strains such as the Rift Valley fever vaccine, when administered during pregnancy, have caused teratogenesis and fetal demise44,45. This also highlights the potential importance of using epitope-based ZIKV vaccines as a risk-reduction strategy.
Among the pathogenic human flaviviruses, ZIKV is most closely related to DENV, and these viruses share a high level of amino-acid sequence homology. Accordingly, similar to our recent DENV study with HLA-B*0702 transgenic _Ifnar_−/− mice38, results of this study implicate a protective role for cross-reactive memory T cells. Despite several decades of research, no study to date has provided direct evidence supporting a pathogenic role for T cells during DENV infection. Instead, consistent with our mouse findings20,35–38,46–48, recent studies have begun to support a protective role for DENV-specific T cells in humans. Specifically, recent studies have revealed that the magnitude and breadth of DENV-specific CD8+ and CD4+ T cell responses are HLA-linked and correlate with clinical disease; and HLA-B*0702 and HLA-A*0101, respectively, represent DENV-protective (that is, associated with resistance to dengue disease) and DENV-susceptible alleles26,28,49. Our identification of a greater number of ZIKV-derived HLA-B*0702-restricted epitopes than HLA-A*0101-restricted epitopes, and identification of ZIKV/DENV cross-reactive HLA-B*0702-restricted but not HLA-A*0101-restricted epitopes, suggests that, similar to DENV, the CD8+ T cell response to ZIKV may be HLA-linked.
In summary, this study of ZIKV T cell immunity has identified HLA-B*0702 and HLA-A*0101 epitopes that are conserved between ZIKV lineages and cross-reactive with a DENV serotype. Our HLA transgenic mouse model results show that pre-existing DENV immunity modulates ZIKV-specific CD8+ T cell response development and that ZIKV-specific and ZIKV/DENV cross-reactive CD8+ T cells play protective roles against ZIKV. These results in transgenic mice are probably relevant to human infection, as the same mouse models have been validated in DENV infection by several independent observations. Our results suggest that ZIKV vaccination approaches should include the induction of both ZIKV-specific and ZIKV/DENV cross-reactive CD8+ T cell responses.
Methods
Mice and ethics statement
_Ifnar_−/− HLA-B*0702 and _Ifnar_−/− HLA-A*0101 transgenic mice were previously generated by intercrossing of HLA-B*0702 and HLA-A*0101 transgenic mice with _Ifnar_−/− C57BL/6 mice21. Mice were bred at the La Jolla Institute for Allergy and Immunology under standard pathogen-free conditions. All experiments involving these mice were approved by the Institutional Animal Care and Use Committee under protocol no. AP028-SS1-0615. Sample sizes were estimated based on experiments in similar studies. Animal experiments were not randomized and blinded.
Epitope prediction and peptide synthesis
The HLA-B*0702- and HLA-A*0101-binding peptides were predicted using the IEDAR website (www.iedb.org) online software. Peptides were chosen if their predictive scores ranked in the top 2% of all candidates. A total of 107 HLA-B*0702-binding and 90 HLA-A*0101-binding epitope candidates were synthesized by Synthetic Biomolecules as crude materials, which were confirmed by mass spectrometry analysis. Six immunodominant HLA-B*0702-binding peptides, five HLA-A*0101-binding peptides and a hepatitis C virus (HCV)-core helper peptide (TPPAYRRPPNAPIL, restricted by mouse MHC molecule I-Ab) were synthesized with a purity of >99% and used for immunizing mice. All peptides were dissolved in DMSO with a concentration of 40 mg ml−1 and stored at −20 °C.
Viral strains and mouse infection
Two ZIKV strains, MR766 (Uganda, 1947) and FSS13025 (Cambodia, 2010), were obtained from the World Reference Center for Emerging Viruses and Arboviruses (WRCEVA). The mouse-adapted DENV2 strain S221 is a triple-plaque purified clone derived from DENV2 D2S10 (ref. 36). Both ZIKV and DENV2 were amplified in C6/36 mosquito cells, and viral titres were measured using baby hamster kidney (BHK)-21 cell-based focus forming assay (FFA)20. For epitope screening, 5-week-old mice (female or male) were infected retro-orbitally (r.o.) with either 1 × 102 focus forming units (FFU) of ZIKV FSS13025 or ZIKV MR766, or 2 × 104 FFU of DENV2 S221 in 200 μl 10% FBS/PBS. Seven days after infection, CD8+ T cells were isolated from splenocytes and used for ELISPOT assay, and the splenocytes were directly used for ICS assay. Additionally, 5-week-old mice were inoculated intraperitoneally (i.p.) with 2 × 103 FFU of DENV2 S221 for 4 weeks. DENV2 S221-immune mice were challenged r.o. with 1 × 104 FFU of ZIKV FSS13025 for 3 days or 7 days and the percentages of peptide-specific IFN-γ+ and/or TNF-α+ CD8+ T cells were detected by ICS.
ZIKV challenge of peptide-immunized mice
Two HLA-B*0702-binding ZIKV-specific peptides (FSS-NS2A133–141 and FSS/MR766-NS2B68–75) and four ZIKV/DENV cross-reactive peptides (FSS/MR766-NS4B426–435, FSS/MR766-NS2A75–84, FSS/MR766-NS3206–215 and FSS/MR766-NS3574–582) were chosen for synthesis. Five HLA-A*0101-binding peptides (FSS/MR-E159–167, FSS/MR-E195–203, FSS/MR-NS123–31, FSS/MR-NS4B231–239 and FSS/MR-NS5509–517) were chosen for synthesis. Mice (both female and male, 5–6 weeks of age) were immunized subcutaneously with a mixture of HCV helper peptide (100 μg per mouse) and three or four HLA-B*0702-binding peptides (50 μg peptide per mouse) emulsified in Complete Freund’s Adjuvant (CFA). Mock group mice received the same immunization strategy but without any ZIKV-specific or ZIKV/DENV cross-reactive peptide. On the 21st day, mice were boosted with the same peptide mixtures emulsified in Incomplete Freund’s Adjuvant (IFA). On the 30th day, all mice were challenged r.o. with 1 × 104 FFU of ZIKV FSS13025. Three days post ZIKV infection, mice were killed and serum and spleen were harvested. The splenocytes were used for ICS assay. After cardiac perfusion with PBS, brain was harvested. The levels of infectious ZIKV in serum and brain were measured using FFA.
ZIKV challenge of CD8+ T cell-depleted, peptide-immunized mice
Both six HLA-B*0702- and five HLA-A*0101-binding peptides were used to immunize corresponding mice using the method as described above. Mock and peptide-immunized mice were injected i.p. with either anti-mouse CD8 monoclonal antibody (250 μg per mouse, rat anti-mouse CD8, clone YTS 169.4, Bio X Cell) or isotype control monoclonal antibody (250 μg per mouse, rat IgG2, clone LTF-2, Bio X Cell) at 3 days and 1 day before ZIKV challenge. Mice were injected r.o. with 1 × 104 FFU ZIKV FSS13025. Three days after infection, mice were killed, and spleen and serum were used for ICS assay and FFA, respectively. After cardiac perfusion with PBS, liver and brain were harvested. ZIKV titres in tissues were measured using FFA.
LPS-Blast preparation
LPS-Blasts were prepared as previously described50. Briefly, spleens were harvested from _Ifnar_−/− HLA-B*0702 or _Ifnar_−/− HLA-A*0101 transgenic mice and homogenized through a 70 μm cell strainer. A single-cell splenocyte suspension was placed into a non-vented culture flask with RPMI-1640 complete medium supplemented with 6 μg ml−1 lipopolysaccharide (LPS) and 7 μg ml−1 dextran sulfate. Cells were incubated for 3 days at 37 °C with 5% CO2. Cells were collected and washed three times with RPMI-1640 medium and adjusted to 4 × 106 per ml.
IFN-γ ELISPOT assay
CD8+ T cells were isolated from splenocytes using magnetic bead positive selection (Miltenyi Biotec) 7 days after virus infection. A total of 2 × 105 CD8+ T cells were stimulated with 1 × 105 LPS-blasts loaded with 10 μg of individual peptide in 96-well flat-bottom plates (Immobilon-P, Millipore) that were coated with anti-IFN-γ mAb (clone AN18, Mabtech) in triplicate. Concanavalin A (ConA) was used as positive control. After 20 h of incubation, biotinylated anti-mouse IFN-γ mAb (R4-6A2; Mabtech), followed by ABC peroxidase (Vector Laboratories) and then 3-amino-9-ethylcarbazole (Sigma-Aldrich) were added into the wells. Responses are expressed as number of IFN-γ SFCs per 1 × 106 CD8+ T cells and were considered positive if the magnitude of response was >20 SFCs and had a stimulation index (SI; ratio of test SFCs to control SFCs) of >2. A peptide inducing a magnitude of >500 SFCs per 106 CD8+ T cells was considered an immunodominant peptide.
ICS assay
Spleens were harvested from virus-infected, mock-immunized or peptide-immunized mice. A total of 1 × 106 splenocytes were plated in each well of 96-well U-bottom plates and stimulated with individual peptide (10 μg crude peptide or 1 μg pure peptide per well) for 6 h. Five hours before the end of incubation, Brefeldin A (GolgiPlug, BD Biosciences) was added to the cells. Splenocytes stimulated with PMA-ionomycin were used as a positive control, while cells without any stimulation were used as negative control. After incubation, cells were first stained with PerCpCy 5.5-conjugated anti-CD3 mAb (Clone 145-2C11, TONBO), PE-Cy7-conjugated anti-CD8 mAb (clone 53–67, BD Biosciences), eFluor 450-conjugated anti-CD44 mAb (clone IM7, eBioscience) and APC eFluor 780-conjugated anti-CD62L mAb (clone Mel-14, eBioscience). Cells were then fixed and permeabilized using Cytofix/Cytoperm solution (BD Biosciences), followed by staining with FITC-conjugated anti-IFN-γ mAb (clone XMG 1.2, TONBO) and APC-conjugated anti-TNF-α mAb (clone MP6-XT22, eBioscience). Samples were run using an LSR II (BD Biosciences) and analysed using FlowJo software × 10.0.7 (Tree Star).
Statistical analyses
All data were analysed with Prism software version 6.0 (GraphPad Software) and expressed as mean ± s.e.m. Grubbs’ test was performed to determine whether any of the values was a significant outlier from the rest. Statistical significance was determined using the non-parametric Mann–Whitney test to compare two groups. P < 0.05 was considered significant.
Data availability
The entire panel of peptides that was screened has been submitted to the Immune Epitope Database and Analysis Resource (IEDB) and can be accessed at IEDB_SUBMISSION_ID_1000716. Other data that support the findings of this study are available from the corresponding author upon request.
Supplementary Material
Supplementary Figure 1
Supplementary Figure 2
Supplementary Figure 3
Supplementary Figure 4
Supplementary Figure 5
Supplementary Table 1
Supplementary Table 2
Supplementary Table 3
Acknowledgments
This research was funded by National Institute of Allergy and Infectious Diseases/National Institutes of Health (NIAID/NIH) grant no. 1R01 AI116813 (S.S.), NIH contracts (nos. HHSN272200900042C and HHSN27220140045C, to A.S.) and the La Jolla Institute for Allergy & Immunology institutional support (to S.S.).
Footnotes
Author contributions
J.W. and S.S. designed the project and experiments. A.S. provided the HLA transgenic mice. J.W., W.W.T. and N.S. performed the experiments. J.W., J.E., K.K. and S.S. interpreted the data and wrote the manuscript.
Additional information
Supplementary information is available for this paper.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Competing interests
The authors declare no competing financial interests.
References
- 1.Lazear HM, Diamond MS. Zika virus: new clinical syndromes and its emergence in the western hemisphere. J Virol. 2016;90:4864–4875. doi: 10.1128/JVI.00252-16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Choumet V, Despres P. Dengue and other flavivirus infections. Rev Sci Tech. 2015;34:473–478. 467–472. [PubMed] [Google Scholar]
- 3.Faye O, et al. Molecular evolution of Zika virus during its emergence in the 20th century. PLoS Negl Trop Dis. 2014;8:e2636. doi: 10.1371/journal.pntd.0002636. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Cugola FR, et al. The Brazilian Zika virus strain causes birth defects in experimental models. Nature. 2016;534:267–271. doi: 10.1038/nature18296. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Miner JJ, et al. Zika virus infection during pregnancy in mice causes placental damage and fetal demise. Cell. 2016;165:1081–1091. doi: 10.1016/j.cell.2016.05.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Li C, et al. Zika virus disrupts neural progenitor development and leads to microcephaly in mice. Cell Stem Cell. 2016;19:120–126. doi: 10.1016/j.stem.2016.04.017. [DOI] [PubMed] [Google Scholar]
- 7.Oehler E, et al. Zika virus infection complicated by Guillain–Barre syndrome— case report, French Polynesia, December 2013. Euro Surveill. 2014;19:2. doi: 10.2807/1560-7917.es2014.19.9.20720. [DOI] [PubMed] [Google Scholar]
- 8.D’Ortenzio E, et al. Evidence of sexual transmission of Zika virus. N Engl J Med. 2016;374:2195–2198. doi: 10.1056/NEJMc1604449. [DOI] [PubMed] [Google Scholar]
- 9.Musso D, et al. Potential sexual transmission of Zika virus. Emerg Infect Dis. 2015;21:359–361. doi: 10.3201/eid2102.141363. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Dupont-Rouzeyrol M, et al. Co-infection with Zika and dengue viruses in 2 patients, New Caledonia, 2014. Emerg Infect Dis. 2015;21:381–382. doi: 10.3201/eid2102.141553. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Screaton G, Mongkolsapaya J, Yacoub S, Roberts C. New insights into the immunopathology and control of dengue virus infection. Nat Rev Immunol. 2015;15:745–759. doi: 10.1038/nri3916. [DOI] [PubMed] [Google Scholar]
- 12.Rothman AL, Medin CL, Friberg H, Currier JR. Immunopathogenesis versus protection in dengue virus infections. Curr Trop Med Rep. 2014;1:13–20. doi: 10.1007/s40475-013-0009-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Weiskopf D, Sette A. T-cell immunity to infection with dengue virus in humans. Front Immunol. 2014;5:93. doi: 10.3389/fimmu.2014.00093. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Halstead SB. Dengue. Lancet. 2007;370:1644–1652. doi: 10.1016/S0140-6736(07)61687-0. [DOI] [PubMed] [Google Scholar]
- 15.Huang X, et al. Antibody-dependent enhancement of dengue virus infection inhibits RLR-mediated type-I IFN-independent signalling through upregulation of cellular autophagy. Sci Rep. 2016;6:22303. doi: 10.1038/srep22303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Barba-Spaeth G, et al. Structural basis of potent Zika–dengue virus antibody cross-neutralization. Nature. 2016;536:48–53. doi: 10.1038/nature18938. [DOI] [PubMed] [Google Scholar]
- 17.Dejnirattisai W, et al. Dengue virus sero-cross-reactivity drives antibody-dependent enhancement of infection with Zika virus. Nat Immunol. 2016;17:1102–1108. doi: 10.1038/ni.3515. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Swanstrom JA, et al. Dengue virus envelope dimer epitope monoclonal antibodies isolated from dengue patients are protective against Zika virus. mBio. 2016;7:e01123–16. doi: 10.1128/mBio.01123-16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Stettler K, et al. Specificity, cross-reactivity, and function of antibodies elicited by Zika virus infection. Science. 2016;353:823–826. doi: 10.1126/science.aaf8505. [DOI] [PubMed] [Google Scholar]
- 20.Elong Ngono A, et al. Mapping and role of the CD8+ T cell response during primary Zika virus infection in mice. Cell Host Microbe. 2017;21:35–46. doi: 10.1016/j.chom.2016.12.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Weiskopf D, et al. Insights into HLA-restricted T cell responses in a novel mouse model of dengue virus infection point toward new implications for vaccine design. J Immunol. 2011;187:4268–4279. doi: 10.4049/jimmunol.1101970. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Weiskopf D, et al. Immunodominance changes as a function of the infecting dengue virus serotype and primary versus secondary infection. J Virol. 2014;88:11383–11394. doi: 10.1128/JVI.01108-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Ashour J, et al. Mouse STAT2 restricts early dengue virus replication. Cell Host Microbe. 2010;8:410–421. doi: 10.1016/j.chom.2010.10.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Aguirre S, et al. DENV inhibits type I IFN production in infected cells by cleaving human STING. PLoS Pathog. 2012;8:e1002934. doi: 10.1371/journal.ppat.1002934. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Yu CY, et al. Dengue virus targets the adaptor protein MITA to subvert host innate immunity. PLoS Pathog. 2012;8:e1002780. doi: 10.1371/journal.ppat.1002780. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Weiskopf D, et al. Comprehensive analysis of dengue virus-specific responses supports an HLA-linked protective role for CD8+ T cells. Proc Natl Acad Sci USA. 2013;110:E2046–E2053. doi: 10.1073/pnas.1305227110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Weiskopf D, et al. The human CD8+ T cell responses induced by a live attenuated tetravalent dengue vaccine are directed against highly conserved epitopes. J Virol. 2015;89:120–128. doi: 10.1128/JVI.02129-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Weiskopf D, et al. Human CD8+ T-cell responses against the 4 dengue virus serotypes are associated with distinct patterns of protein targets. J Infect Dis. 2015;212:1743–1751. doi: 10.1093/infdis/jiv289. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Duangchinda T, et al. Immunodominant T-cell responses to dengue virus NS3 are associated with DHF. Proc Natl Acad Sci USA. 2010;107:16922–16927. doi: 10.1073/pnas.1010867107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Rivino L, et al. Differential targeting of viral components by CD4+ versus CD8+ T lymphocytes in dengue virus infection. J Virol. 2013;87:2693–2706. doi: 10.1128/JVI.02675-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Grant A, et al. Zika virus targets human STAT2 to inhibit type I interferon signaling. Cell Host Microbe. 2016;19:882–890. doi: 10.1016/j.chom.2016.05.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Dowall SD, et al. A susceptible mouse model for Zika virus infection. PLoS Negl Trop Dis. 2016;10:e0004658. doi: 10.1371/journal.pntd.0004658. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Lazear HM, et al. A mouse model of Zika virus pathogenesis. Cell Host Microbe. 2016;19:720–730. doi: 10.1016/j.chom.2016.03.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Teixeira MG, Siqueira JB, Jr, Ferreira GL, Bricks L, Joint G, Unnasch TR. Epidemiological trends of dengue disease in Brazil (2000–2010): a systematic literature search and analysis. PLoS Negl Trop Dis. 2013;7:e2520. doi: 10.1371/journal.pntd.0002520. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Prestwood TR, et al. Gamma interferon (IFN-γ) receptor restricts systemic dengue virus replication and prevents paralysis in IFN-α/β receptor-deficient mice. J Virol. 2012;86:12561–12570. doi: 10.1128/JVI.06743-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Yauch LE, et al. A protective role for dengue virus-specific CD8+ T cells. J Immunol. 2009;182:4865–4873. doi: 10.4049/jimmunol.0801974. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Zellweger RM, et al. CD8+ T cells can mediate short-term protection against heterotypic dengue virus reinfection in mice. J Virol. 2015;89:6494–6505. doi: 10.1128/JVI.00036-15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Elong Ngono A, et al. Protective role of cross-reactive CD8 T cells against dengue virus infection. EBioMedicine. 2016;13:284–293. doi: 10.1016/j.ebiom.2016.10.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Li H, et al. Zika virus infects neural progenitors in the adult mouse brain and alters proliferation. Cell Stem Cell. 2016;19:593–598. doi: 10.1016/j.stem.2016.08.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Yockey LJ, et al. Vaginal exposure to Zika virus during pregnancy leads to fetal brain infection. Cell. 2016;166:1247–1256. doi: 10.1016/j.cell.2016.08.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Beckham JD, Pastula DM, Massey A, Tyler KL. Zika virus as an emerging global pathogen: neurological complications of Zika virus. JAMA Neurol. 2016;73:875–879. doi: 10.1001/jamaneurol.2016.0800. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Dowd KA, et al. Rapid development of a DNA vaccine for Zika virus. Science. 2016;354:237–240. doi: 10.1126/science.aai9137. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Larocca RA, et al. Vaccine protection against Zika virus from Brazil. Nature. 2016;536:474–478. doi: 10.1038/nature18952. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Hunter P, Erasmus BJ, Vorster JH. Teratogenicity of a mutagenised Rift Valley fever virus (MVP 12) in sheep. Onderstepoort J Vet Res. 2002;69:95–98. [PubMed] [Google Scholar]
- 45.Adams Waldorf KM, McAdams RM. Influence of infection during pregnancy on fetal development. Reproduction. 2013;146:R151–R162. doi: 10.1530/REP-13-0232. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Yauch LE, et al. CD4+ T cells are not required for the induction of dengue virus-specific CD8+ T cell or antibody responses but contribute to protection after vaccination. J Immunol. 2010;185:5405–5416. doi: 10.4049/jimmunol.1001709. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Zellweger RM, et al. Role of humoral versus cellular responses induced by a protective dengue vaccine candidate. PLoS Pathog. 2013;9:e1003723. doi: 10.1371/journal.ppat.1003723. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Zellweger RM, Eddy WE, Tang WW, Miller R, Shresta S. CD8+ T cells prevent antigen-induced antibody-dependent enhancement of dengue disease in mice. J Immunol. 2014;193:4117–4124. doi: 10.4049/jimmunol.1401597. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Weiskopf D, et al. Dengue virus infection elicits highly polarized CX3CR1+ cytotoxic CD4+ T cells associated with protective immunity. Proc Natl Acad Sci USA. 2015;112:E4256–E4263. doi: 10.1073/pnas.1505956112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Croft M, Carter L, Swain SL, Dutton RW. Generation of polarized antigen-specific CD8 effector populations: reciprocal action of interleukin (IL)-4 and IL-12 in promoting type 2 versus type 1 cytokine profiles. J Exp Med. 1994;180:1715–1728. doi: 10.1084/jem.180.5.1715. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplementary Figure 1
Supplementary Figure 2
Supplementary Figure 3
Supplementary Figure 4
Supplementary Figure 5
Supplementary Table 1
Supplementary Table 2
Supplementary Table 3
Data Availability Statement
The entire panel of peptides that was screened has been submitted to the Immune Epitope Database and Analysis Resource (IEDB) and can be accessed at IEDB_SUBMISSION_ID_1000716. Other data that support the findings of this study are available from the corresponding author upon request.