Rapid “open-source” engineering of customized zinc-finger nucleases for highly efficient gene modification (original) (raw)
. Author manuscript; available in PMC: 2008 Sep 13.
Abstract
Summary
Custom-made zinc-finger nucleases (ZFNs) can induce targeted genome modifications with high efficiency in cell types including Drosophila, C. elegans, plants, and humans. A bottleneck in the application of ZFN technology has been the generation of highly specific engineered zinc-finger arrays. Here we describe OPEN (Oligomerized Pool ENgineering), a rapid, publicly available strategy for constructing multi-finger arrays, which we show is more effective than the previously published modular assembly method. We used OPEN to construct 37 highly active ZFN pairs which induced targeted alterations with high efficiencies (1 to 50%) at 11 different target sites located within three endogenous human genes (VEGF-A, HoxB13, CFTR), an endogenous plant gene (tobacco SuRA), and a chromosomally-integrated EGFP reporter gene. In summary, OPEN provides an “open-source” method for rapidly engineering highly active zinc-finger arrays, thereby enabling broader practice, development, and application of ZFN technology for biological research and gene therapy.
Introduction
A reliable, publicly available method for creating targeted genome modifications with high efficiency would be broadly useful for biological research and gene therapy (Cathomen and Joung, 2008). Although strategies have been developed using viral vectors or short double- or single-stranded DNA molecules to direct homologous recombination (HR), these methods generally have low efficiencies even under optimal cell culture conditions (Cathomen, 2004). An alternative and more efficient strategy involves introducing a double-stranded break (DSB) into a target locus of interest. Repair of this DSB by cellular mechanisms greatly increases rates of HR-mediated gene targeting with an exogenous DNA molecule (Jasin, 1996) or of gene mutation by error-prone non-homologous end-joining (NHEJ) (Bibikova et al., 2002). However, this approach depends on the capability to create “designer” nucleases targeted to specific genomic sequences of interest.
Engineered ZFNs are customized endonucleases that induce site-specific DSBs and genome modifications (as high as 50%) in Drosophila, somatic C. elegans, plant, and human cells (Alwin et al., 2005; Beumer et al., 2006; Bibikova et al., 2003; Bibikova et al., 2002; Lloyd et al., 2005; Lombardo et al., 2007; Miller et al., 2007; Moehle et al., 2007; Morton et al., 2006; Porteus and Baltimore, 2003; Szczepek et al., 2007; Urnov et al., 2005; Wright et al., 2005). ZFNs function as dimers with each monomer composed of a non-specific cleavage domain from the _Fok_I endonuclease fused to a zinc-finger array engineered to bind a target DNA sequence of interest (Cathomen and Joung, 2008; Durai et al., 2005; Porteus and Carroll, 2005). Because a single zinc-finger domain binds a 3 bp subsite, a ZFN dimer can recognize 18 or 24-bp target sites, depending on the number of zinc-fingers in each ZFN monomer.
Publicly available zinc-finger engineering methods described in the literature can be grouped into two general categories. “Modular assembly” involves joining together single fingers with pre-characterized specificities (Bae et al., 2003; Beerli and Barbas, 2002; Liu et al., 2002; Mandell and Barbas, 2006; Segal et al., 2003). Although easy to perform, modular assembly has an efficacy rate for making functional ZFN pairs that is less than 6% (Ramirez et al., 2008) and can yield ZFNs with low activities and/or high toxicities (Cornu et al., 2008; Pruett-Miller et al., 2008). Alternative approaches involve combinatorial selection-based methods that yield multi-finger domains possessing high DNA-binding affinities and specificities (Greisman and Pabo, 1997; Hurt et al., 2003; Isalan et al., 2001) and high activities and low toxicities when expressed as ZFNs in human cells (Cornu et al., 2008; Pruett-Miller et al., 2008). However, selection-based methods require construction and interrogation of large randomized libraries (typically >108 in size) and therefore remain intractable for all but a few labs that possess the required expertise.
Here we describe the development and validation of OPEN (Oligomerized Pool ENgineering), a facile, robust, and publicly available platform for engineering zinc-finger arrays. OPEN is enabled by an archive of zinc-finger pools constructed by the Zinc Finger Consortium, a group of academic laboratories committed to developing engineered zinc-finger technology (http://www.zincfingers.org). The Consortium used OPEN to rapidly engineer 37 ZFN pairs which induce modifications at 11 different sites located within three endogenous human genes (VEGF-A, HoxB13, CFTR), an endogenous plant gene (tobacco SuRA), and the EGFP reporter gene with efficiencies ranging from 1 to 50%. The publicly available OPEN platform will enable routine practice and further development of ZFN technology.
Results
OPEN -- a rapid and robust strategy for engineering zinc-finger arrays
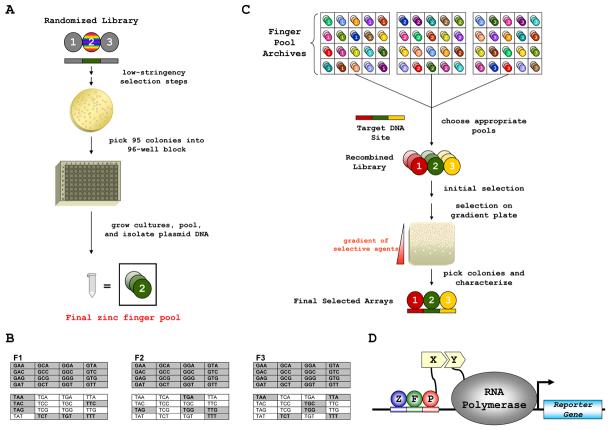
OPEN selections require an archive of pre-selected zinc-finger pools, each containing a mixture of fingers targeted to a different three base pair subsite at a defined position within a three-finger protein (Figure 1A and Experimental Procedures). Fully enabling OPEN will require 192 finger pools (64 potential three bp target subsites for each position in a three-finger protein). In this initial report, we created pools (each containing a maximum of 95 different fingers) targeted to 66 subsites (48 GXX subsites and 18 TXX subsites; Figure 1B). To perform an OPEN selection for a target site, appropriate finger pools from the archive are recombined to create a small library of variants (953=8.6 × 105 members for a three-finger domain) which is interrogated using a bacterial two-hybrid (B2H) selection system in which binding of a zinc-finger domain to its cognate site activates expression of selectable marker genes (Figures 1C & 1D) (Hurt et al., 2003; Joung et al., 2000). To simplify the identification of potential three-finger ZFN sites that can be targeted by OPEN, we created a new version of our web-based ZiFiT software (Sander et al., 2007), ZiFiT 3.0 (Figure S1).
Figure 1. OPEN Method for Engineering Zinc-finger Arrays.
(A) OPEN zinc-finger pool construction. Zinc-finger domains are shown as spheres and associated 3 bp subsites as rectangles. Randomized finger in the library is rainbow colored. Note that the figure illustrates how finger pools for the middle position in a three-finger domain were made, but that pools for amino- or carboxy-terminal fingers were also obtained by building libraries in which finger 1 or finger 3 were randomized, respectively (Experimental Procedures).
(B) GXX and TXX target subsites for which finger pools have been constructed (highlighted in grey).
(C) Schematic overview of OPEN selection for a target DNA site. Zinc-fingers and associated subsites represented as in (A). Details in Supplemental Experimental Procedures.
(D) Schematic of the bacterial two-hybrid (B2H) system. ZFP = zinc-finger protein. X and Y = arbitrary interacting proteins.
Comparing ZFNs made by modular assembly and OPEN
To compare the efficacy of OPEN with modular assembly, we used both strategies to construct multi-finger arrays for five sites in EGFP (ten “half-sites”; Figure S2, Tables S1 and S2) and assessed the DNA-binding activities of these proteins using a quantitative B2H assay (Supplemental Experimental Procedures). Zinc-finger arrays with high affinities and specificities activate transcription by more than three-fold in the B2H system (Hurt et al., 2003). None of the modularly assembled arrays we tested activated transcription by more than three-fold (Figure 2A) even though Western blots showed that all proteins were expressed (data not shown). By contrast, OPEN selections yielded at least one -- in most cases many -- zinc-finger protein that activated transcription by more than three-fold for 9 of the 10 target half-sites (Figure 2A and Table S1).
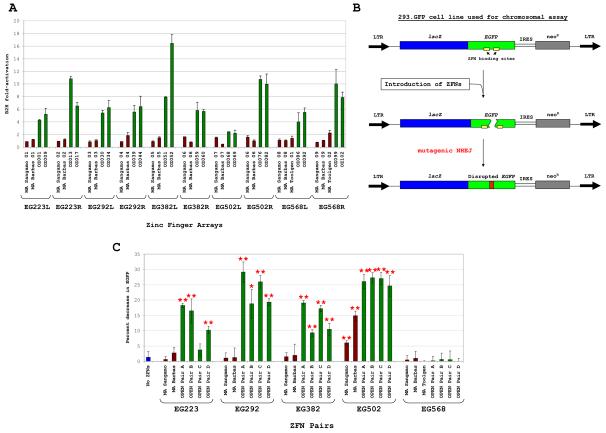
Figure 2. OPEN ZFNs Engineered to Cleave EGFP Gene Sequences.
(A) Quantitative B2H assay of modular assembly (MA; red bars) and OPEN (green bars) zinc finger arrays. Mean fold-activation values (colored bars) and standard deviations (error bars) from three independent assays are shown.
(B) _EGFP_-disruption assay for testing ZFN activities in human cells.
(C) Modularly assembled and OPEN ZFNs assessed using the _EGFP_-disruption assay. Error bars represent standard deviations. Single and double asterisks indicate p values <0.05 or <0.01, respectively.
For each of the five full EGFP target sites, various pairs of modularly assembled and OPEN-selected ZFNs (Table S2) were tested in human cells using an assay in which repair of ZFN-induced DSBs by error-prone NHEJ leads to insertions and deletions in a chromosomally integrated EGFP gene (Figure 2B). Modular assembly yielded ZFN pairs with activities above background for only one of the five sites (EG502) (Figure 2C). By contrast, OPEN yielded ZFN pairs which were active for four of the five full ZFN target sites (EG223, EG292, EG382, and EG502). Although both methods produced active ZFN pairs for the EG502 site, the pairs made by OPEN were more active than those made by modular assembly (Figure 2C). Western blots verified the expression of all ZFNs tested (data not shown).
OPEN selection of zinc-finger arrays that bind to sequences in endogenous human and plant genes
We used ZiFiT 3.0 to identify 14 potential ZFN target sites in three endogenous human genes (VEGF-A, HoxB13, and CFTR) and one endogenous plant gene (tobacco SuRA) (Table S2 and Figure S2). OPEN selections were performed for the 28 “half-sites” within these 14 full ZFN target sites (Tables S1 and S2), and 24 of 28 were deemed successful (Table S1). Using a subset of these finger arrays (Table S2), we constructed ZFN pairs for five sites in VEGF-A, four sites in HoxB13, one site in CFTR, and one site in the SuRA gene.
OPEN ZFNs induce highly efficient mutation of endogenous human and plant genes
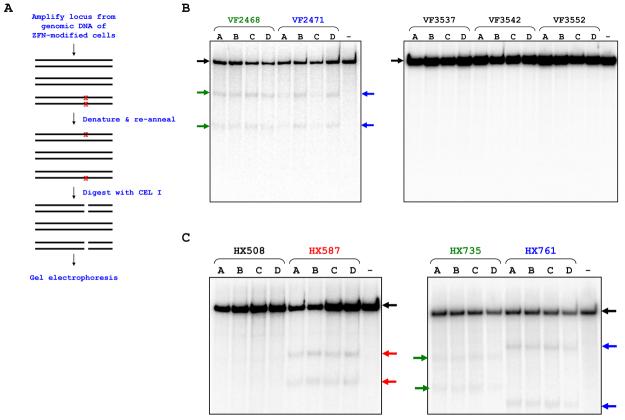
To test whether OPEN ZFNs induce mutations by NHEJ at endogenous human genes, we employed a CEL I nuclease-based mutation detection assay (Figure 3A) (Lombardo et al., 2007; Miller et al., 2007). We found that all four VF2468 and all four VF2471 ZFN pairs we tested generated detectable mutations in the endogenous VEGF-A gene in human 293 cells (Figure 3B). In addition, mutations could be detected at the endogenous HoxB13 gene in 293 cells for all ZFN pairs tested at the HX587, HX735 and HX761 sites (Figure 3C). DNA sequencing of HoxB13 alleles amplified from cells modified by HX587 ZFN pair B (Supplemental Experimental Procedures) revealed mutations at the expected location with an average frequency of 9.6% (Figure S3A). ZFN pairs targeted to three sites in the VEGF-A gene (VF3537, VF3542, VF3552) and to one site in the HoxB13 gene (HX508) failed to induce detectable levels of mutation (Figure 3B and 3C). Due to polymorphisms at the CF877 site in CFTR alleles of 293 cells, we could not assess the CF877 ZFN pairs using the CEL I assay (data not shown). However, DNA sequencing revealed ZFN-induced insertions at the CF877 cleavage site with an average frequency of 1.2% (Figure S3B).
Figure 3. Highly Efficient Mutagenesis of Endogenous Human Genes by OPEN ZFNs.
(A) Schematic of CEL I assay for assaying ZFN-induced mutations.
(B-C) Mutation of the endogenous human VEGF-A gene (B) and HoxB13 gene (C) by OPEN ZFNs. Colored arrows indicate expected CEL I digestion products. Images shown are from representative experiments.
We also tested whether OPEN ZFNs induced mutations in endogenous plant genes. Tobacco protoplasts transformed with a construct encoding the SR2163 ZFN pair were regenerated into individual plants (Experimental Procedures). The SR2163 site is present in both SuRA and SuRB, and each plant was examined for evidence of cleavage at these loci using DNA sequencing. Among 66 transgenic plants surveyed, three had mutationsin SuRA, all of which were deletions of a single base (Figure S3C). In one plant, both alleles of SuRA had the same deletion. No mutated alleles of SuRB were detected. This frequency of mutagenesis by NHEJ (∼2% of potential target alleles) is comparable to what we observed for other OPEN ZFNs in human cells.
OPEN ZFNs induce highly efficient gene targeting at an endogenous human gene
We next tested whether our OPEN ZFNs could induce high efficiency gene targeting at the endogenous human VEGF-A gene. For these experiments, we used VF2468 pair C and VF2471 pair B (Table S2) and directly compared their activities with a previously published ZFN pair (Miller et al., 2007) which cleaves the human IL2Rγ gene. Gene targeting frequencies using _VEGF-A_- or _IL2Rγ_- specific ZFNs and matched donor templates (Figure S4) were measured in human K562 cells using a limited-cycle PCR/restriction digest assay (Experimental Procedures) (Urnov et al., 2005): mean targeting efficiencies of 7.7%, 4.5% and 4.1% were observed with the VF2468, VF2471, and IL2Rγ ZFNs, respectively (Figure 4A). Efficient gene targeting required both the donor construct and ZFN expression vectors (Figure 4A). Control Southern blot assays of cells modified by _VEGF-A_-specific ZFNs confirm that, if anything, the PCR-based assay tended to underestimate gene targeting rates (Figures 4B and S5).
Figure 4. Highly Efficient Gene Targeting of Endogenous Human Loci by OPEN ZFNs.
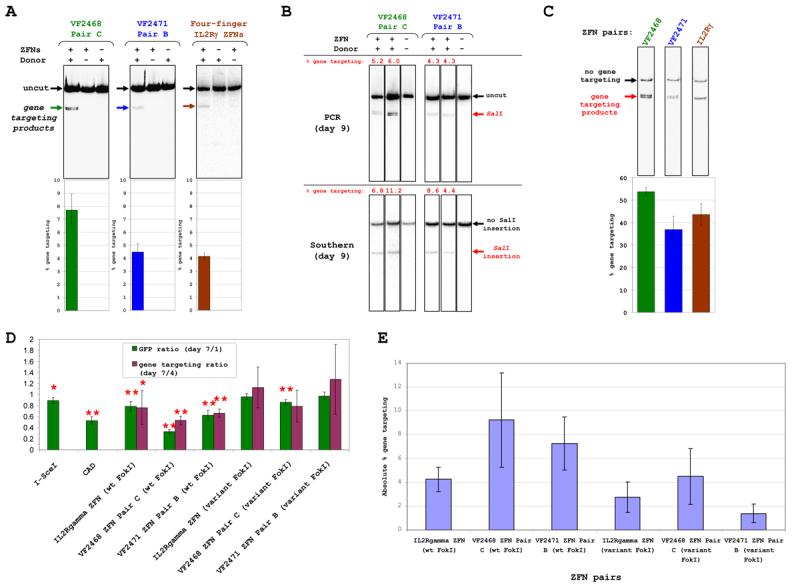
(A) OPEN VEGF-A ZFNs and previously described _IL2Rγ_ZFNs induce efficient gene targeting at endogenous genes in human K562 cells. Top part shows representative gel images from limited-cycle PCR/restriction digest assays and bottom part shows gene targeting frequency means (colored bars) and standard errors (error bars) from multiple experiments.
(B) Gene targeting efficiencies of OPEN VEGF-A ZFNs assessed nine days post-transfection by limited-cycle PCR/restriction digest and Southern blot assays.
(C) Vinblastine enhances gene targeting by OPEN VEGF-A and four-finger _IL2Rγ_ZFNs. Assays performed four days post-transfection. Data presented as in (A).
(D) Toxicities of OPEN VEGF-A and four-finger _IL2Rγ_ZFNs in human K562 cells. Means of GFP (green bars) and gene targeting ratios (purple bars) are shown. Error bars represent standard deviations. Single and double asterisks indicate p values <0.05 or <0.01, respectively.
(E) Gene targeting efficiencies of OPEN VEGF-A and four-finger _IL2Rγ_ZFNs in toxicity experiments of (D). Means and standard deviations (error bars) are shown of PCR-based assays performed four days post-transfection.
Consistent with previous studies (Urnov et al., 2005), we observed higher rates of gene targeting in K562 cells which had been transiently arrested in G2 with vinblastine: 54%, 37%, and 44% mean efficiencies with VF2468, VF2471, and IL2Rγ ZFNs, respectively (Figure 4C). Notably, vinblastine treatment greatly reduced the number of viable cells (data not shown). Sequencing of VEGF-A and IL2Rγ alleles revealed gene targeting events at the expected locations and with the anticipated frequencies (Figure S6), but for all three target sites, we also observed high frequencies of insertion and deletion events at the ZFN cleavage sites (presumably caused by error-prone NHEJ). Unexpectedly, we found that 1.8% and 9.0% of alleles from cells treated with VF2468 and IL2Rγ ZFNs, respectively, contained evidence of both HR-mediated gene targeting and NHEJ-mediated insertion events at a single allele. Interestingly, analysis of single cell clones from vinblastine-treated populations of VEGF-A ZFN-modified cells indicates that gene targeting events are stably maintained (even after 35 days) and strikingly can be induced in as many as four copies of VEGF-A in polyploid cells (Figure S7).
Toxicity profiles of OPEN ZFNs
We compared the relative toxicities of two pairs of three-finger OPEN VEGF-A ZFNs and of a pair of four-finger IL2Rγ ZFNs using cell survival assays in which K562 cells were transfected with ZFN expression vectors, a donor plasmid, and a GFP-expression plasmid. Previous studies demonstrated that toxic ZFNs reduce both the percentage of GFP-positive cells (Cornu et al., 2008; Pruett-Miller et al., 2008) and cells that have undergone gene targeting (Porteus and Baltimore, 2003) over time. All three ZFN pairs we tested showed significant reductions in the relative number of GFP-positive cells by post-transfection day 7 (Figure 4D, green bars) and analogous decreases in the percentage of gene targeting events (Figure 4D, purple bars).
We examined whether toxicity profiles of the IL2Rγ and our OPEN VF2468 and VF2471 ZFNs could be improved by using obligate heterodimeric _Fok_I nuclease domain variants which significantly reduce ZFN-associated toxicity (Miller et al., 2007; Szczepek et al., 2007). Both the variant IL2Rγ ZFN pair and the variant VF2471 ZFN pair showed no significant toxicity as judged by the GFP toxicity assay (Figure 4D, green bars). In addition, the variant VF2468 ZFN pair revealed minimal toxicity (Figure 4D) similar to that of I-_Sce_I, a highly specific meganuclease used previously as a control (Porteus and Baltimore, 2003). Comparable effects were also observed in the relative percentage of gene targeting events (Figure 4D, purple bars). We note that all three variant ZFN pairs induce efficient gene targeting (Figure 4E), demonstrating that the absence of observable toxicity is not due to lack of ZFN activity.
Discussion
The OPEN reagents and methods we describe in this report provide a rapid, highly effective, and publicly available platform for engineering zinc-finger arrays. With OPEN, we created 37 ZFN pairs which mediate highly efficient modification at four different sites within a chromosomally integrated EGFP reporter gene, six different sites within three endogenous human genes (VEGF-A, HoxB13, and CFTR), and one site within an endogenous plant gene (SuRA). The absolute rates of gene modification induced by our OPEN ZFNs ranged from 1 to 50%, and we were able to alter as many as two alleles in a single plant cell and four alleles in a single polyploid human cell.
We found OPEN to be more effective than previously described modular assembly approaches for making zinc-finger arrays. In our studies targeting the EGFP gene, only two of 11 (∼18%) modularly assembled ZFN pairs tested showed activity in human cells, in contrast to 15 of 20 (75%) OPEN ZFN pairs. In addition, at the one target site for which both methods were successful, OPEN ZFNs showed significantly more activity than modularly assembled ZFNs. The low efficacy rate observed with modular assembly is consistent with the results of a large-scale assessment of this method recently conducted by our groups (Ramirez et al., 2008). The higher success rate of OPEN is likely attributable to its consideration of context-dependent effects on DNA-binding among neighboring zinc-fingers (Elrod-Erickson et al., 1996; Isalan et al., 1997; Wolfe et al., 2001; Wolfe et al., 1999) which are largely ignored by modular assembly.
With the set of zinc finger pools described in this report, we estimate that OPEN can be used to engineer three-finger proteins for ∼4.1% ((23*21*22)/(64*64*64)) of all possible 9 bp target sites or ∼0.16% (4.1%*4.1%) of all possible 18 bp ZFN sites. Because ZFNs can bind to sites in which two “half-sites” are separated by “spacer sequences” of five, six or seven base pairs (Bibikova et al., 2001; Porteus and Baltimore, 2003) (K. Wilson and M.H.P., unpublished data), one should be able to find approximately five full ZFN sites in any given kb of random sequence (0.0016 *1000* 3). Thus, with the pools described in this report, one should be able to target multiple ZFN sites within a typical size gene. Important goals for future work will be to generate additional finger pools that expand the targeting range of OPEN and to test whether the approach can also be used to generate arrays composed of more than three-fingers.
While using ZFNs to modify human genes, we observed two limitations that have not been emphasized in previous reports. First, not all zinc-finger arrays that possess sequence-specific DNA-binding activities (as measured in the well-established B2H method) will function as ZFNs in human cells. ZFNs for one of the four sites targeted in the HoxB13 locus, for three of the five sites targeted in the VEGF-A locus, and for one of the five sites targeted in an integrated EGFP reporter gene failed to induce mutagenic NHEJ repair in human cells. In addition, some ZFNs we made to the HoxB13 gene were active in 293 cells (Figure 3C) but not in K562 cells (data not shown). We speculate that the transcriptional status and chromatin configuration of a target site may influence ZFN access to target sites: HoxB13 is transcriptionally active in 293 cells (data not shown) but bears chromatin marks consistent with a repressive state in K562 cells (B. Bernstein, personal communication). Additional studies will be needed to determine whether lack of ZFN activity results from chromatin effects on DNA accessibility or other reasons such as ZFN expression/stability or target site methylation. Second, although the use of vinblastine increased the frequency of gene targeting, DNA sequencing reveals that many alleles still underwent insertions or deletions caused by error-prone NHEJ and that some alleles underwent both a gene targeting event and an insertion. These findings demonstrate limitations in relying solely on PCR- or Southern blot-based assays and suggest that DNA sequencing should always be performed to verify ZFN-induced gene targeting events.
Because OPEN is rapid, reliable, and publicly available, it will foster wider usage and larger-scale applications of engineered zinc-finger technology. OPEN selections are performed in E. coli and do not require specialized equipment. Although our ZFN validation experiments were performed in plant and human cells, OPEN should also be useful for generating ZFNs that function well in other organisms such as zebrafish, mosquitos, Drosophila, and C. elegans. The rapidity and effectiveness of OPEN should enable genome-scale ZFN projects (e.g. developing ZFNs for all human kinase genes or for every zebrafish gene). We note that this report doubles the number of endogenous mammalian genes described in the published literature -- from three (IL2Rg, CCR5, and DHFR) (Lombardo et al., 2007; Santiago et al., 2008; Urnov et al., 2005) to six -- that have been successfully modified using ZFNs. In conclusion, our publicly available OPEN platform will enable scientists to perform the research and development required to move ZFN technology forward for applications in biological research and gene therapy.
Experimental Procedures
Additional details for all methods are provided in Supplemental Experimental Procedures.
Construction of zinc-finger pools
Randomized zinc-finger libraries constructed by cassette mutagenesis were introduced into “B2H selection strains” harboring a 9 bp target site as described (Thibodeau-Beganny and Joung, 2007) and plated on histidine-deficient selective media (NM media) containing 3-AT, a competitive inhibitor of the HIS3 enzyme. Zinc-finger-encoding phagemids were rescued from surviving colonies, re-introduced into fresh B2H selection cells, and plated on NM media containing 3-AT and streptomycin. 95 surviving colonies were inoculated into a 96-well block for growth and plasmids isolated to obtain the final finger pools.
OPEN selections
Each OPEN selection was performed in two steps: First, B2H selection strain cells were infected with randomized zinc-finger phage libraries consisting of three recombined finger pools and then plated on NM media containing 3-AT and streptomycin. Second, zinc finger-encoding phagemids were rescued from surviving colonies, re-introduced into fresh B2H selection strain cells, and plated on NM media containing a gradient of 3-AT and streptomycin. For a small number of the OPEN selections, we performed selections in a single step (see Supplemental Experimental Procedures).
CEL I nuclease assay for NHEJ-mediated mutation
Flp-In T-REx 293 cells (Invitrogen) were transfected with ZFN expression plasmids and genomic DNA was isolated three days post-transfection. Limited-cycle PCR was performed with radiolabeled nucleotides and VEGF-A- or _HoxB13-_specific primers. PCR products were treated with CEL I nuclease and then separated on 10% polyacrylamide gels and visualized using a phosphorimaging screen.
Gene targeting assays
Human K562 cells were transfected with ZFN expression plasmids and donor constructs and genomic DNA harvested four days post-transfection. Limited-cycle PCR was performed with radiolabeled nucleotides and VEGF-A- or _IL2Rγ-_specific primers. PCR products digested with _Sal_I (for VEGF-A) or _BsrB_I (for IL2Rγ) were separated on 10% polyacrylamide gels and visualized using a phosphorimaging screen. Additional details and Southern blot assays are described in Supplemental Experimental Procedures.
Tobacco transformation and assay for mutations
The transformation of tobacco protoplasts by electroporation, selection for kanamycin resistance, and regeneration into plantlets was carried out as previously described (Wright et al., 2005). DNA was prepared from tissue harvested from individual plantlets and SuRA and SuRB alleles were amplified by PCR, gel purified, and sequenced to identify mutations.
ZFN toxicity assays
ZFN expression vectors, donor templates, and plasmid pmaxGFP (Amaxa) were transfected into K562 cells. Cells were assayed for GFP expression at days 1 and 7 post-transfection using a FACScan cytometer and for gene targeting efficiencies at days 4 and 7 post-transfection using the limited-cycle PCR/restriction digest assay.
Supplementary Material
Supplemental M
Acknowledgements
We thank Chong Jin Park for help with statistical calculations and Andrew Hirsh for helpful discussions and comments. J.K.J. is supported by the NIH (R01GM069906, R24GM078369, and R21RR024189), the Cystic Fibrosis Foundation (CFF) (MCCRAY07G0), and the MGH Pathology Service. D.F.V. is supported by the NSF (DBI 0501678). T.C. is supported by the European Commission's 6th Framework Programme (037783 ZNIP). P.B.M. is supported by the CFF (MCCRAY07G0) and the Roy J. Carver Charitable Trust. A.J.I. is supported by the MGH Pathology Service. D.C.S. is supported by the NIH (R01CA112021 and NCI SPORE in Breast Cancer at MGH). M.H.P. is supported by the NIH (R01 HL079295). J.K.J. dedicates this paper to the memory of Robert L. Burghoff and his belief in the power of methods.
References
- Alwin S, Gere MB, Guhl E, Effertz K, Barbas CF, 3rd, Segal DJ, Weitzman MD, Cathomen T. Custom zinc-finger nucleases for use in human cells. Mol Ther. 2005;12:610–617. doi: 10.1016/j.ymthe.2005.06.094. [DOI] [PubMed] [Google Scholar]
- Bae KH, Do Kwon Y, Shin HC, Hwang MS, Ryu EH, Park KS, Yang HY, Lee DK, Lee Y, Park J, et al. Human zinc fingers as building blocks in the construction of artificial transcription factors. Nat Biotechnol. 2003;21:275–280. doi: 10.1038/nbt796. [DOI] [PubMed] [Google Scholar]
- Beerli RR, Barbas CF., 3rd Engineering polydactyl zinc-finger transcription factors. Nat Biotechnol. 2002;20:135–141. doi: 10.1038/nbt0202-135. [DOI] [PubMed] [Google Scholar]
- Beumer K, Bhattacharyya G, Bibikova M, Trautman JK, Carroll D. Efficient gene targeting in Drosophila with zinc-finger nucleases. Genetics. 2006;172:2391–2403. doi: 10.1534/genetics.105.052829. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bibikova M, Beumer K, Trautman JK, Carroll D. Enhancing gene targeting with designed zinc finger nucleases. Science. 2003;300:764. doi: 10.1126/science.1079512. [DOI] [PubMed] [Google Scholar]
- Bibikova M, Carroll D, Segal DJ, Trautman JK, Smith J, Kim YG, Chandrasegaran S. Stimulation of homologous recombination through targeted cleavage by chimeric nucleases. Mol Cell Biol. 2001;21:289–297. doi: 10.1128/MCB.21.1.289-297.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bibikova M, Golic M, Golic KG, Carroll D. Targeted chromosomal cleavage and mutagenesis in Drosophila using zinc-finger nucleases. Genetics. 2002;161:1169–1175. doi: 10.1093/genetics/161.3.1169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cathomen T. AAV vectors for gene correction. Curr Opin Mol Ther. 2004;6:360–366. [PubMed] [Google Scholar]
- Cathomen T, Joung JK. Zinc Finger Nucleases - The Next Generation Emerges. Molecular Therapy. 2008 doi: 10.1038/mt.2008.114. in press. [DOI] [PubMed] [Google Scholar]
- Cornu TI, Thibodeau-Beganny S, Guhl E, Alwin S, Eichtinger M, Joung J, Cathomen T. DNA-binding Specificity Is a Major Determinant of the Activity and Toxicity of Zinc-finger Nucleases. Mol Ther. 2008;16:352–358. doi: 10.1038/sj.mt.6300357. [DOI] [PubMed] [Google Scholar]
- Durai S, Mani M, Kandavelou K, Wu J, Porteus MH, Chandrasegaran S. Zinc finger nucleases: custom-designed molecular scissors for genome engineering of plant and mammalian cells. Nucleic Acids Res. 2005;33:5978–5990. doi: 10.1093/nar/gki912. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Elrod-Erickson M, Rould MA, Nekludova L, Pabo CO. Zif268 protein-DNA complex refined at 1.6 A: a model system for understanding zinc finger-DNA interactions. Structure. 1996;4:1171–1180. doi: 10.1016/s0969-2126(96)00125-6. [DOI] [PubMed] [Google Scholar]
- Greisman HA, Pabo CO. A general strategy for selecting high-affinity zinc finger proteins for diverse DNA target sites. Science. 1997;275:657–661. doi: 10.1126/science.275.5300.657. [DOI] [PubMed] [Google Scholar]
- Hurt JA, Thibodeau SA, Hirsh AS, Pabo CO, Joung JK. Highly specific zinc finger proteins obtained by directed domain shuffling and cell-based selection. Proc Natl Acad Sci U S A. 2003;100:12271–12276. doi: 10.1073/pnas.2135381100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Isalan M, Choo Y, Klug A. Synergy between adjacent zinc fingers in sequence-specific DNA recognition. Proc Natl Acad Sci U S A. 1997;94:5617–5621. doi: 10.1073/pnas.94.11.5617. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Isalan M, Klug A, Choo Y. A rapid, generally applicable method to engineer zinc fingers illustrated by targeting the HIV-1 promoter. Nat Biotechnol. 2001;19:656–660. doi: 10.1038/90264. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jasin M. Genetic manipulation of genomes with rare-cutting endonucleases. Trends Genet. 1996;12:224–228. doi: 10.1016/0168-9525(96)10019-6. [DOI] [PubMed] [Google Scholar]
- Joung JK, Ramm EI, Pabo CO. A bacterial two-hybrid selection system for studying protein-DNA and protein-protein interactions. Proc Natl Acad Sci U S A. 2000;97:7382–7387. doi: 10.1073/pnas.110149297. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu Q, Xia Z, Zhong X, Case CC. Validated zinc finger protein designs for all 16 GNN DNA triplet targets. J Biol Chem. 2002;277:3850–3856. doi: 10.1074/jbc.M110669200. [DOI] [PubMed] [Google Scholar]
- Lloyd A, Plaisier CL, Carroll D, Drews GN. Targeted mutagenesis using zinc-finger nucleases in Arabidopsis. Proc Natl Acad Sci U S A. 2005;102:2232–2237. doi: 10.1073/pnas.0409339102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lombardo A, Genovese P, Beausejour CM, Colleoni S, Lee YL, Kim KA, Ando D, Urnov FD, Galli C, Gregory PD, et al. Gene editing in human stem cells using zinc finger nucleases and integrase-defective lentiviral vector delivery. Nat Biotechnol. 2007;25:1298–1306. doi: 10.1038/nbt1353. [DOI] [PubMed] [Google Scholar]
- Mandell JG, Barbas CF., 3rd Zinc Finger Tools: custom DNA-binding domains for transcription factors and nucleases. Nucleic Acids Res. 2006;34:W516–523. doi: 10.1093/nar/gkl209. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miller JC, Holmes MC, Wang J, Guschin DY, Lee YL, Rupniewski I, Beausejour CM, Waite AJ, Wang NS, Kim KA, et al. An improved zinc-finger nuclease architecture for highly specific genome editing. Nat Biotechnol. 2007;25:778–785. doi: 10.1038/nbt1319. [DOI] [PubMed] [Google Scholar]
- Moehle EA, Rock JM, Lee YL, Jouvenot Y, Dekelver RC, Gregory PD, Urnov FD, Holmes MC. Targeted gene addition into a specified location in the human genome using designed zinc finger nucleases. Proc Natl Acad Sci U S A. 2007;104:3055–3060. doi: 10.1073/pnas.0611478104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morton J, Davis MW, Jorgensen EM, Carroll D. Induction and repair of zinc-finger nuclease-targeted double-strand breaks in Caenorhabditis elegans somatic cells. Proc Natl Acad Sci U S A. 2006;103:16370–16375. doi: 10.1073/pnas.0605633103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Porteus MH, Baltimore D. Chimeric nucleases stimulate gene targeting in human cells. Science. 2003;300:763. doi: 10.1126/science.1078395. [DOI] [PubMed] [Google Scholar]
- Porteus MH, Carroll D. Gene targeting using zinc finger nucleases. Nat Biotechnol. 2005;23:967–973. doi: 10.1038/nbt1125. [DOI] [PubMed] [Google Scholar]
- Pruett-Miller SM, Connelly JP, Maeder ML, Joung JK, Porteus MH. Comparison of Zinc Finger Nucleases for Use in Gene Targeting in Mammalian Cells. Molecular Therapy. 2008 doi: 10.1038/mt.2008.20. doi: 10.1038/mt.2008.1020. [DOI] [PubMed] [Google Scholar]
- Ramirez CL, Foley JE, Wright DA, Muller-Lerch F, Rahman SH, Cornu TI, Winfrey RJ, Sander JD, Fu F, Townsend JA, et al. Unexpected failure rates for modular assembly of engineered zinc-fingers. Nat Methods. 2008;5:374–375. doi: 10.1038/nmeth0508-374. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sander JD, Zaback P, Joung JK, Voytas DF, Dobbs D. Zinc Finger Targeter (ZiFiT): an engineered zinc finger/target site design tool. Nucleic Acids Res. 2007;35:W599–605. doi: 10.1093/nar/gkm349. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Santiago Y, Chan E, Liu PQ, Orlando S, Zhang L, Urnov FD, Holmes MC, Guschin D, Waite A, Miller JC, et al. Targeted gene knockout in mammalian cells by using engineered zinc-finger nucleases. Proc Natl Acad Sci U S A. 2008;105:5809–5814. doi: 10.1073/pnas.0800940105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Segal DJ, Beerli RR, Blancafort P, Dreier B, Effertz K, Huber A, Koksch B, Lund CV, Magnenat L, Valente D, Barbas CF., 3rd Evaluation of a modular strategy for the construction of novel polydactyl zinc finger DNA-binding proteins. Biochemistry. 2003;42:2137–2148. doi: 10.1021/bi026806o. [DOI] [PubMed] [Google Scholar]
- Szczepek M, Brondani V, Buchel J, Serrano L, Segal DJ, Cathomen T. Structure-based redesign of the dimerization interface reduces the toxicity of zinc-finger nucleases. Nat Biotechnol. 2007;25:786–793. doi: 10.1038/nbt1317. [DOI] [PubMed] [Google Scholar]
- Thibodeau-Beganny S, Joung JK. Engineering Cys2His2 Zinc Finger Domains Using a Bacterial Cell-Based Two-Hybrid Selection System. Methods in Molecular Biology. 2007;408:317–334. doi: 10.1007/978-1-59745-547-3_17. [DOI] [PubMed] [Google Scholar]
- Urnov FD, Miller JC, Lee YL, Beausejour CM, Rock JM, Augustus S, Jamieson AC, Porteus MH, Gregory PD, Holmes MC. Highly efficient endogenous human gene correction using designed zinc-finger nucleases. Nature. 2005;435:646–651. doi: 10.1038/nature03556. [DOI] [PubMed] [Google Scholar]
- Wolfe SA, Grant RA, Elrod-Erickson M, Pabo CO. Beyond the “recognition code”: structures of two Cys2His2 zinc finger/TATA box complexes. Structure (Camb) 2001;9:717–723. doi: 10.1016/s0969-2126(01)00632-3. [DOI] [PubMed] [Google Scholar]
- Wolfe SA, Greisman HA, Ramm EI, Pabo CO. Analysis of zinc fingers optimized via phage display: evaluating the utility of a recognition code. J Mol Biol. 1999;285:1917–1934. doi: 10.1006/jmbi.1998.2421. [DOI] [PubMed] [Google Scholar]
- Wright DA, Townsend JA, Winfrey RJ, Jr., Irwin PA, Rajagopal J, Lonosky PM, Hall BD, Jondle MD, Voytas DF. High-frequency homologous recombination in plants mediated by zinc-finger nucleases. Plant J. 2005;44:693–705. doi: 10.1111/j.1365-313X.2005.02551.x. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplemental M